Abstract
Monocyte-chemoattractant protein-1 (MCP-1), also known as CCL2, is a potent chemoattractant of T cells and monocytes, involved in inflammatory and angio-proliferative brain and retinal diseases. Higher expression of MCP-1 is observed in metastatic tumors. Unusual levels of MCP-1 in the brain may be correlated with autism. Immunochemistry where atomic force microscope (AFM) tips functionalized with appropriate antibodies against MCP-1 are used could in principle support medical diagnostics. Useful signals from single molecule experiments may be generated if interaction forces are large enough. The chemokine-antibody unbinding force depends on a relative motion of the interacting fragments of the complex. In this paper the stability of the medically important MCP-1- immunoglobulin G antibody Fab fragment complex has been studied using steered molecular dynamics (SMD) computer simulations with the aim to model possible arrangements of nano-diagnostics experiments. Using SMD we confirm that molecular recognition in MCP1-IgG is based mainly on six pairs of residues: Glu39A - Arg98H, Lys56A - Asp52H, Asp65A - Arg32L, Asp68A - Arg32L, Thr32A - Glu55L, Gln61A - Tyr33H. The minimum external force required for mechanical dissociation of the complex depends on a direction of the force. The pulling of the MCP-1 antigen in the directions parallel to the antigen-antibody contact plane requires forces about 20 %–40 % lower than in the perpendicular one. Fortunately, these values are large enough that the fast lateral force spectroscopy may be used for effective nano-diagnostics purposes. We show that molecular modeling is a useful tool in planning AFM force spectroscopy experiments.

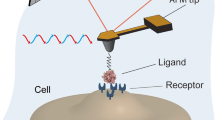
Lateral SMD forces (green arrow) required for mechanical unbinding of MCP-1 chemokine (blue) from Ig G antibody (red/gray) are 20-40% lower than vertical ones (orange arrow)
Similar content being viewed by others
Introduction
The immune system (IS) is a complex network of body organs providing means for defense against pathogens and maintaining integrity of the organism [1]. Due to its medical significance, especially in the contexts of allergies and cancer immunotherapy, IS is vigorously studied [2]. However, our understanding of IS, especially on a molecular level, is still not satisfactory. Recent years brought discoveries of new roles of signaling proteins involved in IS activity modulating the brain [3]. Here we investigate interactions of a small protein MCP-1 which has a diagnostic potential in diseases related to the central nervous system (CNS) and cancer.
Monocyte chemoattractant protein-1 (MCP-1), also known as chemokine ligand 2 (CCL2), is a member of the chemokine family [4]. The chemokines are a type of cytokines, they are divided into two main groups, based on a relative position of the conserved cysteines, which can be either adjacent (CC type) or separated by one amino acid (CXC chemokines). The basic function of those cytokines is to act on chemoattraction in traffic regulation of immune cells. MCP-1 consists of 77 amino acids and belongs to the CC subfamily [5]. It binds to CCR2 and CCR4 receptors [6] but interacts strongly with specialized antibodies as well. MCP-1 is a chemoattrator to monocytes, memory T cells, and dendritic cells [7, 8]. Its main function is to recruit those immune cells to a place of infection or injury. Over a thousand of experimental papers on MCP-1 have been published since its discovery in 1989 [9, 10].
Recent studies show that MCP-1 is also expressed in astrocytes, microglia and neurons [11, 12]. Receptors of MCP-1 are present on surfaces of those CNS cells. The presence of MCP-1 causes migration of leukocytes into the CNS. MCP-1 is present in CNS not only in inflammation but also in the healthy brain [4]. Thus, this chemokine can modulate activity of neurons, astrocytes and microglia [3].
MCP-1 may play a role in many diseases, including multiple sclerosis, rheumatoid arthritis, atherosclerosis, obesity and insulin-resistant diabetes [13]. It has a direct role in angiogenesis and tumor progression [14], promotes prostate cancer tumorigensis and metastasis [15]. The first studies on humans of a new drug CNTO888 (monoclonal antibody) blocking MCP-1 has been very recently published [16]. Other recent studies show that it may also be involved in autism spectrum disorder (ASD) [17–19]. The level of MCP-1 in brain tissues and cerebrospinal fluid [17] in autistic subjects is higher than that in healthy people. Levels of MCP-1 in plasma of children with ASD is elevated [18] as well. Those facts may suggest that the neuroimmune response is a part of the neuropathological processes in ASD and that MCP-1 may play a pathogenic role in this disorder [19] and prompted us to undertake this study.
