Abstract
Copper-containing nitrous oxide reductase catalyzes a 2-electron reduction of the green-house gas N2O to yield N2. It contains two metal centers, the binuclear electron transfer site CuA, and the unique, tetranuclear CuZ center that is the site of substrate binding. Different forms of the enzyme were described previously, representing variations in oxidation state and composition of the metal sites. Hypothesizing that many reported discrepancies in the structural data may be due to radiation damage during data collection, we determined the structure of anoxically isolated Marinobacter nauticus N2OR from diffraction data obtained with low-intensity X-rays from an in-house rotating anode generator and an image plate detector. The data set was of exceptional quality and yielded a structure at 1.5 Å resolution in a new crystal form. The CuA site of the enzyme shows two distinct conformations with potential relevance for intramolecular electron transfer, and the CuZ cluster is present in a [4Cu:2S] configuration. In addition, the structure contains three additional types of ions, and an analysis of anomalous scattering contributions confirms them to be Ca2+, K+, and Cl–. The uniformity of the present structure supports the hypothesis that many earlier analyses showed inhomogeneities due to radiation effects. Adding to the earlier description of the same enzyme with a [4Cu:S] CuZ site, a mechanistic model is presented, with a structurally flexible CuZ center that does not require the complete dissociation of a sulfide prior to N2O binding.
Graphical Abstract
The [4Cu:2S] CuZ site in M. nauticus N 2O reductase. The electron density map shown is contoured at the 5 σ level, highlighting the presence of two sulfide ligands. 705x677mm (72 x 72 DPI)

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bacterial denitrification is the dominant respiratory metabolic pathway in many microoxic or anoxic environments [1]. Denitrifying microorganisms assemble an electron transport chain in the cytoplasmic membrane that couples the stepwise reduction of nitrate (NO3−) via nitrite (NO2−), nitric oxide (NO), and nitrous oxide (N2O) to dinitrogen (N2) to the generation of a proton motive force [2]. The final step of the pathway is a 2-electron reduction of N2O catalyzed by the copper-containing enzyme nitrous oxide reductase (N2OR) [3,4,5]. The process of fertilization in industrial agriculture creates nitrate-rich habitats that support the growth of denitrifying microbes to compete with crop plants, and N2OR then often is inactivated and gaseous N2O is released as a metabolic end product [6], resulting in a steady rise in atmospheric levels of nitrous oxide, in lockstep with the increasing fertilizer use during the past century [7]. With a global warming potential that exceeds the one of CO2 300-fold, nitrous oxide is a potent green-house gas, and it has an additional deleterious effect as an ozone-depleting substance. Its increasing concentration in the atmosphere has thus prompted its designation as the most critical anthropogenic emission in the twenty-first century [8].
Marinobacter nauticus, formerly known as Pseudomonas nautica or Marinobacter hydrocarbonoclasticus [9], is a denitrifying, marine Gammaproteobacterium that forms biofilms on hydrophobic organic matter but can also sustain a pelagic lifestyle [10]. Its N2OR is a 130 kDa homodimer encoded by the nosZ gene that contains two copper sites, CuA and CuZ [11]. CuA, a dinuclear, mixed-valent [Cu1.5+:Cu1.5+] center, transfers a single electron with a midpoint redox potential of + 65 ± 5 mV, at pH 7.6 [12]. It is located in the C-terminal cupredoxin domain of NosZ, similar to the CuA site in cytochrome c oxidase [13]. The N-terminal domain of NosZ, with a 7-bladed β propeller fold, holds the CuZ center, a tetranuclear copper cluster that is unique to N2OR. The enzyme from M. nauticus was the first to be characterized by X-ray crystallography, and an initial description of CuZ as a [4Cu:O] center [14] was subsequently revised to a [4Cu:S] stoichiometry [15], in line with available spectroscopic data [5, 16] and with further structural analyses of the enzymes from Paracoccus denitrificans [17] and “Achromobacter cycloclastes” [18]. In these structures, CuZ is a µ4-S-bridged Cu cluster with a total charge of +3, i.e., formal copper oxidation states of [1Cu2+:3Cu+] [3]. This state is characterized by a single band at 650 nm in electron excitation spectroscopy, arising from an S → Cu LMCT transition resulting in a blue color of the enzyme [19]. This simple spectrum is commonly masked by the more complex signature of the mixed-valent CuA yielding a pink enzyme, but can be revealed when the binuclear site is reduced to a colorless all-cuprous state with ascorbate [20]. The pink form of N2OR has been designated form II [5] and was obtained as the oxidized state of the enzyme when prepared in the presence of dioxygen [17, 18, 20]. A spectroscopically distinct, purple form I was first described for Pseudomonas stutzeri [21], but was later also found in Achromobacter xylosoxidans [22] and M. nauticus [23]. It shows an absorption maximum at 538 nm, and while it can also be reduced with ascorbate to a blue form, this resulting species consists of two bands at 552 nm and 650 nm