Abstract
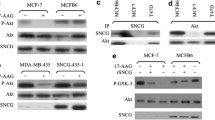
Aberrantly expressed human gamma synuclein (SNCG) interacts with BubR1 and heat shock protein 70 (Hsp70) in late stages of breast and ovarian cancer. This interaction is essential for progression, development and survival of cancer cells. A short, synthetically designed ankyrin-repeat-containing peptide (ANK peptide) was proven to inhibit the activity of SNCG. However, the potential binding site residues of SNCG responsible for its oncogenic function have not been reported so far. The objectives of this study were to generate a three-dimensional model of SNCG and to identify the key residues involved in interaction with BubR1, ANK peptide and Hsp70. Our study is the first attempt to report the specific binding of SNCG with the TPR motif of BubR1 and the 18kDa region of Hsp70. Our findings provide novel insights into the mechanism of interaction of SNCG, and can act as a basis for the ongoing drug design and discovery process aimed at treating breast and ovarian cancer.








Similar content being viewed by others
References
Bruening W, Giasson BI, Klein-Szanto AJ, Lee VM, Trojanowski JQ, Godwin AK (2000) Synucleins are expressed in the majority of breast and ovarian carcinomas and in preneoplastic lesions of the ovary. Cancer 88(9):2154–2163
Lavedan C, Leroy E, Dehejia A, Buchholtz S, Dutra A, Nussbaum RL, Polymeropoulos MH (1998) Identification, localization and characterization of the human gamma-synuclein gene. Hum Genet 103(1):106–112
George JM (2002) The synucleins. Genome Biol 3(1):REVIEWS3002
Lavedan C (1998) The synuclein family. Genome Res 8(9):871–880
Gupta A, Godwin AK, Vanderveer L, Lu A, Liu J (2003) Hypomethylation of the synuclein gamma gene CpG island promotes its aberrant expression in breast carcinoma and ovarian carcinoma. Cancer Res 63(3):664–673
Pan ZZ, Bruening W, Giasson BI, Lee VM, Godwin AK (2002) Gamma-synuclein promotes cancer cell survival and inhibits stress-and chemotherapy drug-induced apoptosis by modulating MAPK pathways. J Biol Chem 277(38):35050–35060
Pan ZZ, Bruening W, Godwin AK (2006) Involvement of RHO GTPases and ERK in synuclein-gamma enhanced cancer cell motility. Int J Oncol 29(5):1201–1205
Davenport J, Harris LD, Goorha R (2006) Spindle checkpoint function requires Mad2-dependent Cdc20 binding to the Mad3 homology domain of BubR1. Exp Cell Res 312(10):1831–1842
Gupta A, Inaba S, Wong OK, Fang G, Liu J (2003) Breast cancer-specific gene 1 interacts with the mitotic checkpoint kinase BubR1. Oncogene 22(48):7593–7599
Hardwick KG, Johnston RC, Smith DL, Murray AW (2000) MAD3 encodes a novel component of the spindle checkpoint which interacts with Bub3p, Cdc20p, and Mad2p. J Cell Biol 148(5):871–882
Zhou Y, Inaba S, Liu J (2006) Inhibition of synuclein-gamma expression increases the sensitivity of breast cancer cells to paclitaxel treatment. Int J Oncol 29(1):289–295
Singh VK, Zhou Y, Marsh JA, Uversky VN, Forman-Kay JD, Liu J, Jia Z (2007) Synuclein-gamma targeting peptide inhibitor that enhances sensitivity of breast cancer cells to antimicrotubule drugs. Cancer Res 67(2):626–633
Jiang Y, Liu YE, Goldberg ID, Shi YE (2004) Gamma synuclein, a novel heat-shock protein-associated chaperone, stimulates ligand-dependent estrogen receptor alpha signaling and mammary tumorigenesis. Cancer Res 64(13):4539–4546
Jiang Y, Liu YE, Lu A, Gupta A, Goldberg ID, Liu J, Shi YE (2003) Stimulation of estrogen receptor signaling by gamma synuclein. Cancer Res 63(14):3899–3903
Souza JM, Giasson BI, Lee VM, Ischiropoulos H (2000) Chaperone-like activity of synucleins. FEBS Lett 474(1):116–119
Roodveldt C, Bertoncini CW, Andersson A, van der Goot AT, Hsu ST, Fernandez-Montesinos R, de Jong J, van Ham TJ, Nollen EA, Pozo D, Christodoulou J, Dobson CM (2009) Chaperone proteostasis in Parkinson’s disease: stabilization of the Hsp70/alpha-synuclein complex by Hip. EMBO J 28(23):3758–3770
Chappell TG, Konforti BB, Schmid SL, Rothman JE (1987) The ATPase core of a clathrin uncoating protein. J Biol Chem 262(2):746–751
