Abstract
Self-splicing inteins are mobile genetic elements invading host genes via nested homing endonuclease (HEN) domains. All HEN domains residing within inteins are inserted at a highly conserved insertion site. A purifying selection mechanism directing the location of the HEN insertion site has not yet been identified. In this work, we solved the three-dimensional crystal structures of two inteins inserted in the cell division control protein 21 of the hyperthermophilic archaea Pyrococcus abyssi and Pyrococcus horikoshii. A comparison between the structures provides the structural basis for the thermo-stabilization mechanism of inteins that have lost the HEN domain during evolution. The presence of an entire extein domain in the intein structure from Pyrococcus horikoshii suggests the selection mechanism for the highly conserved HEN insertion point.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Protein splicing is catalyzed by intervening protein sequences termed inteins (Paulus 2000; Perler et al. 1994). Self-splicing inteins can be divided into three different groups based on their primary structures (Paulus 2000; Gogarten et al. 2002). The first group encompasses canonical inteins bearing a homing endonuclease (HEN) domain that can be one of the various types of HEN domains, including the most common LAGLIDADG family and is either active or degenerated (Barzel et al. 2011). Mini-inteins lacking a HEN domain constitute the second group (Novikova et al. 2016). Naturally occurring mini-inteins generally seem proficient in catalyzing protein splicing, suggesting that protein-splicing HINT (Hedgehog/INTein) and HEN domains are functionally independent of each other. A few mini-inteins are naturally split into two fragments and often regarded as a distinct third group due to their bi-molecular splicing reaction (Wu et al. 1998; Iwai et al. 2006; Dassa et al. 2009). Split inteins catalyze protein splicing in trans, thereby ligating two flanking polypeptide chains into one chain and have been used for various synthetic biological and biotechnological applications (Wood and Camarero 2014; Volkmann and Iwaï 2010).
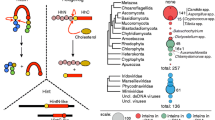
Inteins are particularly prevalent in archaea as they were found in nearly half of all sequenced archaeal genomes, whereas only one percent of eukaryotic genomes contain inteins; these are further limited to unicellular organisms (Novikova et al. 2016). It is believed that inteins have spread across taxa via horizontal gene transfer (HGT) facilitated by the nested HEN domains. Once intein genes are fixed in an organism, nested HEN domains in inteins are constantly facing degeneration due to the lack of activity-driven selection (Gogarten et al. 2002; Gogarten and Hilario 2006). Thus, one might assume that mini-inteins have lost their HEN domains during evolution. Some inteins have developed a mutualism between HINT and HEN domains, thereby avoiding the complete loss of the HEN domain (Iwaï et al. 2017). Comparison of structure and function among various inteins could shed light on how each intein has spread and evolved. In the genomes of Pyrococcus abyssi and Pyrococcus horikoshii OT3, 14 inteins have been reported (Pietrokovski 2001). It remains elusive why some archaeal genomes have accumulated dozens of intein insertions. The investigation of the intein profile of such organisms could provide new insights into the relationship between different inteins and their host organisms and unveil details on the history of horizontal gene transfer events.
As the first step toward understanding the spread and evolution of inteins and cross-talks within the “inteinome”, we determined the crystal structures of mini-inteins inserted at the same insertion site within the putative cell division control protein 21 (Cdc21) from the hyperthermophilic archaea, Pyrococcus abyssi and Pyrococcus horikoshii OT3.
Results
The cell division control protein 21 (Cdc21), belonging to the MCM (mini-chromosome maintenance) protein family, is one of the proteins invaded by several inteins (Pietrokovski 2001). Particularly hyperthermophiles such as Pyrococcus abyssi and Pyrococcus horikoshii contain multiple inteins inserted into the same Cdc21 protein (Pietrokovski 2001). While some thermophilic inteins like the TFIIB intein from Methanocaldococcus jannaschii (MjaTFIIB intein) require the HEN domain or a long linker at the HEN insertion site for protein-splicing activity, mini-inteins from Pyrococcus are capable of protein splicing even at 37 °C (Fig. S1) (Iwaï et al. 2017; Ellilä et al. 2011). To understand the structural basis of the difference between inteins from hyperthermophiles, we investigated three-dimensional structures of mini-inteins from the Pyrococcus genus.
