Abstract
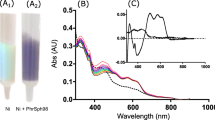
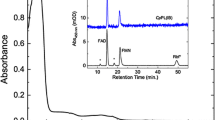
Photolyases are proteins that enzymatically repair the UV-induced DNA damage by a protein–DNA electron transfer mechanism. They repair either cyclobutane pyrimidine dimers or pyrimidine (6-4) pyrimidone photoproducts or just (6-4)-photoproducts. In this work, we report the production and partial characterization of a recombinant (6-4)-photolyase (SphPhrB97) from a bacterial psychrotolerant Antarctic isolate identified as Sphingomonas sp. strain UV9. The spectrum analysis and the in silico study of SphPhrB97 suggest that this enzyme has similar features as compared to the (6-4)-photolyase from Agrobacterium tumefaciens (4DJA; PhrB), including the presence of three cofactors: FAD, DMRL (6,7-dimethyl-8-(1′-D-ribityl) lumazine), and an Fe–S cluster. The homology model of SphPhrB97 predicts that the DNA-binding pocket (area and volume) is larger as compared to (6-4)-photolyases from mesophilic microbes. Based on sequence comparison and on the homology model, we propose an electron transfer pathway towards the FAD cofactor involving the residues Trp342, Trp390, Tyr40, Tyr391, and Tyr399. The phylogenetic tree performed using curated and well-characterized prokaryotic (6-4)-photolyases suggests that SphPhrB97 may have an ancient evolutionary origin. The results suggest that SphPhrB97 is a cold-adapted enzyme, ready to cope with the UV irradiation stress found in a hostile environment, such as Antarctica.






Similar content being viewed by others
References
Albarracín VH, Simon J, Pathak GP et al (2014) First characterisation of a CPD-class i photolyase from a UV-resistant extremophile isolated from high-altitude Andean lakes. Photochem Photobiol Sci. https://doi.org/10.1039/c3pp50399b
An M, Zheng Z, Qu C et al (2018) The first (6–4) photolyase with DNA damage repair activity from the Antarctic microalga Chlamydomonas sp. ICE-L. Mutat Res Fundam Mol Mech Mutagen. https://doi.org/10.1016/j.mrfmmm.2018.03.004
Brzóska K, Mȩczyńska S, Kruszewski M (2006) Iron−sulfur cluster proteins: electron transfer and beyond. Acta Biochim Pol 53(4):685–691
D’Amico S, Claverie P, Collins T, Georlette D, Gratia E, Hoyoux A, Gerday C (2002) Molecular basis of cold adaptation. Philos Trans R Soc Lond B Biol Sci 357(1423):917–925. https://doi.org/10.1098/rstb.2002.1105
Daniel RM, Danson MJ (1995) Did primitive microorganisms use nonhem iron proteins in place of NAD/P? J Mol Evol. https://doi.org/10.1007/BF00160501
Di Tommaso P, Moretti S, Xenarios I et al (2011) T-coffee: a web server for the multiple sequence alignment of protein and RNA sequences using structural information and homology extension. Nucleic Acids Res. https://doi.org/10.1093/nar/gkr245
Dikbas UM, Tardu M, Canturk A et al (2019) Identification and characterization of a new class of (6-4) photolyase from Vibrio cholerae. Biochemistry. https://doi.org/10.1021/acs.biochem.9b00766
Eker APM, Yajima H, Yasui A (1994) DNA photolyase from the fungus Neurospora crassa. Purification, characterization and comparison with other photolyases. Photochem Photobiol 60:125–133. https://doi.org/10.1111/j.1751-1097.1994.tb05078.x
Fortunato AE, Annunziata R, Jaubert M, Bouly JP, Falciatore A (2015) Dealing with light: the widespread and multitasking cryptochrome/photolyase family in photosynthetic organisms. J Plant Physiol 172:42–54
Fujihashi M, Numoto N, Kobayashi Y et al (2007) Crystal structure of archaeal photolyase from sulfolobus tokodaii with two FAD molecules: implication of a novel light-harvesting cofactor. J Mol Biol. https://doi.org/10.1016/j.jmb.2006.10.012
Fullana N, Braña V, José Marizcurrena J et al (2017) Identification, recombinant production and partial biochemical characterization of an extracellular cold-active serine-metalloprotease from an Antarctic Pseudomonas isolate. AIMS Bioeng 4:386–401. https://doi.org/10.3934/bioeng.2017.3.386