Atomic force microscopy (AFM) has been proposed as a useful tool for studies of molecular recognition processes [20, 21]. Protein-antibody interactions may be studied using classical contact mode techniques [22–24]. In these methods the antibody is attached to a silanized tip and a vertical force exerted by the AFM cantilever tip dissociates transient antibody-protein complexes [25, 26]. The maximum force registered during withdrawing the cantilever from the surface covered with a sample, is a measure of the interaction between antibody and protein. The AFM techniques have been successfully used in other antigen-IgG studies [27, 28] but these are very time-consuming procedures. The MCP-1 IgG complex has not been studied experimentally, yet. An alternative, promising in the nano-diagnostics, technique is friction (or lateral) force spectroscopy (FFS) [29, 30]. In contrast to the standard contact mode AFM, in FFS the probe quickly scans the surface laterally and the “unbinding” is enforced by the lateral forces. Usually the AFM tip is functionalized by the antibody and a protein is immobilized on a surface. The process of the enforced dissociation (molecular recognition) may depend on a relative orientation/motion of both proteins. Forces too weak do not generate useful signals.
In order to compare intermolecular interactions met in a classical AFM recognition experiment and in the lateral arrangement, we set-up a computer model of such an experiment. The impact of the direction of the AFM tip motion on mechanically enforced dissociation of antigen—antibody complex was computationally studied. Simulations of MCP-1 in a complex with the Fab fragment of immunoglobulin IgG antibody were performed using the Steered Molecular Dynamics method (SMD) [31–33]. This “virtual experiment” approach has been successful in studies of proteins’ nanomechanics [34–36]. Our numerous SMD ns scale trajectories revealed new atomic details of the molecular recognition phenomena in that medically important system. We show that the lateral dissociation requires substantially (20–40 %) lower forces than the vertical uncoupling of the complex but the later forces are still high enough to make FFS in our system feasible. This finding indicates that after careful calibration of friction based AFM methods a routine FFS nano-diagnostics involving MCP-1 chemokine will be possible.
Methods
As the first step we studied dynamics of a complex of human MCP-1 with 11 K2 Fab fragment [6] (Fig. 1) using the molecular dynamic (MD) method [37]. 11 K2 is a mouse monoclonal antibody against several human MCPs. The complex, extracted from the 2BDN entry in the Protein Data Bank was embedded in an 8 Å thick TIP3P water shell. After 0.4 ns equilibration we performed 10 ns of main MD simulation, using the NAMD 2.7 code [38] and the CHARMM27 force field [39].
A schematic view of MCP-1 (blue) and Fab fragment of Ig G antibody complex. In red light chains (L) are shown, in gray heavy chains (A) are depicted. V denotes variable region of IgG. Yellow arrows represent directions of virtual forces used in simulations. An orange arrow is an example of V direction and a green one represents L direction Note that direction of the z axis was used to define spherical coordinates of force vectors
Next, the SMD [31, 32] method was used in order to apply an external force which should dissociate the MCP-1-antibody complex in two perpendicular directions: the „vertical” force (V, almost parallel to the main axis of the antibody, the direction „z“ in Fig. 1) and the „lateral” one (L, approximately perpendicular to the main axis of the antibody). An increasing external virtual force was attached to all CA atoms of MCP-1 (chain A). During the simulations of stretching all CA atoms of the antibody (chains L and H) were fixed. The last structures obtained from the 3 ns standard MD simulations served as starting points for all SMD simulations. Structures were pulled for 2 ns at a constant speed of 0.025 Å/ps with a spring constant of 278 pN/Å. This value is close to that used in typical FFS experiments. Twenty two pulling directions were used. In addition we have studied a role of disulfide bridges on this molecular recognition process: two 2 ns simulations for each direction were generated for systems with all disulfide bridges converted to cysteines. Thus 3 × 9 V trajectories, and 3 × 13 L trajectories (Fig. 1) were further analyzed. Moreover, for one selected, representative V direction and one L direction ten additional trajectories (2 ns each) were generated in order to calculate values of an average dissociation force and to estimate statistical errors in the maximum force determination. Additionally, a dependence of the calculated forces on the pulling speed was tested. For ten directions, five vertical (V) and five lateral ones (L), we generated trajectories with a constant speed of 0.0025 Å/ps, i.e., ten times slower than before.