Structure of PabCDC21-1 intein
The hypothetical Cdc21 protein from Pyrococcus abyssi (Uniprot: Q9UYR7) harbors the two inteins CDC21-1 (PabCDC21-1) and CDC21-2 (PabCDC21-2) consisting of 164 and 268 residues, respectively. Both are mini-inteins lacking HEN domains. The PabCDC21-1 intein is inserted at the presumable P-loop ATP/GTP-binding motif; while, the PabCDC21-2 intein locates 27 residues downstream of the PabCDC21-1 intein insertion site. According to our experience, inteins with shorter loops at the HEN insertion site tend to crystallize and diffract better than inteins with longer loops (Oeemig et al. 2012). Indeed, we could obtain diffracting crystals of a variant of the PabCDC21-1 intein carrying mutations at the N- and C-terminal catalytic residues (C1A, and N164A) together with three N- and one C-extein residue (Ser-Ala-Lys and Ala, respectively). We solved the three-dimensional structure of PabCDC21-1 intein at 1.60-Å resolution by molecular replacement (Fig. 1). The structure of the PabCDC21-1 intein revealed the HINT fold characteristic for inteins from thermophilic organisms (Aranko et al. 2014a). In contrast to most mesophilic inteins, thermophilic inteins often contain an additional insertion of two β-strands connected with a helix extension between Blocks A and B, approximately 25 residues distant from the N-terminus (Fig. 1a–c) (Aranko et al. 2014a). Because this additional sequence insertion is commonly found among inteins from thermophiles, one could speculate about a possible role in structure stabilization at elevated temperatures, or a functional role specific for thermophilic inteins. The presence of N- and C-extein residues due to the active site mutations allowed us to analyze the conformation of the PabCDC21-1 intein by measuring the distance between the N-scissile peptide and the side chain of the +1 residue (Cα atom of Ala). A large distance of about 8 Å suggests an open conformation as it has been observed in most intein structures containing extein residues (Oeemig et al. 2012; Mizutani et al. 2002).
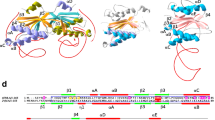
Structure of the Pab and Pho CDC21-1 inteins. a Primary structure comparison between the PabCDC21-1 and PhoCDC21-1 inteins. Conserved sequence Blocks A and B are indicated. b The crystal structure of the PabCDC21-1 intein. The final electron density map, contoured at 1.3 σ-level, is shown for the active site residues. c The crystal structure of the PhoCDC21-1 intein together with the fused extein (SUMO domain). The SUMO N-extein domain is colored in orange. a–c The insertion typical for thermophilic inteins is colored in red
Structure of PhoCDC21-1 intein
Pyrococcus horikoshii OT3 also contains the CDC21-1 intein (PhoCDC21-1) at the same site as the PabCDC21-1 intein within the P-loop region (UniProt: O58310). The PhoCDC21-1 intein consists of 168 residues, which is four residues longer than the PabCDC21-1 intein. Both share a pairwise sequence identity of 64% (108/168). We introduced the C1A mutation into the PhoCDC21-1 intein without additional C-extein sequence and fused it to an N-terminal H6-SUMO domain (yeast SMT3) for convenient purification (Guerrero et al. 2015). Typically, three residues between the cleavage site of the Ubiquitin-like specific protease 1 (Ulp1) and the first residue of an intein are sufficient for proteolytic cleavage of the SUMO purification tag (Guerrero et al. 2015). Interestingly, the three-residue “SGK” linker used between the H6-SUMO and PhoCDC21-1 intein in the fusion protein was too short for efficient cleavage by Ulp1. However, a very similar linker sequence was sufficient for the proteolytic removal of the H6-SUMO fusion from the PabCDC21-1 intein. Thus, we performed crystallization trials with the H6-SUMO-PhoCDC21-1 fusion protein without removal of H6-SUMO domain and were successful in obtaining diffracting crystals. We were able to solve the structure at 2.65-Å resolution by molecular replacement using the coordinates of the PabCDC21-1 intein and the SUMO domain (PDB: 1EUV) as search models. Overall, the crystal structure of the PhoCDC21-1 intein is very similar to that of PabCDC21-1 but includes the SUMO domain as N-extein (Fig. 1b–c).