and is thus distinct from the one described above [4]. In 2011, the purple N2OR from P. stutzeri was crystallized under anoxic conditions [24], revealing that here the enzyme contains a CuZ site of composition [4Cu:2S] [25]. The second sulfur atom in this site, SZ2, is labile and seems to be depleted—to varying degrees—in different batches of protein. This revised stoichiometry of the tetranuclear site explained the spectroscopic findings and clarified that the CuZ site of purple N2OR form I is a [4Cu:2S] cluster, while the desulfurated [4Cu:S] cluster of the pink form II enzyme was assigned as the CuZ* species defined previously by spectroscopy [4, 5]. In purple N2OR as isolated, CuZ is present in a [2Cu2+:2Cu+] state that cannot be reduced with ascorbate but converts to a [1Cu2+:3Cu+] state upon incubation with dithionite [3]. This state is distinct from the one found for CuZ* with respect to its EPR spectra. Based on a recombinant production system for active N2OR in E. coli that yielded catalytically active, purple N2OR with a [4Cu:2S] CuZ site [26], the systematic replacement of the seven histidine ligands to CuZ led to a complete loss of metalation in six cases, but to an unexpected spectrum resembling a form II enzyme in the case of the H382A variant [27]. However, rather than the [4Cu:2S] CuZ cluster commonly associated with this spectrum, the crystal structure of H382A N2OR contained a [3Cu:2S] center that was catalytically inactive but retained three copper ions liganded by sulfides, and it still produced the CT band at 650 nm. Interestingly, sulfide SZ2 was present in this cluster, but in the absence of the CuZ1 ion it bound in the native enzyme, it had shifted its position toward a nearby lysine. The implication was that the difference between the reported CuZ and CuZ* sites may not be the complete loss of sulfide SZ2 [28], but rather its dissociation from Cu1 that also leads to the loss of the 552 nm CT band, rendering the second sulfide invisible in UV/Vis spectra. If so, this may reconcile different hypotheses about the mode of N2O binding to the CuZ site and the mechanism of its reduction, as a mechanistic proposal for N2OR had suggested substrate binding to Cu1 and Cu4 of CuZ [29,30,31], which seemed incompatible with the presence of SZ2, but may well be feasible if the shift of SZ2 would render both Cu1 and Cu4 three-coordinate [27].
Considering these results, we revisited M. nauticus N2OR, whose initial analysis had a [4Cu:S] CuZ site with H2O bound at the Cu1–Cu4 edge (PDB 1QNI), providing the basis for the proposed mechanism [29]. The positive midpoint potential of copper sites in proteins makes them particularly prone to photoreduction or even more severe forms of radiation damage during diffraction data collection at high-flux X-ray sources, such as synchrotrons. We therefore analyzed the anoxically isolated, purple form of MnN2OR using the far lower photon flux of a rotating anode X-ray generator while retaining cryogenic conditions during data collection, aiming to generate a diffraction data set that was unperturbed by radiation-induced effects and more accurately represented the enzyme in its active form as isolated from its native host.
Experimental procedures
Cell growth and protein purification
M. nauticus was grown under denitrifying conditions in the presence of nitrate, as described elsewhere [16]. Cell disruption was carried out in an anoxic chamber with an N2/H2 (95%/5%) atmosphere (Coy Labs), where H2 was used to react with residual oxygen on a platinum catalyst surface to maintain pO2 < 1 ppm. During protein purification, samples, buffers, and columns were kept anoxic using modified Schlenk techniques. Approximately 180 g of cells (wet weight) were resuspended in 10 mM Tris/HCl buffer at pH 7.6 and broken in a microfluidizer (microfluidics). From the resulting crude extract, cell debris and membranes were sedimented by centrifugation at 100,000×g for 1 h. With the supernatant, all steps were carried out without breaks to avoid storage of the sample or freeze–thaw cycles. The soluble fraction was loaded onto a DEAE-FF anion exchange column (GE Healthcare) equilibrated with 10 mM Tris/HCl buffer at pH 7.6 and washed with the same buffer until A280nm reached baseline. The column was developed with a linear gradient from 10 to 500 mM Tris/HCl buffer, pH 7.6, from which N2O reductase eluted at approximately 300 mM Tris/HCl. The pooled fractions of the elution peak were then dialyzed against 10 mM Tris/HCl buffer at pH 7.6, and subsequently loaded onto a second anion exchange column (HiTrapQ, GE Healthcare) that was equilibrate in the same buffer. The column was again washed to baseline and developed with a linear gradient from 10 to 500 mM Tris/HCl buffer, pH 7.6, with elution of N2OR at approximately 300 mM Tris/HCl. As a final step, the eluate was loaded onto a Superdex 200 size exclusion column (Cytiva) equilibrated with 300 mM Tris/HCl buffer at pH 7.6. The pure enzyme eluted as a symmetric peak and was concentrated by ultrafiltration (50 kDa MWCO, Sartorius) to 14 mg mL−1, while at the same time, the buffer was exchanged to 10 mM Tris/HCl, pH 7.6, for crystallization. Protein was determined using the bicinchoninic acid method [32], using bovine serum albumin as protein standard.