Graf D, Wesslowski J, Ma H et al (2015) Key amino acids in the bacterial (6-4) photolyase PhrB from Agrobacterium fabrum. PLoS ONE. https://doi.org/10.1371/journal.pone.0140955
Haase I, Fischer M, Bacher A, Schramek N (2003) Temperature-dependent presteady state kinetics of lumazine synthase from the hyperthermophilic eubacterium Aquifex aeolicus. J Biol Chem. https://doi.org/10.1074/jbc.M303090200
Kavakli IH, Ozturk N, Gul S (2019) DNA repair by photolyases. In: Advances in protein chemistry and structural biology, vol. 115. Academic Press Inc., pp 1–19
Klar T, Kaiser G, Hennecke U et al (2006) Natural and non-natural antenna chromophores in the DNA photolyase from Thermus Thermophilus. ChemBioChem. https://doi.org/10.1002/cbic.200600206
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol. https://doi.org/10.1093/molbev/msw054
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. https://doi.org/10.1038/227680a0
Lamparter T, Zhang F, Graf D, Wesslowski J, Oberpichler I, Schünemann V, Scheerer P (2011) A Prokaryotic (6-4) photolyase with a DMRL chromophore and an iron–sulfur cluster. In: Encyclopedia of inorganic and bioinorganic chemistry pp 1–13
Letunic I, Bork P (2016) Interactive tree of life (iTOL) v3: an online tool for the display and annotation of phylogenetic and other trees. Nucleic Acids Res 44(W1):W242–W245
López-Maury L, Marguerat S, Bähler J (2008) Tuning gene expression to changing environments: from rapid responses to evolutionary adaptation. Nature Rev Genet 9(8):583–593
Ma H, Holub D, Gillet N et al (2019) Two aspartate residues close to the lesion binding site of Agrobacterium (6-4) photolyase are required for Mg2+ stimulation of DNA repair. FEBS J. https://doi.org/10.1111/febs.14770
Marizcurrena JJ, Morel MA, Braña V et al (2017) Searching for novel photolyases in UVC-resistant Antarctic bacteria. Extremophiles. https://doi.org/10.1007/s00792-016-0914-y
Marizcurrena JJ, Martínez-López W, Ma H et al (2019a) A highly efficient and cost-effective recombinant production of a bacterial photolyase from the Antarctic isolate Hymenobacter sp. UV11. Extremophiles 23:49–57. https://doi.org/10.1007/s00792-018-1059-y
Marizcurrena JJ, Morales D, Smircich P, Castro-Sowinski S (2019b) Draft genome sequence of the UV-resistant Antarctic bacterium Sphingomonas sp. Strain UV9. Microbiol Resour Announc. https://doi.org/10.1128/mra.01651-18
Marizcurrena JJ, Acosta S, Canclini L et al (2020) A natural occurring bifunctional CPD/(6–4)-photolyase from the Antarctic bacterium Sphingomonas sp. UV9. Appl Microbiol Biotechnol. https://doi.org/10.1007/s00253-020-10734-5
Munshi S, Rajamoorthi A, Stanley RJ (2017) Characterization of a cold-adapted DNA photolyase from C. psychrerythraea 34H. Extremophiles. https://doi.org/10.1007/s00792-017-0953-z
Notredame C, Higgins DG, Heringa J (2000) T-coffee: A novel method for fast and accurate multiple sequence alignment. J Mol Biol. https://doi.org/10.1006/jmbi.2000.4042
Oberpichler I, Pierik AJ, Wesslowski J et al (2011) A photolyase-like protein from Agrobacterium tumefaciens with an iron-sulfur cluster. PLoS ONE. https://doi.org/10.1371/journal.pone.0026775
Ozturk N (2017) Phylogenetic and functional classification of the photolyase/cryptochrome family. Photochem Photobiol 93(1):104–111
Portero LR, Alonso-Reyes DG, Zannier F et al (2019) Photolyases and Cryptochromes in UV-resistant bacteria from high-altitude Andean lakes. Photochem Photobiol. https://doi.org/10.1111/php.13061
Puig S, Granger C, Garre A, Trullàs C, Sanmartin O, Argenziano G (2019) Review of clinical evidence over 10 years on prevention and treatment of a film-forming medical device containing photolyase in the management of field cancerization in actinic keratosis. Dermatol Ther 9(2):259–270