Comparison between Pab and Pho CDC21-1 inteins
The HINT fold of the Pab and Pho CDC21-1 inteins is almost identical with an RMSD of 1.1 Å for 162 pairs of aligned Cα atoms, as expected from the high sequence identity of 64% (Fig. 2a). The main difference between the two structures is a four-residue loop extension located at the canonical HEN insertion site of the PhoCDC21-1 intein (Fig. 2b, c). The HINT domain subdivides into two subdomains, which are presumably the result of gene duplication and fusion, resulting in a C2-symmetry. PabCDC21-1 has an improved C2-symmetry interface with the loop regions at the C35 and N35 sites with hydrophobic residues L53, L58, and V60 for the N-terminal, and I121, L124, and V126 for the C-terminal subdomain (the C35 and N35 numbering for the split sites is based on the NpuDnaE intein as described in Aranko et al. (2014a) for convenience) (Fig. 2b). The 4-residue loop extension in the PhoCDC21-1 intein creates additional interactions, presumably stabilizing the two subdomains further. The additional interactions are made by hydrophobic and charged residues (Fig. 2c). The presence of stabilizing interactions within the extended loop is in line with previous observations in other thermophilic inteins, the Mja and Mvu TFIIB inteins, which showed that the artificial deletion of the HEN domain caused a significant deterioration of the protein splicing activity. However, introducing longer loops partially restored the splicing activity by compensating the removal of the HEN domain for the splicing activity (Fig. 3) (Iwaï et al. 2017). Similar results were also obtained for non-thermophilic inteins when HEN domains were removed (Hiraga et al. 2005). These observations suggest that the proper arrangement of the two subdomains in the HINT domain is critical for efficient protein splicing catalysis. Utilizing the loop at the C35 split site could be a general strategy of inteins to stabilize active conformations and might be of special importance for thermophilic inteins.
Comparison between PabCDC21-1 and PhoCDC21-1 inteins. a Stereoview of the superposition of the two crystal structures, PabCDC21-1 intein (light blue) and PhoCDC21-1 intein (magenta). b Schematic illustrations of the intramolecular interactions between the N35 (blue) and C35 (red) loops in the PabCDC21-1 intein. c Schematic illustrations of the intramolecular interactions between the N35 (blue) and C35 (red) loops in the PhoCDC21-1 intein. b–c An arrowhead indicates the HEN insertion site
Cartoon model of the stabilization mechanisms in inteins from thermophilic organisms. a The HINT fold has presumably emerged from gene duplication and fusion events. Stabilization by fixing the two subdomain orientations with the nested HEN domain (Iwaï et al. 2017). b The loop at the HEN insertion site adopts the stabilization function of the two HINT subdomains to countervail thermal motions at the elevated temperature in the case of hyperthermophilic CDC21 mini-inteins. The stabilizing interactions are highlighted in red
Unexpected interactions between the N-extein and PhoCDC21-1 intein
Despite very similar structures and sequences of the PabCDC21-1 and PhoCDC21-1 inteins, we could not proteolytically remove the fused SUMO domain from the PhoCDC21-1 intein using Ulp1 protease. Therefore, we crystallized the fusion protein. We observed clear electron densities for the entire SUMO domain in addition to the PhoCDC21-1 intein, except for the 30 N-terminal residues containing the His-tag. To our knowledge, this is the first intein structure with an entire extein domain in addition to the junction regions. Of interest are interactions that we observed between the SUMO domain and the PhoCDC21-1 intein (Fig. 4). These interactions were presumably absent in the H6-SUMO–PabCDC21-1 intein, leading to efficient digestion by Ulp1 protease. The interacting residues at the SUMO/intein interface locate mainly on one subdomain of the PhoCDC21-1 intein (residues 37–42, 71–75, and 146–147). The location also coincides with the β-turn of the extension found in inteins from hyperthermophiles (Figs. 1 and 4). On the SUMO domain, interacting residues mainly cover the C-terminal side. Because the amino-acid sequences at the interfaces are similar between the Pho and Pab CDC21-1 inteins, these interactions could be rather weak and depend on the covalent linkage of the SUMO domain to the PhoCDC21-1 intein as well as on the crystal packing. The poorer X-ray diffraction obtained for the crystal of the H6-SUMO-PhoCDC21-1 intein fusion could also be explained by the relatively large deviations of the SUMO structure with respect to the PhoCDC21-1 intein structure presumably because of the weak interactions. We previously observed that cis-splicing of the PhoCDC21-1 intein strongly depends on the fused exteins when artificial exteins are used—even while maintaining identical junction sequences (Ellilä et al. 2011). The observed extein–intein interactions confirm the structural basis of this extein-context dependency in addition to the well-known junction-sequence dependency common among inteins (Iwai et al. 2006; Oeemig et al. 2012; Aranko et al. 2009; Topilina et al. 2015).