Electrostatic molecular potentials were calculated using the APBS method [40–43].
The analysis of results was performed using the VMD code [44] and homemade scripts.
Results and discussion
A classical MD
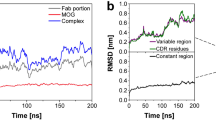
Since MCP-1 chemokine, despite its medical significance, has not been previously analyzed using classical MD modeling, we have studied dynamics of the complex on a 10 ns timescale. Except for the flexible terminal ends the chemokine has a rigid structure. Mean square atomic displacements of amino acids with respect to average positions (B-factor simulation) correlate rather well with the temperature B factors (Fig. 2).
A comparison of calculated mean square atomic displacements of MCP-1 cytokine amino acids with experimental temperature B-factors [6]
As expected, the N-terminal end (Ala4-Thr10) exhibit very large flexibility. This region is responsible for dimerization of MCP-1 cytokine [45]. One may notice that in the Cys12 – Ile32 fragment the model is more stable than X-ray measurements indicate. Probably in the computer model of an isolated complex the Arg18 residue is more strongly stabilized by the intramolecular electrostatic interactions than in a crystal setting. Both the simulations and the X-ray experiment indicate that the most stable region is Ala40 – Thr45 which corresponds to β2 internal β stand. Amino acids identified in the SMD simulations as being involved in molecular recognition process (Thr32, Glu39, Lys56, Gln61, Asp65) exhibit low fluctuations as well. This means that the MCP-1/IgG interface is quite well stabilized. The calculated B-factors for the Fab fragment are also in a very good agreement with the experimental data [6], Fig. 3.
A comparison of calculated mean square atomic displacements of heavy chain of Fab IgG antibody fragment with experimental temperature B-factors [6]
In Fig. 3 one can see that the most flexible part of the antibody heavy chain corresponds to a large loop in the region from Ala128(H) to Ser138(H). Interestingly, also in the antibody we observe a polar region which is more stable in the simulations than in a real crystal: Glu175-Ser176-Asp177. We explain this stabilization by a relaxation in an isolated model complex which leads to new salt bridges. Probably such relaxation is absent in the crystal due to packing interactions.
Steered molecular dynamics—mechanically enforced dissociation
All calculated force spectra of the enforced dissociation of MCP1-Fab complexes show qualitatively the same features—(i) a steep rise of the force up to a certain maximum value, (ii) a gradual decrease of the interaction force (iii) a separation phase characterized by a force close to 1 nN corresponding to the hydrodynamic drag. Representative curves for L and V dissociation modes are shown in Fig. 4. All calculated maxima of unbinding forces are collected in Table 1.
In reducing chemical conditions S-S protein bridges may break apart, ditiothreitol is commonly used for that purpose. Also some enzymes, such as thioredoxin or glutaredoxin facilitate transfer of electrons and make this redox reaction possible. Another source of the absence of stabilizing S-S covalent bonds may be just a point mutation in a cysteine position. Such perturbation may have significant impact on the molecular recognition process, particularly in CC chemokines. In order to check how such a structural modification may affect “in vitro” AFM experiments, we have repeated SMD simulation for models in which S-S bridges were disrupted both in MCP-1 chemokine and in the IgG antibody Fab fragment (7 bridges in total are absent now). Results for the reduced forms of the complex are presented in Table 1 as well (First, Second).
Thus, in total 69 output files were analyzed for the value of the force necessary to dissociate the complex.
In the standard forms (S-S bonds present), for 2 ns simulations, when the vertical (V) direction of the pulling force vector was applied, the lowest calculated force value was 1768 pN, while the highest value was 2911 pN. The average value of the maximum force observed during V direction SMD simulation was 2302 ± 366 pN. For laterally oriented pulling vectors the forces were lower: the highest value was 2229 pN, the lowest was 1391 pN, the average 1799 ± 301 pN. Respectively, for the simulations with ten times slower pulling speed the average for the vertical forces was 1437 ± 254 pN, and for the lateral force simulations the average was 962 ± 169 pN. Thus the process of mechanical unbinding requires lower forces if it proceeds in the direction L parallel to the MCP-1- antibody contact plane.
To study possible correlations between the value of the force necessary to separate antibody and MPC-1 and the dragging force direction we transformed force vectors into standard spherical coordinates (see Fig. 1). Forces with respect to values of the φ angle (only for “lateral” L cases) are given in Fig. 5a and forces dependence on the θ angle are shown in Fig. 5b.