Crystallization and data collection
MnN2OR was crystallized by sitting drop vapor diffusion in an anoxic glove box (Coy Labs) at 303 K, in an N2/H2 atmosphere with < 1 ppm O2. 2 µL of protein solution (14 mg mL−1) were mixed with 2 µL of a reservoir solution containing 15% (w/v) polyethylene glycol 8000, 0.6 M NaCl, and 0.1 M imidazole/malate buffer at pH 7.2 and equilibrated against 50 µL of the same reservoir solution in a sealed crystallization plate (SwissSci). Large plate-like crystals were obtained in less than 1 day. For data collection, a crystal was transferred into a harvesting buffer that consisted of the reservoir solution with 15% (v/v) of 2R,3R-butane diol as a cryoprotectant. The crystal was then flash-cooled in liquid nitrogen before being removed from the anoxic chamber. Diffraction data were collected on a rotating copper anode X-ray generator (Rigaku MicroMax 007HF with a marresearch mar345dtb image plate system) at a wavelength of λ = 1.5418 Å and a temperature of 100 K.
Structure solution and refinement
The data set was integrated with XDS [33] and scaled with AIMLESS [34] from the CCP4 suite [35]. Molecular replacement with MOLREP [36] was used to locate two copies of the dimeric enzyme in the asymmetric unit, using the previous structure of MnN2OR (PDB 1QNI) as a search model. Model building was conducted in COOT [37], and REFMAC5 [38] was used for refinement. The quality of the final model was evaluated with MolProbity [39]. Data collection and refinement statistics are summarized in Table 1.
Results and discussion
Diffraction data from a plate-like single crystal of MnN2OR were collected using Cu Kα radiation (λ = 1.5418 Å) on a rotating anode X-ray generator with a mar345dtb image plate detector at a temperature of 100 K. The crystal belonged to the triclinic space group P1 and diffracted beyond the detector edge at the minimal possible crystal-to-detector distance, yielding data to 1.5 Å resolution with a mean I/σ(I) of 2.6 in the highest resolution shell (Table 1). The data set proved to be of outstanding quality and the excellent electron density map compared favorably to the best available data obtained from synchrotron sources, while presumably not suffering from radiation-induced changes in the architecture of the metal sites. Notably, the quality of the map also far exceeded that of the earlier, synchrotron-derived structure from the same organism that was refined to 2.4 Å resolution (PDB 1QNI) [14, 15]. This crystal form contained two N2OR dimers in the asymmetric unit and thus four independent observations of each feature.
Overall structure
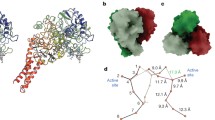
The low-dose structure of MnN2OR aligns with the previous model with a root-mean-squared deviation (r.m.s.d.) of 0.25 Å for 7727 atoms [14, 15]. The additional level of detail offered by the improved resolution allows for a detailed comparison with structures of the enzyme from other organisms. MnN2OR forms the typical head-to-tail homodimer (Fig. 1A), with the N-terminal, seven-bladed β-propeller domain of one monomer in close contact with the C-terminal cupredoxin domain of the other (Fig. 1B). At the interface of both domains, the CuA site closely approaches the CuZ cluster of the other subunit to form a combined active site that is accessible via a hydrophobic substrate channel running along the subunit interface, with a water-filled cavity behind the CuZ cluster (Fig. 1C). These features were previously observed in a structure of PsN2OR prepared under anoxic conditions after pressurization with N2O gas, in which the substrate N2O was bound between the two copper sites [25]. The gaseous substrate N2O and product N2 enter and exit via this hydrophobic channel, while the second product, H2O, can exit through the water-filled cavity [4].