Interactions between the PhoCDC21-1 intein and the N-extein (SUMO). The upper panel shows the view from the dorsal side of a horseshoe-crab shape. The lower panel shows the side-view. The N- and C-termini of the intein locate at the ventral side. The inserted β-strand and extended α-helix commonly found in inteins from hyperthermophiles face to the dorsal side. Residues contributing to the interaction between the HINT domain (gray) and the N-terminal SUMO extein (orange) are shown as teal and magenta sticks, respectively. The residues are highlighted in the primary sequences below. An Ala to Thr mutation in the SUMO domain in the used vector is underlined
Discussion
Many inteins contain an active or degenerated homing endonuclease domain, which has presumably played (or still plays) an important role in the propagation of intein genes via horizontal gene transfer (Gogarten et al. 2002; Barzel et al. 2011; Okuda et al. 2003; Koufopanou and Burt 2005). Even though many mini-inteins including PabCDC21-1 and PhoCDC21-1 are capable of protein splicing and hence independent of HEN domains for protein splicing, some inteins have developed a mutualism between the HEN and HINT domains. Examples include the TFIIB inteins from Methanococcus jannaschii and Methanocaldococcus vulcanius, both of which require the HEN domain for efficient protein splicing activity (Iwaï et al. 2017). The endonuclease domain nested in the MjaTFIIB intein is a degenerated HEN domain but presumably plays a role in fixing the orientation of the two pseudo-domains of the HINT fold, because deletion of the HEN domain caused a uniquely open conformation (Iwaï et al. 2017). The crystal structures of the PabCDC21-1 and PhoCDC21-1 inteins showed that both mini-inteins do not have large flexible loop insertions at the HEN insertion site, unlike the MjaTFIIB intein. The structure of the PabCDC21-1 intein revealed extensive hydrophobic interactions between the C2-symmetry-related N35 and C35 loops, increasing the subdomain interactions at the elevated temperature optimal for the growth of Pyrococcus abyssi (Fig. 2b). The four-residue insertion at the HEN insertion site of the PhoCDC21-1 intein further enhances the intramolecular stabilization between the two subdomains compared with the PabCDC21-1 intein (Fig. 2c). The extended loop at the HEN insertion site bridges the two subdomains via additional hydrophobic and electrostatic interactions which likely substitute the stabilization role of the HEN domain in inteins like MjaTFIIB (Fig. 3). The precise orientation of the subdomains stabilized by the loops might be of particular importance at elevated temperatures where hyperthermophiles inhabit.
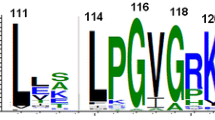
It has been puzzling why the HEN insertion site (C35 site) is extremely conserved among inteins. There is no obvious purifying selection of the HEN domains to exclusively locate at that specific position (Gogarten and Hilario 2006). If a free-standing endonuclease domain randomly invaded into inteins, inteins should exist in which HEN domains have inserted at other sites than the C35 site (Iwaï et al. 2017). For example, the C2-symmetry-related N35 site could be an equally suited insertion point. The structures of the hyperthermophilic CDC21-1 inteins might provide a hint for the high conservation of the HEN insertion site. The HINT fold has often been referred to as a flat horseshoe shape (Hall et al. 1997). Because of the C2 symmetry in the HINT fold, one can define an upper (dorsal) and lower (ventral) side (Figs. 4 and 5). The HINT fold, thus, rather resembles a horseshoe-crab shape with a carapace than a horseshoe shape due to the C2-symmetry (Fig. 4). The ventral side is where the splicing reaction takes place and the N- and C-termini of inteins locate (Figs. 4 and 5). In this architecture, the HEN insertion site locates on the dorsal side (Fig. 5a). The C2-symmetry-related site (N35) of the HEN insertion site points toward the ventral side of the intein structure. Any large insertion at the N35 site would clash with the extein region (host protein). We believe that this steric incompatibility with the host protein sequences (exteins) defines the conservation of the C35 HEN insertion site and restricts random insertions at other positions, including the N35 site. Indeed, all crystal structures of inteins with HEN domains (PI-PfuI, PI-PkoII, and PI-SceI) have open space below the ventral side of the HINT fold, providing room for their insertion into host proteins (Fig. 5b) (Mizutani et al. 2002; Ichiyanagi et al. 2000; Matsumura et al. 2006). The protein size of self-splicing inteins widely ranges from 123 to > 1000 residues, presumably because of repeated invasion and degeneration events of homing endonuclease domains (Novikova et al. 2016). Further investigations of the “inteinome” by elucidating intein structures will shed light on the historical invasion, fixation, and adaptation of inteins as molecular fossils of the intein emergence and evolution.
