Abstract
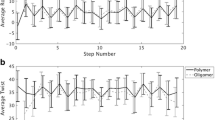
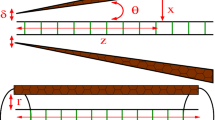
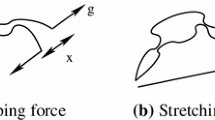
We investigated the melting transition of deoxyribonucleic acid (DNA) embedded in a Langevin fluctuation–dissipation thermal bath. Torsional effects were taken into consideration by introducing a twist angle \(\varphi \) between neighboring base pairs stacked along the molecule backbone. We use the Barbi–Cocco–Payrard model to numerically study the impact of the twist angle on the melting temperature, considering four different sequences composed of 69 base pairs. According to the outcomes of our simulation, for all heterogeneous sequences, an increase in twist angle leads to a linear rise in melting temperature with a positive slope. For angles greater than the so-called equilibrium angle, the DNA chain becomes very rigid against opening and accordingly high temperatures are required to initiate the melting process. We also investigate the opening probability of bubbles, the bubble lifetime profiles and bubble length along the different DNA sequences.
Graphical abstract







Similar content being viewed by others
Data availability statement
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
I. H. G. S. Consortium, Nature 431, 931–945 (2004)
B. Alexandrov, N.K. Voulgarakis, K.Ø. Rasmussen, A. Usheva, A.R. Bishop, J. Phys. Condens. Matter 21, 034107 (2008)
B.S. Alexandrov, V. Gelev, S.W. Yoo, A.R. Bishop, K. Rasmussen, A. Usheva, PLoS Comput. Biol. 5, e1000313 (2009)
Y. Lubelsky, J. Prinz, L. DeNapoli, Y. Li, J. Belsky, D. Macalpine, Genome Res. 24, 1102 (2014)
B. Alberts, A. Johnson, J. Lewis et al., Molecular Biology of the Cell, 4th edn. (Garland Science, New York, 2002). (DNA Replication Mechanisms)
M. Peyrard, A.R. Bishop, Phys. Rev. Lett. 62, 2755 (1989)
T. Dauxois, M. Peyrard, A.R. Bishop, Phys. Rev. E 47, 684 (1993)
S. Zdravkovi’c, J. Nonlinear Math. Phys. 18, 463 (2011)
M. Zoli, J. Chem. Phys. 135, 115101 (2011)
M. Peyrard, Nonlinearity 17, R1 (2004)
S. Ares, N.K. Voulgarakis, K.O. Rasmussen, A.R. Bishop, Phys. Rev. Lett. 94, 035504 (2005)
M. Peyrard, S. Cuesta-L’opez, D. Angelov, J. Phys. Condens. Matter 21, 034103 (2008)
O. Farzadian, M.D. Niry, Phys. A 450, 95–103 (2016)
O. Farzadian, M.D. Niry, Phys. A 505, 49–60 (2018)
M. Hillebrand, G. Kalosakas, A.R. Bishop, C. Skokos, J. Chem. Phys. 155, 095101 (2021)
M. Hillebrand, G. Kalosakas, C. Skokos, A.R. Bishop,Phys. Rev. E 102, 062114 (2020)
M. Barbi, S. Cocco, M. Peyrard, Phys. Lett. A 253, 358 (1999)
S. Cocco, R. Monasson, Phys. Rev. Lett. 83, 5178 (1999)
A. Campa, Phys. Rev. E 63, 021901 (2001)
M. Barbi, S. Lepri, M. Peyrard, N. Theodorakopoulos, Phys. Rev. E 68, 061909 (2003)
M. Zoli, J. Chem. Phys. 138, 205103 (2013)
M. Zoli, Soft Matter 10, 4304 (2014)
M. Manghi, N. Destainville, Phys. Rep. 631, 1 (2016)
J.C. Wang, Proc. Natl. Acad. Sci. 76, 200 (1979)
M. Zoli, J. Chem. Phys. 148, 214902 (2018)