Low-dose structure of M. nauticus N2OR. A The head-to-tail homodimer of N2OR with protomer A in green, protomer B in blue. The CuA site of one protomer forms a combined active site with the CuZ site of the other. B Cartoon representation of a NosZ monomer with the distinct two-domain architecture, colored from blue at the N-terminus to red at the C-terminus. The N-terminal domain forms a seven-bladed β-propeller with CuZ at its center, while CuA is bound to the C-terminal domain, which attains the cupredoxin fold. The positions of the ions discussed in the text are indicated. C A hydrophobic channel along the dimer interface allows for access of the substrate N2O and egress of product N2, while the second product water is released into a water-filled exit channel that leads back to the protein surface. D The seven blades of the N-terminal domain and CuZ at the hub of the propeller. Except for blade IV, each blade contributes a histidine ligand to CuZ. Blade II contributes two consecutive histidines, to a total of seven ligands to the four copper ions of the site
The electron transfer site CuA
Originally described as one of the two spectroscopically distinct copper sites in respiratory cytochrome c oxidase, the exact nature of CuA remained under debate (‘embattled’, in the words of Helmut Beinert) for years [40, 41]. In EPR, CuA appeared with an unusually small hyperfine splitting around g⊥ = 2.18 that in most cases was not even resolved into what was expected to be a four-line pattern for a mononuclear copper, and the presence of hemes a and a3 in the oxidase further obfuscated the signature of the copper site. In 1989, Dooley and Zumft showed that PsN2OR also contains a CuA-type site that is not masked by the presence of heme, and after some exchange on the nature of CuA [42, 43], Kroneck, Zumft, and others established along various lines the dinuclear, valence-delocalized nature of the site even before a first crystal structure of cytochrome c oxidase became available [44, 45]. In structures of cytochrome c oxidases, N2O reductases and diverse natural and engineered CuA domains [19, 46,47,48], two Cu ions form a central rhomb with the thiols of two cysteines, and the distorted tetrahedral ligand environment of the coppers is completed by a histidine at each metal, plus a methionine for CuA1, the ion more distal from the CuZ cluster (Fig. 2), and a backbone carbonyl for CuA2. The same arrangement was found and modeled in both CuA sites within the MnN2OR dimer (Fig. 2A, B), with C614 and C618 as bridging cysteines, H579 and M625 as ligands to CuA1, and H622 and the backbone carbonyl of W616 as ligands to CuA2 (Fig. 2C). The initial analysis of MnN2OR had established these residues and modeled all six occurrences of CuA in the asymmetric unit in the same conformation [14]. This uniform nature of CuA was first challenged with the analysis of PsN2OR under anoxic conditions [25], where the corresponding CuA sites now attained two distinct conformations. Typically, in one of the two copies of the monomer, the histidine ligand to CuA2 (H583 in this organism) had its imidazole moiety rotated away from CuA1 by 130° to form a short hydrogen bond to a nearby serine (S550). In the following, this will be designated the OUT-conformation. Only in structures with bound N2O, this histidine was consistently flipped back to the copper (the IN conformation), and we hypothesized at the time that this may represent a mechanism to gate electron transfer during catalysis [25]. Upon establishment of a recombinant production system for a functional PsN2OR in E. coli, [26] the metal-replete form I structure had both CuA sites in the OUT-conformation at CuA1. For the low-dose structure reported here, generated from protein isolated from actively denitrifying M. nauticus, the conformations of the CuA sites were therefore of particular interest.
The CuA site in MnN2OR. A 2Fo–Fc difference electron density map around the CuA site in protomer A, contoured at the 1σ (grey) and 5σ (blue) levels. Orientation as in (C). B The corresponding representation for the CuA site in protomer B. C CuA in protomer A in the IN conformation of H579 with the ligands of the first coordination sphere. D Bond distances (in Å) and angles in the 2Cu:2S diamond core of CuA in protomer A, showing distortion from an ideal geometry, in particular in a long bond from CuA2 to the Sγ atom of C614. E Contrary to [2Fe:2S] clusters, the ligand field of the copper ions is not very close to tetrahedral. While CuA1 remains largely tetrahedral, with an extended bond to M625, the environment of CuA2 is better described as trigonal pyramidal, with the backbone carbonyl of Q616 as an axial ligand and the Cu ion in the plane of the other three, C614, C618, and H622. F CuA with H579 in the OUT-conformation in protomer B, rendering CuA1 three-coordinate. G The 2Cu:2S core now shows much stronger distortion, with the CuA2–C614-Sγ bond elongated to 2.83 Å. H With the release of H579, CuA2 moves almost fully into the plane of the remaining ligands, leading to a trigonal planar coordination environment. CuA2 remains largely unchanged. Panels (A), (B), (C), and (F) are rendered for wall-eyed stereo viewing
