Abstract
Context
RARγ is a therapeutic target for many skin diseases and has potential in cancer treatment. In the current study, we put forward a comprehensive structure–activity relationship study of third and fourth generations of RARγ agonists, addressing multiple crystal structures of RARγ complexes and approved drugs. Adapalene and Trifarotene, through hybrid strategies including protein contacts Atlas analysis, molecular docking, dynamics simulations, MM-GBSA, ASM, and pharmacophore modeling. Our result revealed crucial amino acids Arg267, Ser278, Phe288, Phe230, Met272, Leu271, and Leu268 within the RARγ pocket, as well as pharmacophore features such as two hydrophobic groups, two aromatic rings, and negative ionic features, which are essential for the binding of RARγ agonists. Based on this study, the binding mechanism of RARγ agonists was elucidated, which will be helpful for the rational design of new RARγ agonists for skin diseases and cancer treatment.
Methods
In this study, Schrödinger suite 2021–2 with OPLS_4 force field, Discovery Studio program 3.0, LigandScout 4.3, and PyMOL are utilized in the investigation.
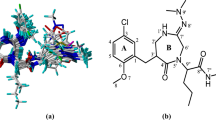
Graphical abstract













Similar content being viewed by others
Data availability
Availability of data and materials is not relevant to the content of my submission.
References
di Masi A, Leboffe L, De Marinis E, Pagano F, Cicconi L, Rochette-Egly C, Lo-Coco F, Ascenzi P, Nervi C (2015) Retinoic acid receptors: from molecular mechanisms to cancer therapy. Mol Aspects Med 41:1–115. https://doi.org/10.1016/j.mam.2014.12.003
Philipp Königshofer, Ksenia Brusilovskaya, Oleksandr Petrenko, Benedikt Silvester Hofer, Philipp Schwabl, Michael Trauner, Thomas Reiberger, Nuclear receptors in liver fibrosis. Biochim Biophys Acta (BBA) - Mol Basis Dis1 1867(12):166235. https://doi.org/10.1016/j.bbadis.2021.166235
Liu Z, Hu Q, Rosenfeld MG (2014) Complexity of the RAR-mediated transcriptional regulatory programs. Subcell Biochem 70:203–225. https://doi.org/10.1007/978-94-017-9050-5_10
A unique secondary-structure switch controls constitutive gene repression by retinoic acid receptor - PubMed. https://pubmed.ncbi.nlm.nih.gov/20543827/ (accessed 2022–06–27).
Nadendla E, Teyssier C, Delfosse V, Vivat V, Krishnasamy G, Gronemeyer H, Bourguet W, Germain P (2015) An unexpected mode of binding defines BMS948 as a full retinoic acid receptor β (RARβ, NR1B2) selective agonist. Plos One 10(5):e0123195. https://doi.org/10.1371/journal.pone.0123195
Klaholz BP, Mitschler A, Belema M, Zusi C, Moras D (2000) Enantiomer discrimination illustrated by high-resolution crystal structures of the human nuclear receptor HRARgamma. Proc Natl Acad Sci U S A 97(12):6322–6327. https://doi.org/10.1073/pnas.97.12.6322
Li B, Cai S-Y, Boyer JL (2021) The role of the retinoid receptor, RAR/RXR heterodimer, in liver physiology. Biochim Biophys Acta Mol Basis Dis 1867(5):166085. https://doi.org/10.1016/j.bbadis.2021.166085
Tan Y, Wang X, Song H, Zhang Y, Zhang R, Li S, Jin W, Chen S, Fang H, Chen Z, Wang K (2021) A PML/RARα direct target atlas redefines transcriptional deregulation in acute promyelocytic leukemia. Blood 137(11):1503–1516. https://doi.org/10.1182/blood.2020005698
Wu M-J, Kim MR, Chen Y-S, Yang J-Y, Chang C-J (2017) Retinoic acid directs breast cancer cell state changes through regulation of TET2-PKCζ pathway. Oncogene 36(22):3193–3206. https://doi.org/10.1038/onc.2016.467
Fukasawa H, Kagechika H, Shudo K (2006) Retinoid therapy for autoimmune diseases. Nihon Rinsho Meneki Gakkai Kaishi 29(3):114–126. https://doi.org/10.2177/jsci.29.114
Kassir M, Karagaiah P, Sonthalia S, Katsambas A, Galadari H, Gupta M, Lotti T, Wollina U, Abdelmaksoud A, Grabbe S, Goldust M (2020) Selective RAR agonists for acne vulgaris: a narrative review. J Cosmet Dermatol 19(6):1278–1283. https://doi.org/10.1111/jocd.13340
Kadigamuwa C, Choksi S, Xu Q, Cataisson C, Greenbaum SS, Yuspa SH, Liu Z-G (2019) Role of retinoic acid receptor-γ in DNA damage-induced necroptosis. iScience 17:74–86. https://doi.org/10.1016/j.isci.2019.06.019
Bauzone M, Souidi M, Dessein A-F, Wisztorski M, Vincent A, Gimeno J-P, Monté D, Van Seuningen I, Gespach C, Huet G (2021) Cross-talk between YAP and RAR-RXR drives expression of stemness genes to promote 5-FU resistance and self-renewal in colorectal cancer cells. Mol Cancer Res MCR 19(4):612–622. https://doi.org/10.1158/1541-7786.MCR-20-0462