Cyr DM, Langer T, Douglas MG (1994) DnaJ-like proteins: molecular chaperones and specific regulators of Hsp70. Trends Biochem Sci 19(4):176–181
McGinnis S, Madden TL (2004) BLAST: at the core of a powerful and diverse set of sequence analysis tools. Nucleic Acids Res 32(Web Server issue):W20–W25
Zhang Y (2008) I-TASSER server for protein 3D structure prediction. BMC Bioinform 9:40
Corpet F (1988) Multiple sequence alignment with hierarchical clustering. Nucleic Acids Res 16(22):10881–10890
Sali A, Blundell TL (1993) Comparative protein modelling by satisfaction of spatial restraints. J Mol Biol 234(3):779–815
Vriend G (1990) WHAT IF: a molecular modeling and drug design program. J Mol Graph 8(1):52–56, 29
Lovell SC, Davis IW, Arendall WB 3rd, de Bakker PI, Word JM, Prisant MG, Richardson JS, Richardson DC (2003) Structure validation by Calpha geometry: phi, psi and Cbeta deviation. Proteins 50(3):437–450
Colovos C, Yeates TO (1993) Verification of protein structures: patterns of non bonded atomic interactions. Protein Sci 2(9):1511–1519
Liang J, Edelsbrunner H, Woodward C (1998) Anatomy of protein pockets and cavities: measurement of binding site geometry and implications for ligand design. Protein Sci 7(9):1884–1897
Lee B, Richards FM (1971) The interpretation of protein structures: estimation of static accessibility. J Mol Biol 55(3):379–400
Rocchia W, Sridharan S, Nicholls A, Alexov E, Chiabrera A, Honig B (2002) Rapid grid-based construction of the molecular surface and the use of induced surface charge to calculate reaction field energies: applications to the molecular systems and geometric objects. J Comput Chem 23(1):128–137
Dominguez C, Boelens R, Bonvin AM (2003) HADDOCK: a protein-protein docking approach based on biochemical or biophysical information. J Am Chem Soc 125(7):1731–1737
Huey R, Morris GM, Olson AJ, Goodsell DS (2007) A semiempirical free energy force field with charge-based desolvation. J Comput Chem 28(6):1145–1152
Shen MY, Sali A (2006) Statistical potential for assessment and prediction of protein structures. Protein Sci 15(11):2507–2524
Van der Spoel D, Lindahl E, Hess B, Groenhof G, Mark AE, Berendsen HJ (2005) GROMACS: fast, flexible, and free. J Comput Chem 26(16):1701–1718
Ulmer TS, Bax A, Cole NB, Nussbaum RL (2005) Structure and dynamics of micelle-bound human alpha-synuclein. J Biol Chem 280(10):9595–9603
Bolanos-Garcia VM, Kiyomitsu T, D’Arcy S, Chirgadze DY, Grossmann JG, Matak-Vinkovic D, Venkitaraman AR, Yanagida M, Robinson CV, Blundell TL (2009) The crystal structure of the N-terminal region of BUB1 provides insight into the mechanism of BUB1 recruitment to kinetochores. Structure 17(1):105–116
Kohl A, Binz HK, Forrer P, Stumpp MT, Pluckthun A, Grutter MG (2003) Designed to be stable: crystal structure of a consensus ankyrin repeat protein. Proc Natl Acad Sci USA 100(4):1700–1705
Morshauser RC, Hu W, Wang H, Pang Y, Flynn GC, Zuiderweg ER (1999) High-resolution solution structure of the 18 kDa substrate-binding domain of the mammalian chaperone protein Hsc70. J Mol Biol 289(5):1387–1403
Thompson JD, Gibson TJ, Higgins DG (2003) Multiple sequence alignment using ClustalW and ClustalX. In: Baxevanis AD (ed) Current Protocols in Bioinformatics. Wiley, San Francisco, pp 2.3.1–2.3.22
DeLano WL (2002) The PyMOL Molecular Graphics System. DeLano Scientific LLC, San Carlos, CA. http://www.pymol.org
Krishna M, John K, Helen K, Bryan F (1994) XYPLOT-Data Analysis Program, Creative Consulting for Research & Education. http://ccreweb.org/software/xyplot/xyplot.html
Acknowledgments
Research in the Laboratory of Structure Biology is supported by grants from University Grants Commission (UGC), Government of India, New Delhi, India. R.K. thanks the Centre for Bioinformatics, Pondicherry University, India.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Manivel, P., Muthukumaran, J., Kannan, M. et al. Insight into residues involved in the structure and function of the breast cancer associated protein human gamma synuclein. J Mol Model 17, 251–263 (2011). https://doi.org/10.1007/s00894-010-0718-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00894-010-0718-4