The two N2OR dimers of the asymmetric unit indeed differed in the conformation of residue H579, which was a ligand to CuA1 in the A protomer of both dimers (IN conformation) (Fig, 2A, C), but rotated toward S546 in protomer B (OUT-conformation) (Fig. 2B, F). These positions corresponded to the two conformations observed in the first PsN2OR structures, and the precise definition of both sites now allowed to resolve distortions and asymmetries of the metal centers. In the IN conformation of protomer A, the Cu2S2 rhomb was already slightly distorted: At a short Cu–Cu distance of 2.47 Å (compared to 2.5 Å in the earlier analysis with synchrotron data at 2.4 Å resolution [14]), the CuA2–Sγ-C614 distance was extended to 2.6 Å, and the CuA1–Sγ-C618 distance was slightly elongated to 2.37 Å. This conformation is more similar to all other reported CuA sites except for P. stutzeri. CuA1 had a nearly ideally tetrahedral ligand field with three sulfurs and a nitrogen, and all bond distances between 2.27 Å and 2.46 Å (Fig. 2E). The latter distance to the Sδ of M625 was far shorter than for typical methionine ligands in type I copper centers [49, 50]. In contrast, the geometry of CuA2 diverged farther from an ideal tetrahedron, with the metal residing almost exactly in the plane of ligands C614, C618, and W616 (Fig. 2E), implying that it is better described as trigonal pyramidal. Interestingly, the second longest bond distance after the one to C618 was 2.51 Å to the backbone carbonyl of W616, suggesting a rather weak Cu–O bond. When in protomer B residue H579 rotated into the OUT-conformation (Fig. 2F), the resulting geometrical distortion of the Cu2S2 core was quite significant (Fig. 2G). With slight changes in distances and angles, the rhomb became more asymmetric, and the CuA2–Sγ-C614 bond weakened further, its length increasing to 2.83 Å. The Cu–Cu distance contracted only slightly to 2.44 Å. The most obvious difference, however, was geometric. While the environment of CuA2 changed only with respect to C618, the missing H579 now caused CuA1 to move almost fully into the plane of C614, C618, and M625, so that the coordination geometry became trigonal planar, while the environment of CuA2 remained trigonal pyramidal (Fig. 2H). For PsN2OR, we previously suggested that H579 serves as a gatekeeper for electron and proton transfer in the enzyme [25, 51, 52], and the distortions of CuA observed here very likely also reflect in distinct changes of the midpoint potential of the site. Furthermore, the methionine ligand to CuA1 was frequently designated as a weak axial ligand at this position, and the influence of the protein matrix was suggested to keep it bound, while at the same time lowering the midpoint redox potential [53]. More recently, an NMR analysis revealed significant spin density on the methionine, in line with a stronger binding and an involvement in electron transfer [54]. The present analysis adds an additional level of complexity to the electronic properties of CuA, in that the methionine ligand to CuA1 remains bound to the metal in both states but undergoes changes both in bond length and geometry.
CuZ is a [4Cu:2S] cluster
Adjacent to CuA but in the other protomer of the N2OR dimer, CuZ is located at the hub of the N-terminal seven-bladed β-propeller domain, coordinated by seven histidine ligands: H132, H133, H181, H323, H378, H429, and H490. The fourth propeller blade that also contains the Ca2+-binding loop (vide infra) is the only one that does not contribute a ligand to CuZ, while the second blade provides two. The first blade also provides a histidine, although its innermost β-strand is contributed by the C-terminus of the domain (Fig. 1D). Contrary to the IN- and OUT-conformations of the CuA sites, the four individual observations of CuZ were indistinguishable in the low-dose structure, where CuZ consistently was a [4Cu: 2S] site, with a largely symmetric Cu2S2 diamond core that is reminiscent of CuA (Fig. 3A). With four metal ions, two sulfides, and seven histidine ligands, CuZ is a uniquely asymmetrical metal site, and the architectures reported in previous structure determinations varied substantially, in particular with respect to the CuZ1–CuZ4 edge of the cluster [51]. In the present structure, CuZ seemed unperturbed, which was also reflected in a perfect alignment of all seven histidines with the coordinated coppers, which was not always the case in previous structures (Fig. 3A). Five of the six ions of CuZ were almost perfectly in a plane, with only CuZ3 outside at 2.16 Å distance (Fig. 3B). Each Cu ion was liganded by two histidines, with the exception of CuZ4, to which H490 was the only protein ligand (Fig. 3B,C). The Cu2S2 rhomb in CuZ is formed by CuZ1, CuZ4 and the two sulfides SZ1 and SZ2. The Cu–S bonds to SZ1 were shorter than those to SZ2, and the CuZ1–CuZ4 distance of 3.1 Å was substantially longer than the metal–metal distance in CuA (Fig. 3C). Due to this, the angles in this rhomb were closer to orthogonal than in CuA. Arguably, the most unusual feature of this unique cluster is the arrangement of four Cu ions around the central sulfide SZ1. Three of the metal ions, CuZ2, CuZ3, and CuZ4, form a nearly equilateral triangle that is completed to a tetrahedron by SZ1, and the bond distances in this unit are remarkably short. The three copper ions had distances of 2.16 ± 0.03 Å to SZ1, and the distances of 2.61 ± 0.01 Å for CuZ2–CuZ4, 2.64 ± 0.01 Å for CuZ2–CuZ3, and 2.81 ± 0.01 Å for CuZ3–CuZ4 are indicative of metal–metal bonding. They fall in the range of distances also observed in CuA sites in other analyses, while these distances in CuA are shorter in the present structure (Fig. 2D, G). Notably, this tetrahedral structure was retained in a H382A variant of recombinant PsN2OR [27], although CuZ1 was lost due to the removal of the histidine that corresponds to H378 in MnN2OR (Fig. 3E). In this structure, even the second sulfide SZ2 was still present, now as a dangling ligand with a long hydrogen bond to K545 (K450 in MnN2OR). It may well be the stability of this tetrahedral core that allows for the flexibility of sulfide SZ2, as observed in previous analyses.