Structural features of the HEN insertion site on the horseshoe-crab shape of inteins. a A side-view of the PabCDC21-1 intein with the C2-symmetry around the HEN insertion site. The HEN insertion locates on the dorsal side of the horseshoe-crab shape. The thermophilic insertion is colored in red. b The locations of the HEN domains in the three crystal structures of PI-PfuI (PDB: 1DQ3), PI-PkoII (PDB: 2CW8), and PI-SceI (PDB: 1JVA) (Mizutani et al. 2002; Ichiyanagi et al. 2000; Matsumura et al. 2006). The HEN and HINT domains are colored in yellow and gray, respectively. The horseshoe-crab shape is indicated as red shape on top of the HINT domains
Methods
Cloning of PabCDC21-1 intein
The gene encoding the PabCDC21-1 intein was amplified from genomic DNA of Pyrococcus abyssi, DSM-25543, using the two oligonucleotides I725: 5′-AAAGGATCCGCGAAATGCGTCGATTATGAAACTG and I726: 5′-AAAGGTACCAAGTTGGCTGTTGTGAACGAAGATC. The PCR product was cloned between BamHI/KpnI of pHYRSF53 (Addgene #64696), resulting in plasmid pBHRSF151. The plasmid pBHRSF151 encodes a cis-splicing precursor with a H6-SUMO domain (yeast SMT3) and Chitin-Binding Domain (CBD) as N- and C-exteins, respectively. For structural studies, the C1A and N164A mutations were introduced into the PabCDC21-1 intein, and the C-terminal extein at the +1 position was replaced with a single Ala residue (T + 1A). The fusion protein was constructed by cloning the PCR product amplified from pBHRSF151 using the two oligonucleotides J303: 5′-AAAGGATCCGCGAAAGCCGTCGATTATGAAACTG and J304: 5′-GCGGTACCTTATGCAGCGTGAACGAAGATCCCATT between BamHI and KpnI sites of pHYRSF53, resulting in plasmid pBHRSF399 encoding the H6-SUMO-fused PabCDC21-1 (C1A, N164A, T + 1A) intein.
Cloning of PhoCDC21-1 intein
For the structural investigation, the PhoCDC21-1 intein with the C1A mutation was constructed as an N-terminally His-tagged SUMO fusion similar to the PabCDC21-1 intein variant described above. The gene of the PhoCDC21-1 intein was amplified from the previously published cis-splicing vector pSKDuet21 (Addgene #41695) in which the intein is flanked by two B1 domains of IgG binding protein G (GB1) with N-terminal His tag (Ellilä et al. 2011). For PCR amplification, the two oligonucleotides HK937: 5′-AGGATCCGGTAAGGCCGTTGACTACGATACAG and HK938: 5′-TTGGTACCTTAATTGTGCACGAATATTCCG were used. The PCR product was cloned into pHYRSF53-36 using the restriction enzymes BamHI and KpnI, resulting in pCARSF52 (Guerrero et al. 2015; Aranko et al. 2009).
Production and purification of Pab and Pho mini-inteins
The PabCDC21-1 and PhoCDC21-1 inteins were produced by expression of the above-described plasmids pBHRSF399 and pCARSF52 in E. coli strain T7 Express (New England Biolabs, Ipswich, USA). pCARSF52 was co-transformed with plasmid pRARE (Merck Millipore, Darmstadt, Germany) to account for rare codons. The transformed cells were grown at 37 °C in 0.5 L and 2 L LB expression cultures supplemented with 25 μg mL−1 kanamycin for the expression of PabCDC21-1 and PhoCDC21-1 inteins, respectively. Additionally, 5 μg mL−1 chloramphenicol was added for the plasmid pRARE for the expression of PhoCDC21-1 intein. The cultures were induced with a final concentration of 1 mM isopropyl-β-d-thiogalactoside (IPTG) for 3 h when OD600 reached 0.6. The induced cells were harvested by centrifugation at 4700 × g for 10 min, 4 °C and lyzed in 20 mL Buffer A (50 mM sodium phosphate, pH 8.0, 300 mM NaCl) using continuous passaging through an EmulsiFlex-C3 homogenizer (Avestin Inc, Ottawa, Canada) at 15,000 psi for 10 min, 4 °C. Lysates were cleared by centrifugation at 38,000 ×g for 60 min, 4 °C. The PabCDC21-1 intein was purified using 5 mL HisTrap HP columns (GE Healthcare, Chicago, USA) as previously described, including the removal of the N-terminal H6-SUMO fusion domain (Guerrero et al. 2015). For the PhoCDC21-1 intein, the second purification step involving proteolytic digestion with the SUMO-specific protease Ulp1 was omitted. Proteins were dialyzed against deionized water and concentrated using Macrosep® Advance Centrifugal Devices 10 K MWCO (PALL Corporation, New York, USA).