M. Zoli, Phys. A 492, 9039 (2018)
I. Omelyan, I. Mryglod, R. Folk, Comput. Phys. Commun 151, 272 (2003)
H.A. Forbert, S.A. Chin, Phys. Rev. E 63, 016703 (2000)
G. Kalosakas, K.Ø. Rasmussen, A.R. Bishop, C.H. Choi, A. Usheva, EPL 68, 127 (2004)
A.E. Bergues-Pupo, J.M. Bergues, F. Falo, Phys. Rev. E 87, 022703 (2013)
R. Tapia-Rojo, J.J. Mazo, F. Falo, Phys. Rev. E 82, 031916 (2010)
M. Hillebrand, G. Kalosakas, A. Schwellnus, C. Skokos, Phys. Rev. E 99, 022213 (2019)
G. Kalosakas, S. Ares, J. Chem. Phys. 130, 235104 (2009)
B.S. Alexandrov, V. Gelev, Y. Monisova, L.B. Alexandrov, A.R. Bishop, K.Ø. Rasmussen, A. Usheva, Nucleic Acids Res. 37, 2405 (2009)
R. Wells, J. Larson, R. Grant, B. Shortle, C. Cantor, J. Mol. Biol. 54, 465 (1970)
B.S. Alexandrov, L.T. Wille, K.O. Rasmussen, A.R. Bishop, K.B. Blagoev, Phys. Rev. E 74, 050901 (2006)
O. Dahlena, T.S. van Erp, J. Chem. Phys. 142, 235101 (2015)
R.B. Inman, R.L. Baldwin, J. Mol. Biol. 8(4), 452–469 (1964)
B.J. Wood, J.R. Ramkaransingh, T. Fojo, M.M. Walther, S.K. Libutti, Cancer 94, 443–451 (2002)
T. Vanagas, A. Gulbinas, J. Pundzius, G. Barauskas, Medicina (Kaunas) 46, 13–17 (2010)
Acknowledgements
The authors acknowledge the ORAU grant with PN 17098 and the state-targeted program “Center of Excellence for Fundamental and Applied Physics” (BR05236454) by the Ministry of Education and Science of the Republic of Kazakhstan.
Author information
Authors and Affiliations
Contributions
All authors made a significant contribution to the work reported, whether that is in the conception, study design, execution, acquisition of data, analysis or interpretation.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflicts to disclose.
Appendix: Rescaled equation of motion
Appendix: Rescaled equation of motion
The full analytical expression in Eq. (3) is given by
Introducing the dimensionless stretching of the base pairs as \(\tilde{r}_n=\alpha r_n\), \(\tilde{R}_0=\alpha R_0\) and substituting \(b=\frac{\beta }{\alpha }\) and \(a_n=\frac{\alpha _n}{\alpha }\), we can rewrite the equation of motion as
Next, we introduce the dimensionless time \(\tau =\sqrt{\frac{D\alpha ^2}{m}}t\) and the substitutions \(\lambda _n=\frac{D_n\alpha _n}{D\alpha }\), so that
Finally, in the former equation we rewrite the noise term as
which is justified due to Dirac delta function and Gaussian noise properties, \( \langle \xi _n(A\tau )\xi _n(A\tau ^{\prime }) \rangle = A^{-1} \delta (\tau -\tau ^{\prime })\) and \(\xi (A\tau )\rightarrow \frac{1}{\sqrt{A}}\xi (\tau )\), respectively.
By considering
and substituting Eq. (11) into Eq. (10), we are led to Eq. (5) in the text, namely
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Farzadian, O., Oikonomou, T., Moradkhani, M. et al. Model for melting transition of twisted DNA in a thermal bath. Eur. Phys. J. B 96, 23 (2023). https://doi.org/10.1140/epjb/s10051-023-00492-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1140/epjb/s10051-023-00492-4