The [4Cu:2S] CuZ site in MnN2OR. A Stereo image of a 2Fo-Fc difference electron density map around the CuZ site in protomer B, contoured at the 1σ (grey) and 5σ (blue) levels. B Front view of CuZ, highlighting the plane formed by CuZ1, CuZ2, CuZ4 and the two sulfides. C Top view of CuZ, showing the Cu2S2 rhomb of CuZ1 and CuZ4 and the two sulfides, with distances in Å an angles. D Sulfide SZ1 coordinates all four copper ions. It forms a tetrahedron with short bond distances to CuZ1, CuZ2, and CuZ3, with short Cu–Cu distances indicative of metal–metal bonding. E In an H382A variant of PsN2OR, the removal of histidine 382 led to the loss of CuZ1. The remainder of CuZ, with the now dangling SZ2 and the Cu3S tetrahedron remained largely unchanged [27]
In the synchrotron data structure of MnN2OR, the position of sulfide SZ2 was modeled as a water molecule bridging CuZ1 and CuZ4 [14, 15]. However, the position of this ligand coincided very well with SZ2, and the six observations of CuZ in the asymmetric unit of these crystals yield bond distances of 2.7 ± 0.2 Å to CuZ1 and 2.3 ± 0.1 Å to CuZ4 (stating only a single decimal at 2.4 Å resolution). These values agree with the Cu–SZ2 distances in the present analysis (Fig. 3C), but are longer than typical water ligands to copper in the range of 1.9–2.1 Å. A structurally flexible or partly occupied SZ2 could have been interpreted as a bridging water at moderate resolution and without additional information from anomalous scattering.
Further ion-binding sites in N2O reductase
All known structures of N2O reductases contain additional ion-binding sites beside the copper centers CuA and CuZ. Contrary to many other observations of surface-bound ions, these are specific sites, buried within the protein, and likely of functional importance. Although we only used diffraction data collected at a single energy, its high quality allowed us to evaluate the distinct anomalous scattering contribution of individual atoms as a means of identification. According to the Sasaki tables for anomalous scattering, the anomalous contribution of Ca2+ at the Cu Kα energy (8014 eV) is 1.29 e−, while that of K+ is 1.08 e− [55], and there is a similar difference in the scattering contribution for Cl– and S2− (Fig. 4A). Note that all these ions are isoelectronic and therefore virtually indistinguishable in common difference electron density maps. The anomalous scattering signal is notoriously weak, but high data accuracy and a crystal form with four independent observations of each ion-binding site in the two dimers of the asymmetric unit allowed for a reliable evaluation according to the tabled values. For all sites, the refined B-factors agreed with their surroundings, indicating full occupancy (Fig. 4A).
