Abstract
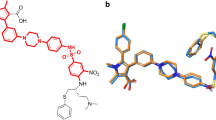
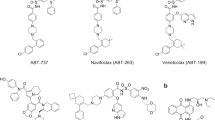
B-cell lymphoma/leukemia gene-2(Bcl-2) protein family known for regulating cell cycle arrest and subsequent cell death is highly expressed in a variety of cancers. Among them, the Bcl-xL and Bcl-2 are two essential proteins in the Bcl-2 family. In the present work, the differences in binding modes as between the two proteins and two ligands ABT-263/43b were investigated and compared. And the computational alanine scanning combined with the recently developed interaction entropy (AS-IE) method was employed for predicting their binding free energies and finding those amino acids that were more critical during the binding process. The result showed that the binding free energy calculated by the AS-IE method was more in line with experimental values than the molecular mechanics/Poisson-Boltzmann surface area (MM/PBSA) method. Besides, no significant difference was found between Bcl-xL and ABT-263/43b in the binding free energy, which Bcl-xL showed slightly weaker binding free energy to 43b because of the fewer number of key residues with interactions. Nonetheless, compared with the Bcl-2 and 43b complex, the Bcl-2 and ABT-263 system had greater number of key residues interacting with ABT-263, in particular, contribute favorably, resulting in a stronger binding ability for the Bcl-2 and ABT-263 systems. The van der Waals and hydrogen bond contributions were significant in the four protein–ligand complexes. Overall, Tyr108 was found to be the common key residues in the Bcl-xL–ligand complex, while Tyr105, Glu100, and Glu143 were established as the common key residue in the Bcl-2–ligand systems. We hope that the predicted hot spot residues and their energy distributions can guide the design of peptide and small-molecule drugs targeting Bcl-xL and Bcl-2.






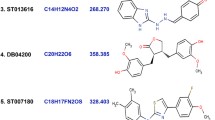
Similar content being viewed by others
Data availability
N/A
Code availability
N/A
References
Kerr JF, Wyllie AH, Currie AR (1972) Apoptosis: a basic biological phenomenon with wide-ranging implications in tissue kinetics. Br J Cancer 26(4):239–257
Kerr JF, Winterford CM, Harmon BV (1994) Apoptosis. Its significance in cancer and cancer therapy. Cancer 73(8):2013–2026
Tsujimoto Y, Cossman J, Jaffe E, Croce C (1985) Involvement of the bcl-2 gene in human follicular lymphoma. Science 228(4706):1440–1443
Soderquist RS, Crawford L, Liu E, Lu M, Agarwal A, Anderson GR et al (2018) Systematic mapping of BCL-2 gene dependencies in cancer reveals molecular determinants of BH3 mimetic sensitivity. Nat Commun 9(1):3513
Inoue-Yamauchi A, Jeng PS, Kim K, Chen HC, Han S, Ganesan YT et al (2017) Targeting the differential addiction to anti-apoptotic BCL-2 family for cancer therapy. Nat Commun 8:16078
Ku B, Liang CY, Jung JU, Oh B-H (2011) Evidence that inhibition of BAX activation by BCL-2 involves its tight and preferential interaction with the BH3 domain of BAX. Cell Res 21(4):627–641
Awan FT, Kay NE, Davis ME, Wu WT, Geyer SM, Leung N et al (2009) Mcl-1 expression predicts progression-free survival in chronic lymphocytic leukemia patients treated with pentostatin, cyclophosphamide, and rituximab. Blood 113(3):535–537
Davids MS, Letai A (2012) Targeting the B-cell lymphoma/leukemia 2 family in cancer. J Clin Oncol 30(25):3127–3135
Adams JM, Cory S (2007) The Bcl-2 apoptotic switch in cancer development and therapy. Oncogene 26(9):1324–1337
Mattson MP (2000) Apoptosis in neurodegenerative disorders. Nat Rev Mol Cell Biol 1(2):120–130
Jian X, Fu Z, Zhang YL, Che XC, Lu R, Yao Z (2012) Subcellular location of antitumor tripeptide-tyroserleutide in human hepatocellular carcinoma cells. Exp Transl Stroke Med 3(2):195–199
Konopleva M, Contractor R, Tsao T, Samudio I, Ruvolo PP, Kitada S et al (2006) Mechanisms of apoptosis sensitivity and resistance to the BH3 mimetic ABT-737 in acute myeloid leukemia. Cancer Cell 10(5):375–388
Tse C, Shoemaker AR, Adickes J, Anderson MG, Chen J, Jin S et al (2008) ABT-263: a potent and orally bioavailable Bcl-2 family inhibitor. Cancer Res 68(9):3421–3428
Varnes JG, Gero T, Huang S, Diebold RB, Ogoe C, Grover PT et al (2014) Towards the next generation of dual Bcl-2/Bcl-xL inhibitors. Bioorg Med Chem Lett 24(14):3026–3033
Souers AJ, Leverson JD, Boghaert ER, Ackler SL, Catron ND, Chen J et al (2013) ABT-199, a potent and selective BCL-2 inhibitor, achieves antitumor activity while sparing platelets. Nat Med 19(2):202–208
Oltersdorf T, Elmore SW, Shoemaker AR, Armstrong RC, Augeri DJ, Belli BA et al (2005) An inhibitor of Bcl-2 family proteins induces regression of solid tumours. Nature 435(7042):677–681
