Abstract
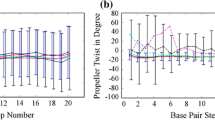
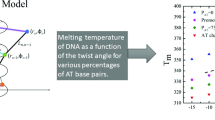
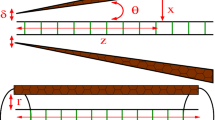
Genomic DNA of higher organisms exists as extremely long polymers, while in bacteria and other lower organisms it is circular with no terminal base pairs. Temperature-induced melting of the DNA double helix by localized strand separation has been unattainable by molecular dynamic simulations due to more rapid fraying of the terminal base pairs in oligomeric DNA. However, local-sequence-dependent unfolding of the DNA double helix is extremely important for understanding various biochemical phenomena, and can be addressed by simulating a model polymeric DNA duplex. Here, we present simulations of polymeric B-DNA of sequence d(CGCGCGCGAATTCGCGCGCG)2 at elevated temperatures, along with its equivalent oligomeric constructs for comparison. Initiation of temperature-induced DNA melting was observed with higher fluctuations of the central d(AATT) region only in the model polymer. The polymeric construct shows a definite melting start site at the weaker A/T stretch, which propagates slowly through the CG rich regions. The melting is reflected in the hydrogen bond breaking, i.e. basepair opening, and by disruption of stacking interaction between successive basepairs. Melting at higher temperature of the oligomer, however, was only through terminal fraying, as also reported earlier.








Similar content being viewed by others
References
Mergny JL, Lacroix L (2003) Oligonucleotides 13:515
Stucki M, Stagljar I, Jonsson ZO, Hubscher U (2001) Prog Nucleic Acid Res Mol Biol 65:261
Kannan S, Zacharias M (2009) Phys Chem Chem Phys 11:10589
Zeng Y, Zocchi G (2006) Biophys J 90:4522
van Erp TS, Cuesta-Lopez S, Peyrard M (2006) Eur Phys J E Soft Matter 20:421
Mandal M, Mukhopadhyay C (2014) Phys Chem Chem Phys 16:21706
Das A, Mukhopadhyay C (2009) Proteins 75:1024
Dastidar SG, Mukhopadhyay C (2005) Phys Rev E Stat Nonlinear Soft Matter Phys 72:051928
Roy S, Basu S, Dasgupta D, Bhattacharyya D, Banerjee R (2015) PLoS One 10:e0142173
Samanta S, Mukherjee S, Chakrabarti J, Bhattacharyya D (2009) J Chem Phys 130:115103
Kundu S, Mukherjee S, Bhattacharyya D (2012) J Biosci 37:445
Bevan DR, Li LP, Pedersen LG, Darden TA (2000) Biophys J 78:668
Cheng Y, Korolev N, Nordenskiold L (2006) Nucleic Acids Res 34:686
Mukherjee S, Kundu S, Bhattacharyya D (2014) J Comput Aided Mol Des 28:735
Luan B, Aksimentiev A (2008) Phys Rev Lett 101:118101
Chandrasekaran R, Arnott S (1996) J Biomol Struct Dyn. 13:1015
MacKerell AD, Banavali NK (2000) J Comput Chem 21:105
Foloppe N, MacKerell AD (2000) J Comput Chem 21:86
Brooks BR, Bruccoleri RE, Olafson BD, States DJ, Swaminathan S, Karplus M (1983) J Comput Chem 4:187
Korolev N, Lyubartsev AP, Laaksonen A, Nordenskiold L (2003) Nucleic Acids Res 31:5971
Dai L, Mu Y, Nordenskiold L, van der Maarel JR (2008) Phys Rev Lett 100:118301
Halder S, Bhattacharyya D (2010) J Phys Chem B 114:14028
Feller SE, Zhang YH, Pastor RW, Brooks BR (1995) J Chem Phys 103:4613
Kale L, Skeel R, Bhandarkar M, Brunner R, Gursoy A, Krawetz N, Phillips J, Shinozaki A, Varadarajan K, Schulten K (1999) J Comput Phys 151:283
Nelson M, Humphrey W, Kufrin R, Gursoy A, Dalke A, Kale L, Skeel R, Schulten K (1995) Computer Phys Commun 91:111
Bansal M, Bhattacharyya D, Ravi B (1995) Computer Appli Biosci 11:281
Mukherjee S, Bansal M, Bhattacharyya D (2006) J Comput Aided Mol Des 20:629
Dickerson R, Bansal M, Calladine CR (1989) EMBO J 8:1
Olson WK, Bansal M, Burley SK, Dickerson RE, Gerstein M, Harvey SC, Heinemann U, Lu XJ, Neidle S, Shakked Z, Sklenar H, Suzuki M, Tung CS, Westhof E, Wolberger C, Berman HM (2001) J Mol Biol 313:229
Pingali PK, Halder S, Mukherjee D, Basu S, Banerjee R, Choudhury D, Bhattacharyya D (2014) J Comput Aided Mol Des 28:851
Mukherjee S, Bhattacharyya D (2013) J Biomol Struct Dyn 31:896
Calladine CR (1982) J Mol Biol 161:343
Drew HR, Samson S, Dickerson RE (1982) Proc Natl Acad Sci USA 79:4040
Dickerson RE, Kopka ML, Pjura P (1983) Proc Natl Acad Sci USA 80:7099
Drsata T, Perez A, Orozco M, Morozov AV, Sponer J, Lankas F (2013) J Chem Theo Comput 9:707
Acknowledgments
We are thankful for partial financial support from the BARD project of Department of Atomic Energy, Government of India and Department of Biotechnology, Government of India.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
ESM 1
(DOC 2492 kb)
Rights and permissions
About this article
Cite this article
Kundu, S., Mukherjee, S. & Bhattacharyya, D. Melting of polymeric DNA double helix at elevated temperature: a molecular dynamics approach. J Mol Model 23, 226 (2017). https://doi.org/10.1007/s00894-017-3398-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-017-3398-5