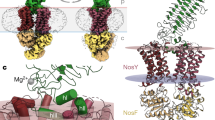
Additional ion-binding sites in MnN2OR. A With the same number of electrons, Ca2+, K+, Cl–, and S2– cannot be distinguished in 2Fo–Fc difference electron density maps, but their anomalous scattering contribution at Cu K∝ energy differs slightly (left). The integrated anomalous difference density of the ion positions (right) scales very well with these expected differences. The depicted N2OR dimer highlights the positions of the ions and corresponds to panel (E). B Stereo representation of the Ca2+ site in the β-propeller domain, with the typical bidentate coordination by two carboxylate ligands. C Stereo rendering of the K+ site. The octahedral ligand field includes four waters, but only one carboxylate oxygen. D Stereo image of the Cl−-binding site located directly adjacent to CuZ [see (E)]. R184 as a positively charged ligands supports the assignment as an anion. All electron density maps are 2Fo-Fc difference maps contoured at the 1σ (grey) and 5σ (blue) levels. E The relative positions of the ion sites with respect to the CuA and CuZ sites. Note that the K+ ion is placed in the dimer interface, while Ca2+ stabilizes a loop that clamps the β-propeller domain of protomer A to the cupredoxin domain of protomer B [56]
The first site in the enzyme is a Ca2+ ion bound in the N-terminal β-propeller domain, connecting β-strands 2 and 3 of the fourth blade (Figs. 1D, 4B). With bound Ca2+, the loop contacts the C-terminal cupredoxin domain of the other monomer, stabilizing its interaction with the β-propeller, while in an apo-structure of Shewanella denitrificans N2OR, the loop was disordered in the absence of the cation [56]. Ca2+ binding raised the Tm of the protein, suggesting that this binding event occurs after the insertion of the copper sites to provide stability to the mature enzyme [56]. Interestingly, a recent structure of apo-N2OR bound to the copper site maturation complex NosDFY already had Ca2+ bound, although both copper centers were absent [57]. However, this structure was only obtained with an inactive E154Q variant of the ABC protein NosF that rendered the complex incompetent for copper uptake and transfer to N2OR. In the present analysis of MnN2OR, the Ca2+ site was fully defined and showed a typical coordination environment that in essence was octahedral but included two bidentate ligands, E260 and D274. Further ligands to Ca2+ were the amide oxygen of N321, the backbone amide oxygens of residues Y257 and T268, and a water molecule (Fig. 4B).
In the initial structural models for N2OR from P. denitrificans and M. nauticus, a further ion-binding site was modeled as a second Ca2+ [14, 15, 17, 18]. This was contested in the analysis of PsN2OR, where the conserved feature was interpreted as a coordinated K+ ion [25]. As calcium and potassium ions are not readily discriminated in an electron density map, the point of distinction made was the coordination environment that is the exact same in the present structure, showing a nearly undistorted octahedral geometry. Four of the six ligands to the ion were water molecules, and only one acidic residue, E465, acted as a monodentate ligand, compensating the charge of K+. The sixth ligand was the backbone carbonyl of the highly conserved K450, whose Nζ amino group is close to CuZ. It was suggested to play a functional role during catalysis, as a proton donor for the reaction [58,59,60] and in supporting conformational flexibility of the CuZ site [27] (Fig. 4C). The integrated anomalous scattering contribution for the Ca2+ sites exceeded that for K+ by exactly the expected amount (Fig. 4A). Together with the finding that upon Ca2+ reconstitution of the otherwise similar S. denitrificans N2OR, only one site is occupied by the divalent cation [56], we conclude that this site, as in PsN2OR [25], binds K+ specifically. The K+ site is located on the surface of the CuZ domain, in the interface with the CuA domain of the other N2OR protomer (Fig. 4E). Although four of its ligands are water molecules, these are buried within the protein matrix of the dimer, stabilized by a tight hydrogen-bonding network and thus not readily exchangeable. The K+ site furthermore is in close (if indirect) contact with CuA, via hydrogen bonds from water ligands, on one hand to the Nε1 atom of the side chain of W616 that coordinates CuA through its backbone carbonyl, and on the other hand to the backbone carbonyl of F617, a conserved residue that together with M623 controls access to the N2O binding site on CuZ [25]. A conserved and possibly functionally essential residue is E488 that forms a hydrogen bond to a water ligand of K+, but also to the backbone amide of F617. It also forms a salt bridge to the side chain of K450, orienting it near CuZ and providing a compensating charge (Fig. 4E). Taken together, F617, E488, and K450 form an efficient electron conduit between the CuA and the CuZ site. Alternatively, we have suggested that the exact same arrangement in PsN2OR (residues F621, E492, and K454) acts as a proton conduit that runs along the CuA site and is gated by the conformational change of the distal histidine ligand to CuA, H579 [52]. The present analysis of the MnN2OR structure is fully in line with this. The second ion-binding site in N2OR thus holds a K+ ion that plays a very specific role in connecting the CuA and CuZ sites, integrating them into a single functional active site within N2OR. For N2OR assembly, this choice of ions is deliberate. Calcium binding locks the structure in a closed state that no longer allows for the insertion of copper during the maturation of the enzyme in the periplasm [56], while the position of K+ indicates that it already is inserted upon dimerization of apo-NosZ. This step occurs in the cytoplasm prior to the export of the enzyme as a folded dimer via the Tat system, and the K+ site is no longer accessible after the dimer has formed [56]. It is, therefore, important that the correct ion binds in the proper sequence in the right cellular compartment.