Data presented in Fig. 5 clearly show that for the purely vertical separations the process requires the highest forces. In more lateral directions some lower force paths for dissociation may be easily found. The average later force is 22 % (2 ns simulations) and 40 % (for ten times slower pulling speed) lower than the vertical one. One should remember, that the SMD calculated forces, even with the low pulling speed, are a factor of 8–10 higher than those usually measured in AFM FS experiments. The reason is that, due to limited computer power, the loading rate used in SMD calculations has to be typically much higher than that in an experiment. However, calculated SMD forces correctly reproduce experimental trends: the lower the pulling speed—the lower the force. The simulations with ten times lower velocities confirm, that there is a difference in maximum unbinding forces for V and L arrangements of the dissociating components of the complex. Given all difficulties related to quantitative reproduction of force spectra from the virtual SMD experiments, we conclude that lateral motion of the functionalized AFM tip may require less force than a standard (V) contact mode molecular recognition study. These findings await for experimental confirmation, so far no AFM experiments for this system have been performed yet.
Molecular recognition and bioinformatics analysis of MCP-1
Detailed analysis of trajectories gave unique information on molecular interactions between the antibody and MCP-1 and their evolutions along alternative stretching paths. The flat interface region in a static crystal has been previously characterized [6]: it consists of the end of β1 strand, the β1–β2 loop, the beginning of β2 strand, the loop between β3 and α1, and the α1 helix. The formation of the complex buries about 15 % of the MCP-1 solvent accessible surface. Most of the contacts with MCP-1 are made through complementarity determining regions (CDR) H1 and H3 of the heavy chain of IgG and CDRs L1 and L3 of the light chain [6]. Important for recognition is Phe101H residue embedded in a mainly hydrophobic pocket of MCP-1 composed of Arg30A, Thr32A, Glu39A, Val41A, Pro55A and Met64A.
During stretching we observe that the complex is stabilized by strong salt bridges (in bold) and hydrogen bonds between: Glu39A - Arg98H, Lys56A - Asp52H, Asp65A - Arg32L, Asp68A - Arg32L, Thr32A - Glu55L, Gln61A - Tyr33H, where A denotes MCP-1, L - a light chain of IgG Fab fragment and H—the heavy chain of Fab. These interactions are shown in Fig. 6. Enlisted salt bridges/H-bonds were present in the majority of analyzed trajectories prior to an enforced dissociation both in V and L subsets. However, the scenario of consecutive breaking of these bonds obviously depends on the direction of the dragging force, particularly for L type vectors. Our trajectories provide data for interpretation of fine details induced by local events in future AFM force spectra.
An interface between MCP-1 and Fab IgG at the maximum force (V5) of SMD simulation. Amino acids involved in double interactions are shown in ball-and-stick representation. Thr32A-Glu55L pair is shown in orange, Glu39A- Arg98H pair is shown in green, Gln61A-Tyr33H pair is shown in pink. Standard coloring of atoms was applied for residues interacting with two amino acids from the partner protein. An important Phe101 residue is also shown in gray
Reduction of cysteines does not affect this list of stronger intermolecular interactions in MCP-1/Fab complex. Interestingly, additional H-bonds interacting pairs Asp68A-Ile31L, Lys56A-Trp32H were observed only for the complex with S-S bonds present.
Besides perfect matching of VdW surfaces and a strong hydrophobic handle created by Phe101, electrostatic interactions may also play a significant role for long range recognition in this system. We have used APBS program [40–43] to calculate a map of the molecular electrostatic potential (MEP) of MCP-1 and the Fab fragment. Rigid separated structures extracted from the 2BDN data were used for calculations. Results are presented in Fig. 7.
There are at least three regions (a, b, c, Fig. 7) with higher values of MEP. The regions in MCP-1 have corresponding counterparts in the Fab system of opposite charge. Thus electrostatics contribute to the stability of this complex as well. These calculations help to identify regions crucial for effective recognition of the important MCP-1 chemokine.
The fragment of MCP-1 specifically recognized by an antibody is called epitope. In order to check to what extent amino acids present at the interface in our cytokine are conserved in other proteins we performed PSI-BLAST search in standard non-redundant protein sequences database and used ClustalX2 code [46, 47] to make alignments of ten most similar sequences. Results visualized with Jalview 2.7 program [48, 49] are presented in Fig. 8.