Crystallization of Pab and Pho mini-inteins
A 316 μM solution (6 mg/mL) of PabCDC21-1 intein (C1A, N164A, T + 1A) and 1.24 mM solution (40 mg/mL) of H6-SUMO-PhoCDC21-1 intein (C1A) were used for crystallization trials. Drops of 200 nl (100 nl concentrated protein and 100 nl of mother liquor) were set up in 96-well MRC (Molecular Dimensions, Suffolk, UK) crystallization plates using a Mosquito LCP® (TTP Labtech, Melbourn, UK). Diffracting crystals were obtained with mother liquor compositions of 100 mM sodium acetate pH 4.6, 30% (w/v) polyethylene glycol 4000, and 200 mM ammonium acetate for the PabCDC21-1 intein, and 1.8 M tri-ammonium citrate pH 7 for the PhoCDC21-1 intein. 20% glycerol was added for the latter on top of the drop, which served as a cryoprotectant when freezing crystals in liquid nitrogen.
Diffraction data collection and processing
PabCDC21-1 and PhoCDC21-1 crystals belonged to the space groups 20 (C2221) and 80 (I41), respectively. PabCDC21-1 intein diffraction data were collected on beamline I03 at the Diamond Light Source, Oxfordshire, UK and were subsequently indexed, integrated, and scaled to 1.60-Å resolution using the program XDS (Kabsch 2010; Allan et al. 2015). Diffraction data for the crystal of the PhoCDC21-1 mini-intein were collected on beamline I04-1 at Diamond Light Source and were processed to 2.65-Å resolution.
Structure determination and refinement
The structures of the PabCDC21-1 and PhoCDC21-1 inteins were solved by molecular replacement. The search model used for the PabCDC21-1 intein was based on the coordinates of the thermophilic TvoVMA intein (PDB: 4O1S) (Aranko et al. 2014b). The initial solution from Phaser was provided to ARP/WARP for auto-building (McCoy et al. 2007; Langer et al. 2008). The model was built with Coot, followed by rounds of refinement using the software Refmac5 and Phenix (Emsley et al. 2010; Murshudov et al. 2011; Adams et al. 2002). The entire polypeptide chain could be traced into the electron density map without breaks for all 168 residues. The quality of the final structure was validated using the MolProbity webserver (Table 1) (Chen et al. 2010).
For the molecular replacement solution of the PhoCDC21-1 intein, the coordinates of the PabCDC21-1 intein and the SUMO domain (PDB: 1EUV) were used as search models. The structure was solved with Phenix, followed by iterative rounds of the model building using ARP/WARP and Coot (McCoy et al. 2007; Langer et al. 2008). The initial 30 residues including the N-terminal His tag lacked sufficient density and were not modeled. The resulting model was rebuilt in place using Phenix and refined with Coot, Refmac5, Phenix, and, PDB-REDO (Emsley et al. 2010; Murshudov et al. 2011; Adams et al. 2002; Joosten et al. 2014). The final model was validated using the MolProbity webserver (Table 1) (Chen et al. 2010).