Heneghan RE, Starnes BW, Nathan DP, Zierler RE (2016) Renal duplex ultrasound findings in fenestrated endovascular aortic repair for juxtarenal aortic aneurysms. J Vasc Surg 63(4):915–921. https://doi.org/10.1016/j.jvs.2015.10.090
Zhang C, Zhang J, Wang J, Yan Y, Zhang C (2020) Alpha-fetoprotein accelerates the progression of hepatocellular carcinoma by promoting Bcl-2 gene expression through an RA-RAR signalling pathway. J Cell Mol Med 24(23):13804–13812. https://doi.org/10.1111/jcmm.15962
le Maire A, Teyssier C, Balaguer P, Bourguet W, Germain P (2019) Regulation of RXR-RAR heterodimers by RXR- and RAR-specific ligands and their combinations. Cells 8(11):E1392. https://doi.org/10.3390/cells8111392
Renaud JP, Rochel N, Ruff M, Vivat V, Chambon P, Gronemeyer H, Moras D (1995) Crystal structure of the RAR-gamma ligand-binding domain bound to all-trans retinoic acid. Nature 378(6558):681–689. https://doi.org/10.1038/378681a0
Foster RH, Brogden RN, Benfield P (1998) Tazarotene. Drugs 55(5):705–711. https://doi.org/10.2165/00003495-199855050-00008. (discussion 712)
Tolaymat L, Dearborn H, Zito PM (2022) Adapalene. In StatPearls; StatPearls Publishing, Treasure Island (FL)
Cosio T, Di Prete M, Gaziano R, Lanna C, Orlandi A, Di Francesco P, Bianchi L, Campione E (2021) Trifarotene: a current review and perspectives in dermatology. Biomedicines 9(3):237. https://doi.org/10.3390/biomedicines9030237
Tan J, Thiboutot D, Popp G, Gooderham M, Lynde C, Del Rosso J, Weiss J, Blume-Peytavi U, Weglovska J, Johnson S, Parish L, Witkowska D, Sanchez Colon N, Alió Saenz A, Ahmad F, Graeber M, Stein Gold L (2019) Randomized phase 3 evaluation of trifarotene 50 Μg/g cream treatment of moderate facial and truncal acne. J Am Acad Dermatol 80(6):1691–1699. https://doi.org/10.1016/j.jaad.2019.02.044
Klaholz BP, Mitschler A, Moras D (2000) Structural basis for isotype selectivity of the human retinoic acid nuclear receptor11edited by T. Richmond J Mol Biol 302(1):155–170. https://doi.org/10.1006/jmbi.2000.4032
Klaholz BP, Moras D (2002) C-H·O Hydrogen bonds in the nuclear receptor RARγ—a potential tool for drug selectivity. Structure 10(9):1197–1204. https://doi.org/10.1016/S0969-2126(02)00828-6
Structure-based design of Trifarotene (CD5789), a potent and selective RARγ agonist for the treatment of acne - PubMed. https://pubmed.ncbi.nlm.nih.gov/29706423/ (accessed 2022–06–27).
Sastry GM, Adzhigirey M, Day T, Annabhimoju R, Sherman W (2013) Protein and ligand preparation: parameters, protocols, and influence on virtual screening enrichments. J Comput Aided Mol Des 27(3):221–234. https://doi.org/10.1007/s10822-013-9644-8
Friesner RA, Banks JL, Murphy RB, Halgren TA, Klicic JJ, Mainz DT, Repasky MP, Knoll EH, Shelley M, Perry JK, Shaw DE, Francis P, Shenkin PS (2004) Glide: a new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy. J Med Chem 47(7):1739–1749. https://doi.org/10.1021/jm0306430
Santos LHS, Ferreira RS, Caffarena ER (2019) Integrating molecular docking and molecular dynamics simulations. Methods Mol Biol Clifton NJ 2053:13–34. https://doi.org/10.1007/978-1-4939-9752-7_2
Hata H, Phuoc Tran D, MarzoukSobeh M, Kitao A (2021) Binding free energy of protein/ligand complexes calculated using dissociation parallel cascade selection molecular dynamics and Markov state model. Biophys Physicobiology 18:305–316. https://doi.org/10.2142/biophysico.bppb-v18.037
Holman, N. D. M.; Wilkinson, A. J.; Smith, M. C. M. (2021) Alanine-scanning mutagenesis of protein mannosyl-transferase from streptomyces coelicolor reveals strong activity-stability correlation. Microbiol Read Engl 167 (10). https://doi.org/10.1099/mic.0.001103.
Acknowledgements
The work was financially supported by the Overseas Expertise Introduction Project for Discipline Innovation (Grant No. D20029), Program for Innovative Talents of Higher Education of Liaoning (2012520005), and Education Department of Liaoning (2020LJC05).
Funding
The work was financially supported by the Overseas Expertise Introduction Project for Discipline Innovation (Grant No. D20029), Program for Innovative Talents of Higher Education of Liaoning (2012520005), and Education Department of Liaoning (2020LJC05).
Ethics declarations
Ethical approval
Ethical approval is not relevant to the content of my submission.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, H., Hu, B., Luan, J. et al. Structural requirement of RARγ agonism through computational aspects. J Mol Model 29, 108 (2023). https://doi.org/10.1007/s00894-023-05507-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-023-05507-6