Robert X, Gouet P (2014) Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res. https://doi.org/10.1093/nar/gku316
Saxowsky TT, Doetsch PW (2006) RNA polymerase encounters with DNA damage: transcription-coupled repair or transcriptional mutagenesis?. Chem Rev 106(2):474–488
Schrödinger LLC (2010) The PyMOL molecular graphics system, Version 1(5), 0
Sievers F, Higgins DG (2018) Clustal omega for making accurate alignments of many protein sequences. Protein Sci 27:135–145. https://doi.org/10.1002/pro.3290
Steffen C, Thomas K, Huniar U et al (2010) AutoDock4 and autodocktools4: automated docking with selective receptor flexibility. J Comput Chem. https://doi.org/10.1002/jcc
Studier FW (2005) Protein production by auto-induction in high-density shaking cultures. Protein Expr Purif. https://doi.org/10.1016/j.pep.2005.01.016
Tian W, Chen C, Lei X et al (2018) CASTp 3.0: computed atlas of surface topography of proteins. Nucleic Acids Res. https://doi.org/10.1093/nar/gky473
von Zadow A, Ignatz E, Pokorny R et al (2016) Rhodobacter sphaeroides CryB is a bacterial cryptochrome with (6-4) photolyase activity. FEBS J. https://doi.org/10.1111/febs.13924
Wang X, Shi C, Chen G et al (2019) Characterization of recombinant glutathione reductase from Antarctic yeast Rhodotorula mucilaginosa. Polar Biol. https://doi.org/10.1007/s00300-019-02603-3
Wilson TJ, Crystal MA, Rohrbaugh MC et al (2011) Evidence from thermodynamics that DNA photolyase recognizes a solvent-exposed CPD lesion. J Phys Chem B. https://doi.org/10.1021/jp208129a
Yarosh DB, Rosenthal A, Moy R (2019) Six critical questions for DNA repair enzymes in skincare products: a review in dialog. Clin Cosmet Investig Dermatol 12:617
Zhang F, Scheerer P, Oberpichler I et al (2013) Crystal structure of a prokaryotic (6-4) photolyase with an Fe-S cluster and a 6,7-dimethyl-8-ribityllumazine antenna chromophore. Proc Natl Acad Sci USA. https://doi.org/10.1073/pnas.1302377110
Zhang F, Ma H, Bowatte K, Kwiatkowski D, Mittmann E, Qasem H, Yang X (2017) Crystal structures of bacterial (6-4) photolyase mutants with impaired DNA repair activity. Photochem Photobiol 93(1):304–314
Zhong D (2015) Electron transfer mechanisms of DNA repair by photolyase. Annu Rev Phys Chem. https://doi.org/10.1146/annurev-physchem-040513-103631
Zimmermann L, Stephens A, Nam SZ et al (2018) A completely reimplemented MPI bioinformatics toolkit with a new hhpred server at its core. J Mol Biol. https://doi.org/10.1016/j.jmb.2017.12.007
Acknowledgements
The authors thank the Uruguayan Antarctic Institute for the logistic support during the stay in the Antarctic Base Artigas. S. Castro-Sowinski and J. J. Marizcurrena are members of the National Research System (SNI, Sistema Nacional de Investigadores). This work was partially supported by PEDECIBA (Programa de Desarrollo de las Ciencias Básicas), CSIC (Project C667), and ANII (Project FMV_3_2016_1_1226654). The work of JJM was supported by ANII and CAP (Comisión Académica de Posgrado, UdelaR).
Author information
Authors and Affiliations
Contributions
JJM and SCS conceived and designed research. JJM conducted experiments and analyzed data. TL conducted phylogenetic analysis. JJM, TL, and SCS wrote the manuscript. All the authors read and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest with the contents of this article.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Communicated by A. Driessen.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Marizcurrena, J.J., Lamparter, T. & Castro-Sowinski, S. A (6-4)-photolyase from the Antarctic bacterium Sphingomonas sp. UV9: recombinant production and in silico features. Extremophiles 24, 887–896 (2020). https://doi.org/10.1007/s00792-020-01202-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00792-020-01202-z