For proteins in this set over 80 % similarity to MCP-1 is observed. Amino acids important for strong interactions identified by SMD simulations were analyzed in greater detail. Two distinct groups of polar epitope amino acids are present in MCP-1: conserved set (Thr32, Glu39, Gln61, Asp68) and a specific set: Lys56 and Asp65. In the conserved set the same amino acids are present in nearly all similar proteins. Residues from the specific set are characteristic only for MCP-1 chemokine. This finding corresponds well with the observed 160-fold decrease of IgG antibody affinity to a Lys56Asn MCP-1 mutant described in [6]. Thus Lys56 and Asp65 are particularly strongly involved in specific recognition of this chemokine by our IgG antibody. It is worth noting that this region is distinct from that responsible for an interaction of MCP-1 with its natural membrane receptor. Mutational studies have shown, that only a small subset of surface residues of human MCP-1 is important for effective interaction with CCR2 receptor: Tyr13, Arg24, Lys38, Lys49 [50]. None of these residues is involved in interaction with Fab fragment of IgG antibody studied here.
Conclusions
The presented computer modeling study was intended to check whether the lateral FFS AFM approach using antibody functionalized tips can be used in antigen-antibody molecular recognition studies and nano-diagnostics involving chemokines. Our results of over forty 2 ns and ten 10 ns long SMD simulations have shown that the interaction force between the Fab fragment of IgG antibody and MCP-1 is significant in lateral unbinding, thus such measurements should be possible. Computer experiments indicate that irrespective of a particular direction of the lateral unbinding forces, their values are systematically lower by about 20-40 % than the vertical ones. Ten times lower pulling speed data confirm qualitatively this conclusion. The lower forces require better sensitivity in the experiments than a standard, time consuming tapping modes, if FFS technique is to be used in the nano-diagnostics. This may be achieved by careful selection of high affinity antibodies against MCP-1 or a better design of the AFM hardware. The SMD force required to dissociate the complex studied here was even bigger than the force required to unfold the whole contactin 4 protein calculated with the same SMD protocol [51].
The strong intermolecular interactions arise from a set of 4–6 strong salt bridges and hydrogen bonds. MEP maps based of the CHARMM force field for MCP-1 and the antibody show high degree of electrical complementarity. Substantial hydrophobic interactions of Phe101 IgG residue, also noticed earlier [6], contribute to the molecular recognition process as well. Recent synthesis of human MCP-1 using a combination of solid phase peptide synthesis and native chemical ligation further enhances potential for medical applications of this interesting protein [52].
In summary, our modeling results indicate that the AFM FFS diagnostics using antibody functionalized tips for fast lateral scanning of a sample with immobilized cytokine may be a promising alternative to time consuming classical AFM measurements.
References
Paul WE (2008) Fundamental immunology. Kluwer/Lippincott Williams & Wilkins, Philadelphia
Weiner LM, Surana R, Wang S (2010) Monoclonal antibodies: versatile platforms for cancer immunotherapy. Nat Rev Immunol 10(5):317–327
Garay PA, McAllister AK (2010) Novel roles for immune molecules in neural development: implications for neurodevelopmental disorders. Front Synaptic Neurosci 2:136. doi:10.3389/fnsyn.2010.00136
Comerford I, McColl SR (2011) Mini-review series: focus on chemokines. Immunol Cell Biol 89(2):183–184
Deshmane SL, Kremlev S, Amini S, Sawaya BE (2009) Monocyte chemoattractant protein-1 (MCP-1): an overview. J Interferon Cytokine Res 29(6):313–326
Reid C, Rushe M et al (2006) Structure activity relationships of monocyte chemoattractant proteins in complex with a blocking antibody. Protein Eng Des Sel 19(7):317–324