Cis-splicing analysis
Cis-splicing of PabCDC21-1 and PhoCDC21-1 inteins was analyzed by expressing the above-described constructs pBHRSF151 and pSKDuet21 with identical conditions as done for the production of samples for crystallization, except that the culture volume was reduced to 5 mL. Culture samples for SDS-PAGE analysis were taken before and 1 and 3 h after induction and cultures were harvested by centrifugation as described above. Cell pellets were resuspended and subsequently lysed in 400 µL B-PER reagent (Thermo Scientific, Waltham, USA) for 15 min under continuous shaking at 1000 rpm. Lysates were cleared by centrifugation at 12,300 × g for 5 min. His-tagged splicing products were IMAC purified at room temperature (RT) using Ni-NTA spin columns (Qiagen, Hilden, Germany) according to the protocol of the manufacturer. Proteins were eluted in 100 µL elution buffer (50 mM sodium phosphate, 300 mM NaCl, 250 mM imidazole, pH 8.0) and heated at 60 °C for 1 h before samples were analyzed by SDS-PAGE on 16.5% acrylamide gels and visualized using Coomassie Blue staining.
Accession numbers
Coordinates and structure factors have been deposited to the Protein Data Bank with accession numbers 6RPP for the PabCDC21-1 intein and 6RPQ for the PhoCDC21-1 intein.
Abbreviations
- CBD:
-
Chitin-binding domain
- GB1:
-
B1 domain of IgG binding protein G
- HEN:
-
Homing endonuclease
- HGT:
-
Horizontal gene transfer
- HINT:
-
Hedgehog/INTein
- IMAC:
-
Immobilized metal ion affinity chromatography
- IPTG:
-
Isopropyl-β-d-1-thiogalactopyranoside
- MCM:
-
Mini-chromosome maintenance
- Mja :
-
Methanococcus jannaschii
- Mvu :
-
Methanocaldococcus vulcanius M7
- Pab :
-
Pyrococcus abyssi
- PDB:
-
Protein data bank
- Pho :
-
Pyrococcus horikoshii OT3
- RMSD:
-
Root mean square deviation
- RT:
-
Room temperature
- SUMO:
-
Yeast SMT3 domain
- Ulp1:
-
Ubiquitin-like-specific protease 1
References
Adams PD et al (2002) PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr D 58:1948–1954
Allan DR et al (2015) Status of the crystallography beamlines at diamond light source. Eur Phys J Plus 130:56
Aranko AS, Züger S, Buchinger E, Iwaï H (2009) In vivo and in vitro protein ligation by naturally occurring and engineered split DnaE inteins. PLoS ONE 4:e5185
Aranko AS, Wlodawer A, Iwaï H (2014a) Nature’s recipe for splitting inteins. Protein Eng Des Sel 27:263–271
Aranko AS et al (2014b) Structure-based engineering and comparison of novel split inteins for protein ligation. Mol BioSyst 10:1023–1034
Barzel A, Naor A, Privman E, Kupiec M, Gophna U (2011) Homing endonucleases residing within inteins: evolutionary puzzles awaiting genetic solutions. Biochem Soc Trans 39:169–173
Brünger AT (1992) The free R value: a novel statistical quantity for assessing the accuracy of crystal structures. Nature 355:472–475
Chen VB et al (2010) MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr D 66:12–21
Dassa B, London N, Stoddard BL, Schueler-Furman O, Pietrokovski S (2009) Fractured genes: a novel genomic arrangement involving new split inteins and a new homing endonuclease family. Nucleic Acids Res 37:2560–2573
Ellilä S, Jurvansuu JM, Iwaï H (2011) Evaluation and comparison of protein splicing by exogenous inteins with foreign exteins in Escherichia coli. FEBS Lett 585:3471–3477
Emsley P, Lohkamp B, Scott WG, Cowtan K (2010) Features and development of Coot. Acta Crystallogr D 66:486–501
Gogarten JP, Hilario E (2006) Inteins, introns, and homing endonucleases: recent revelations about the life cycle of parasitic genetic elements. BMC Evol Biol 6:94
Gogarten JP, Senejani AG, Zhaxybayeva O, Olendzenski L, Hilario E (2002) Inteins: structure, function, and evolution. Annu Rev Microbiol 56:263–287
Guerrero F, Ciragan A, Iwaï H (2015) Tandem SUMO fusion vectors for improving soluble protein expression and purification. Protein Exp. Purif. 116:42–49
Hall TM et al (1997) Crystal structure of a Hedgehog autoprocessing domain: homology between Hedgehog and self-splicing proteins. Cell 91:85–97
Hiraga K, Derbyshire V, Dansereau JT, Van Roey P, Belfort M (2005) Minimization and stabilization of the Mycobacterium tuberculosis recA intein. J Mol Biol 354:916–926