McCammon JA, Gelin BR, Karplus M (1977) Dynamics of folded proteins. Nature 267(5612):585–590
Huang KF, Dong SH, Zhong SS, Li H, Duan LL (2019) Study on the amyloid Aβ42 with accelerated molecular dynamics simulations. Commun Theor Phys 71(9):1121–1126
Dong SH, Luo S, Huang KF, Zhao XY, Duan LL, Li H (2021) Insights into four helical proteins folding via self-guided Langevin dynamics simulation. Mol Phys e1874558
Xue WW, Yang FY, Wang PP, Zheng GX, Chen YZ, Yao XJ et al (2018) What contributes to serotonin–norepinephrine reuptake inhibitors’ dual-targeting mechanism? The key role of transmembrane domain 6 in human serotonin and norepinephrine transporters revealed by molecular dynamics simulation. ACS Chem Neurosci 9(5):1128–1140
Sattarinezhad E, Bordbar AK, Fani N (2017) Virtual screening of piperine analogs as survivin inhibitors and their molecular interaction analysis by using consensus docking, MD simulation, MMPB/GBSA and alanine scanning techniques. J Biomol Struct Dyn 35(8):1824–1832
Nicholls A, Honig B (1991) A rapid finite difference algorithm, utilizing successive over relaxation to solve the Poisson-Boltzmann equation. J Comput Chem 12(4):435–445
Moreira I, Fernandes P, Ramos M (2007) Computational alanine scanning mutagenesis-an improved methodological approach. J Comput Chem 28:644–654
Duan LL, Liu X, Zhang JZH (2016) Interaction entropy: a new paradigm for highly efficient and reliable computation of protein-ligand binding free energy. J Am Chem Soc 138(17):5722–5728
Huang KF, Luo S, Cong YL, Zhong SS, Zhang JZH, Duan LL (2020) An accurate free energy estimator: based on MM/PBSA combined with interaction entropy for protein-ligand binding affinity. Nanoscale 12(19):10737–10750
Cong YL, Huang KF, Li YC, Zhong SS, Zhang JZH, Duan LL (2020) Entropic effect and residue specific entropic contribution to the cooperativity in streptavidin-biotin binding. Nanoscale 12(13):7134–7145
Li YC, Cong YL, Feng GQ, Zhong SS, Zhang JZH, Sun HY et al (2018) The impact of interior dielectric constant and entropic change on HIV-1 complex binding free energy prediction. Struct Dyn 5(6):064101
Luo S, Huang KF, Zhao XY, Cong YL, Zhang JZH, Duan LL (2021) Inhibition mechanism and hot-spot prediction of nine potential drugs for SARS-CoV-2 Mpro by large-scale molecular dynamic simulations combined with accurate binding free energy calculations. Nanoscale 13(17):8313–8332
Yan YN, Yang MY, Ji CG, Zhang JZH (2017) Interaction entropy for computational alanine scanning. J Chem Inf Model 57(5):1112–1122
Liu X, Peng L, Zhou YF, Zhang YZ, Zhang JZH (2018) Computational alanine scanning with interaction entropy for protein–ligand binding free energies. J Chem Theory Comput 14(3):1772–1780
Hornak V, Abel R, Okur A, Strockbine B, Roitberg A, Simmerling C (2006) Comparison of multiple Amber force fields and development of improved protein backbone parameters. Proteins 65(3):712–725
Ryckaert JP, Ciccotti G, Berendsen HJC (1977) Numerical integration of the cartesian equations of motion of a system with constrains: molecular dynamics of n-alkanes. J Comput Phys 23(3):327–341
Pastor RW, Brooks BR, Szabo A (1988) An analysis of the accuracy of Langevin and molecular dynamics algorithms. Mol Phys 65(6):1409–1419
Ryckaert JP, Ciccotti G, Berendsen HJC (1977) Numerical integration of the cartesian equations of motion of a system with constraints: molecular dynamics of n-alkanes. J Comput Phys 23(3):327–341
Homeyer N, Gohlke H (2012) Free energy calculations by the molecular mechanics Poisson-Boltzmann surface area method. Mol Inf 31(2):114–122
Sanner MF, Olson AJ, Spehner JC (1996) Reduced surface: an efficient way to compute molecular surfaces. Biopolymers 38(3):305–320
Nguyen DT, Case DA (1985) On finding stationary states on large-molecule potential energy surfaces. J Phys Chem 89(19):4020–4026
Funding
This work was supported by the National Natural Science Foundation of China (Grant no. 11774207).
Author information
Authors and Affiliations
Contributions
Hao Li performed the data analysis and drafted the manuscript. Shuheng Dong performed the MD simulations and prepared figures. Lili Duan supervised, investigated, and designed the study.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Li, H., Dong, S. & Duan, L. Difference in the binding mechanisms of ABT-263/43b with Bcl-xL/Bcl-2: computational perspective on the accurate binding free energy analysis. J Mol Model 27, 317 (2021). https://doi.org/10.1007/s00894-021-04924-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-021-04924-9