Carr MW, Roth SJ, Luther E, Rose SS, Springer TA (1994) Monocyte chemoattractant protein 1 acts as a T-lymphocyte chemoattractant. Proc Natl Acad Sci U S A 91(9):3652–3656
Xu L, Warren M, Rose W, Gong W, Wang J (1996) Human recombinant monocyte chemotactic protein and other CC chemokines bind and induce directional migration of dendritic cells in vitro. J Leukoc Biol 60(3):365–371
Yoshimura T, Robinson EA, Tanaka S, Appella E, Kuratsu J-l, Leonard EJ (1989) Purification and amino acid analysis of two human glioma-derived monocyte chemoattractants. J Exp Med 169(4):1449–1459
Van Coillie E, Van Damme J, Opdenakker G (1999) The MCP/eotaxin subfamily of CC chemokines. Cytokine Growth Factor Rev 10(1):61–86
Banisadr G, Gosselin RD, Mechighel P, Kitabgi P, Rostne W, Parsadaniantz SM (2005) Highly regionalized neuronal expression of monocyte chemoattractant protein 1 (MCP 1/CCL2) in rat brain: evidence for its colocalization with neurotransmitters and neuropeptides. J Comp Neurol 489(3):275–292
De Haas A, van Weering HRJ, De Jong E, Boddeke HWGM, Biber KPH (2007) Neuronal chemokines: versatile messengers in central nervous system cell interaction. Mol Neurobiol 36(2):137–151
Panee J (2012) Monocyte chemoattractant protein 1 (MCP-1) in obesity and diabetes. Cytokine 60(1):1–12
Salcedo R, Ponce ML et al (2000) Human endothelial cells express CCR2 and respond to MCP-1: direct role of MCP-1 in angiogenesis and tumor progression. Blood 96(1):34–40
Zhang J, Patel L, Pienta KJ (2010) CC chemokine ligand 2 (CCL2) promotes prostate cancer tumorigenesis and metastasis. Cytokine Growth Factor Rev 21(1):41–48
Sandhu SK, Papadopoulos K et al (2013) A first-in-human, first-in-class, phase I study of carlumab (CNTO 888), a human monoclonal antibody against CC-chemokine ligand 2 in patients with solid tumors. Cancer Chemother Pharmacol 71(4):1041–1050
Vargas D, Nascimbene C, Krishnan C, Zimmerman A, Pardo C (2005) Neuroglial activation and neuroinflammation in the brain of patients with autism. Ann Neurol 57(1):67–81
Ashwood P, Krakowiak P, Hertz-Picciotto I, Hansen R, Pessah IN, Van de Water J (2011) Associations of impaired behaviors with elevated plasma chemokines in autism spectrum disorders. J Neuroimmunol 232(1–2):196–199
Pardo CA, Eberhart CG (2007) The neurobiology of autism. Brain Pathol 17(4):434–447
Parot P, Dufrêne YF, Hinterdorfer P, Le Grimellec C, Navajas D, Pellequer JL, Scheuring S (2007) Past, present and future of atomic force microscopy in life sciences and medicine. J Mol Recognit 20(6):418–431
Francis LW, Lewis PD, Wright CJ, Conlan RS (2010) Atomic force microscopy comes of age. Biol Cell 102(2):133–143
Hinterdorfer P, Baumgartner W, Gruber HJ, Schilcher K, Schindler H (1996) Detection and localization of individual antibody-antigen recognition events by atomic force microscopy. Proc Natl Acad Sci U S A 93(8):3477
Ros R, Schwesinger F, Anselmetti D, Kubon M, Schäfer R, Plückthun A, Tiefenauer L (1998) Antigen binding forces of individually addressed single-chain Fv antibody molecules. Proc Natl Acad Sci U S A 95(13):7402
Allen S, Chen X et al (1997) Detection of antigen-antibody binding events with the atomic force microscope. Biochemistry 36(24):7457–7463
Li G, Xi N, Wang DH (2006) Probing membrane proteins using atomic force microscopy. J Cell Biochem 97(6):1191–1197
Dufrêne YF, Hinterdorfer P (2008) Recent progress in AFM molecular recognition studies. Pflügers Archiv Eur J Physiol 456(1):237–245
Lv Z, Wang J, Chen G, Deng L (2010) Imaging recognition events between human IgG and rat anti-human IgG by atomic force microscopy. Int J Biol Macromol 47(5):661–667
Lv Z, Wang J, Deng L, Chen G (2009) Preparation and characterization of covalently binding of rat anti-human IgG monolayer on thiol-modified gold surface. Nanoscale Res Lett 4(12):1403–1408