Ichiyanagi K, Ishino Y, Ariyoshi M, Komori K, Morikawa K (2000) Crystal structure of an archaeal intein-encoded homing endonuclease PI-PfuI. J Mol Biol 300:889–901
Iwai H, Züger S, Jin J, Tam PH (2006) Highly efficient protein trans-splicing by a naturally split DnaE intein from Nostoc punctiforme. FEBS Lett 580:1853–1858
Iwaï H et al (2017) Structural basis for the persistence of homing endonucleases in transcription factor IIB inteins. J Mol Biol 429:3942–3956
Joosten RP, Long F, Murshudov GN, Perrakis A (2014) The PDB_REDO server for macromolecular structure model optimization IUCrJ 1:213–220
Kabsch W (2010) XDS. Acta Crystallogr D 66:125–132
Karplus PA, Diederichs K (2012) Linking crystallographic model and data quality. Science 336:1030–1033
Koufopanou V, Burt A (2005) Degeneration and domestication of a selfish gene in yeast: molecular evolution versus site-directed mutagenesis. Mol Biol Evol 22:1535–1538
Langer G, Cohen SX, Lamzin VS, Perrakis A (2008) Automated macromolecular model building for X-ray crystallography using ARP/wARP version 7. Nat Prot 3:1171–1179
Matsumura H et al (2006) Crystal structure of intein homing endonuclease II encoded in DNA polymerase gene from hyperthermophilic archaeon Thermococcus kodakaraensis strain KOD1. Proteins 63:711–715
McCoy AJ, Grosse-Kunstleve RW, Adams PD, Winn MD, Storoni LC, Read RJ (2007) Phaser crystallographic software. J Appl Cryst 40:658–674
Mizutani R et al (2002) Protein-splicing reaction via a thiazolidine intermediate: crystal structure of the VMA1-derived endonuclease bearing the N and C-terminal propeptides. J Mol Biol 316:919–929
Murshudov GN et al (2011) REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr D 67:355–367
Novikova O et al (2016) Intein clustering suggests functional importance in different domains of life. Mol Biol Evol 33:783–799
Oeemig JS, Zhou D, Kajander T, Wlodawer A, Iwaï H (2012) NMR and crystal structures of the Pyrococcus horikoshii RadA intein guide a strategy for engineering a highly efficient and promiscuous intein. J Mol Biol 421:85–99
Okuda Y et al (2003) Occurrence, horizontal transfer and degeneration of VDE intein family in Saccharomycete yeasts. Yeast 20:563–573
Paulus H (2000) Protein splicing and related forms of protein autoprocessing. Annu Rev Biochem 69:447–496
Perler FB et al (1994) Protein splicing elements: inteins and exteins—a definition of terms and recommended nomenclature. Nucleic Acids Res 22:1125–1127
Pietrokovski S (2001) Intein spread and extinction in evolution. Trends Genet 17:465–472
Topilina NI, Novikova O, Stanger M, Banavali NK, Belfort M (2015) Post-translational environmental switch of RadA activity by extein-intein interactions in protein splicing. Nucleic Acids Res 43:6631–6648
Volkmann G, Iwaï H (2010) Protein trans-splicing and its use in structural biology: opportunities and limitations. Mol BioSyst 6:2110–2121
Wood DW, Camarero JA (2014) Intein applications: from protein purification and labeling to metabolic control methods. J Biol Chem 289:14512–14519
Wu H, Hu Z, Liu XQ (1998) Protein trans-splicing by a split intein encoded in a split DnaE gene of Synechocystis sp. PCC6803. Proc Natl Acad Sci USA 95:9226–9231
Acknowledgements
We thank B. Haas, C. Albert, A. D. Hietikko, and Dr. V. Manole for technical help in protein and plasmid preparations. We thank Drs. T. Kajander and V. Manole for technical help at the crystallization facility. This work was supported in part by the Academy of Finland (131413, 137995, 277335), Novo Nordisk Foundation (NNF17OC0025402 to H. M. B., NNF17OC0027550 to H. I.) and Biocenter Finland, HiLIFE-INFRA for crystallization and NMR facilities at the Institute of Biotechnology. Open access funding provided by University of Helsinki including Helsinki University Central Hospital.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by H. Atomi.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Beyer, H.M., Mikula, K.M., Kudling, T.V. et al. Crystal structures of CDC21-1 inteins from hyperthermophilic archaea reveal the selection mechanism for the highly conserved homing endonuclease insertion site. Extremophiles 23, 669–679 (2019). https://doi.org/10.1007/s00792-019-01117-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00792-019-01117-4