Lekka M, Kulik AJ, Jeney S, Raczkowska J, Lekki J, Budkowski A, Forró L (2005) Friction force microscopy as an alternative method to probe molecular interactions. J Chem Phys 123:014702
Chirasatitsin S, Engler AJ (2010) Detecting cell-adhesive sites in extracellular matrix using force spectroscopy mapping. J Phys: Condens Matter 22:194102
Grubmüller H, Heymann B, Tavan P (1996) Ligand binding: molecular mechanics calculation of the streptavidin-biotin rupture force. Science 271(5251):997–999
Marszalek PE, Lu H, Li H, Carrion-Vazquez M, Oberhauser AF, Schulten K, Fernandez JM (1999) Mechanical unfolding intermediates in titin modules. Nature 402(6757):100–103
Hanasaki I, Haga T, Kawano S (2008) The antigen–antibody unbinding process through steered molecular dynamics of a complex of an Fv fragment and lysozyme. J Phys: Condens Matter 20:255238
Mikulska K, Pepłowski Ł, Nowak W (2012) Nanomechanics of Ig-like domains of human contactin (BIG-2). J Mol Model 17:2313–2323
Pepłowski Ł, Sikora M, Nowak W, Cieplak M (2011) Molecular jamming—The cystine slipknot mechanical clamp in all-atom simulations. J Chem Phys 134:085102
Mikulska K, Strzelecki J, Balter A, Nowak W (2012) Nanomechanical unfolding of α-neurexin-a major component of the synaptic junction. Chem Phys Lett 512:134–137
Berendsen HJC, Postma JPM, van Gunsteren WF, DiNola A, Haak JR (1984) Molecular dynamics with coupling to an external bath. J Chem Phys 81(8):3684–3690
Phillips JC, Braun R et al (2005) Scalable molecular dynamics with NAMD. J Comput Chem 26(16):1781–1802
Brooks BR, Brooks C III et al (2009) CHARMM: the biomolecular simulation program. J Comput Chem 30(10):1545–1614
Baker NA, Sept D, Joseph S, Holst MJ, McCammon JA (2001) Electrostatics of nanosystems: application to microtubules and the ribosome. Proc Natl Acad Sci U S A 98(18):10037
Dolinsky TJ, Czodrowski P, Li H, Nielsen JE, Jensen JH, Klebe G, Baker NA (2007) PDB2PQR: expanding and upgrading automated preparation of biomolecular structures for molecular simulations. Nucleic Acids Res 35(suppl 2):W522–W525
Dolinsky TJ, Nielsen JE, McCammon JA, Baker NA (2004) PDB2PQR: an automated pipeline for the setup of Poisson–Boltzmann electrostatics calculations. Nucleic Acids Res 32(suppl 2):W665–W667
Holst M, Saied F (1993) Multigrid solution of the Poisson—Boltzmann equation. J Comput Chem 14(1):105–113
Humphrey W, Dalke A, Schulten K (1996) VMD: visual molecular dynamics. J Mol Graph 14(1):33–38
Lubkowski J, Bujacz G, Boqué L (1997) The structure of MCP-1 in two crystal forms provides a rare example of variable quaternary interactions. Nat Struct Mol Biol 4(1):64–69
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25(17):3389–3402
Larkin M, Blackshields G et al (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23(21):2947–2948
Waterhouse AM, Procter JB, Martin DMA, Clamp M, Barton GJ (2009) Jalview Version 2—a multiple sequence alignment editor and analysis workbench. Bioinformatics 25(9):1189–1191
Clamp M, Cuff J, Searle SM, Barton GJ (2004) The jalview java alignment editor. Bioinformatics 20(3):426–427
Hemmerich S, Paavola C et al (1999) Identification of residues in the monocyte chemotactic protein-1 that contact the MCP-1 receptor, CCR2. Biochemistry 38(40):13013–13025
Strzelecki J, Mikulska K, Lekka M, Kulik A, Balter A, Nowak W (2009) AFM force spectroscopy and steered molecular dynamics simulation of protein contactin 4. Acta Phys Pol, A 116:156–159
Grygiel TLR, Teplyakov A et al (2010) Synthesis by native chemical ligation and crystal structure of human CCL2. Pept Sci 94(3):350–359
Acknowledgments
This work was supported by Polish Funds for Science – grant no. N519 578138 and grant no. N202 262038.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Gogolinska, A., Nowak, W. Molecular basis of lateral force spectroscopy nano-diagnostics: computational unbinding of autism related chemokine MCP-1 from IgG antibody. J Mol Model 19, 4773–4780 (2013). https://doi.org/10.1007/s00894-013-1972-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00894-013-1972-z