One additional ion-binding site was consistently observed in all structures of N2OR enzymes reported to date. Located close to the CuZ site, a significant electron density peak appeared that could not be modeled as water. Due to its position close to the positively charged R184 (M. nauticus numbering) and an electron density peak with approximately twice the magnitude of a water (after occupancy refinement), this feature was modeled as a chloride anion. Besides R184, the only other amino acid that directly ligated this ion was H378, a ligand to CuZ1 (Fig. 4D). The role of this ion is unclear, although its charge may help in shaping the electrostatic environment of the CuZ site. Moreover, chloride is not easy to distinguish from the isoelectronic hydrosulfide anion, HS−, adding to the enigma of the possible flexibility of SZ2, as discussed above. The average anomalous signal peak intensity at this position is slightly higher than that for all cysteine sulfides in the structure, but as the varying B-factors of these amino acids in different parts of the structure play strongly into this measurement and the anomalous signal strength for chloride should only be 14% higher than for sulfide, the distinction is not a clear as for the case of K+ versus Ca2+. We therefore used only the most well-defined sulfur atoms in the structure, the cysteine Sγ atoms, but not the Sδ of methionines. Here, the sulfur anomalous signal perfectly coincided with the tabled values, while the four presumed Cl− sites showed a stronger signal that corresponded to the Sasaki values for this element (Fig. 4A). Our analysis of the anomalous scattering properties of all ion sites supports an assignment as Cl– rather than S2–, but its influence on the properties of CuZ—if any—remains to be understood.
Implications for the catalytic mechanism of N2OR
The electron-transferring CuA and the unique CuZ combine at a dimer interface to create an intricate active site for the reduction of inert N2O. The observation of two distinct conformations of CuA in the present analysis is in line with our earlier findings on PsN2OR [25, 26], but it has not been reported for any of the multiple other descriptions of mixed-valent CuA sites. Most intriguingly, the precise definition of two conformations for CuA implies that the C2-symmetric N2OR dimer must pack in the crystal lattice in a non-random fashion, as otherwise the two conformations of the site should be randomly distributed, and the resulting electron density maps should represent an average of both conformations. We initially assumed that the IN- and OUT-conformations of CuA might lead to changes at the protein surface that could then affect crystal packing, but we could not identify any significant differences between the two protomers. Both the P. stutzeri and the present M. nauticus enzyme crystallized in the lowest-symmetry space group, triclinic P1, resulting in a rather simple crystal packing that may prevent that different conformations average out, so that it remains possible that this feature could not be discerned in other structures. Note also that the serine required for the OUT-conformation (S546 in MnN2OR) is conserved in N2OR sequences, but not in the CuA domains of oxidases or the engineered sites in generic cupredoxins. Mechanistically, a gating mechanism for electron transfer to and through CuA is highly interesting. CuA is a single-electron transfer site, so that the 2-electron reduction of N2O to N2 should proceed in two consecutive reduction steps, possibly with an enzyme-bound intermediate, formally [N2O]−. In this case, the role of CuZ would be to bind and activate N2O as a prerequisite for electron transfer from CuA. The finding that crystals of N2OR bind N2O between the CuZ and CuA sites upon pressurization with the gas is well in line with this model [25]. However, we so far failed to isolate and analyze a one-electron-reduced intermediate of the reaction, which should be formed if enzyme with reduced CuA was incubated with N2O in the absence of further reductant. In addition, N2OR exhibited higher catalytic activity when CuZ was reduced to an all-cuprous state with 4 Cu(I) [60]. Turnover from this highly reduced state led to a one-electron-oxidized form, designated CuZ0. While at the same redox level as the CuZ* state, only CuZ0 could be reduced from CuA using a physiological reductant [60]. The mechanistic implication is that for the reduction of N2O, one electron is provided by CuZ and the other by CuA. This results in a concerted 2-electron reduction, presumably without a long-lived intermediate. The observed transition from an OUT to an IN conformation upon N2O binding [25] may be key to this process, as the geometric changes reported here likely affect the midpoint potential for CuA and potentially serve as a trigger for electron transfer to the substrate.
The observed uniformity of the unique CuZ cluster indicates that the heterogeneity seen before at this site may indeed have been due to radiation damage or photoreduction processes during data collection. Available structures mostly differ in the ligand or ligands bridging CuZ1 and CuZ4, but not in the tetrahedral core formed by CuZ2-4 and SZ1. Previously, this was interpreted as lability of SZ2 that was postulated to fully dissociate from the cluster to facilitate N2O binding as a bridging ligand to CuZ1 and CuZ4 [29, 61]. The H382A variant of PsN2OR then revealed that the spectroscopic differences between form I and II of the enzyme, or the CuZ and CuZ* form of the active site cluster, are also reproduced if SZ2 dissociates only from CuZ1 to interact with the nearby lysine [27]. This flexibility can rationalize that SZ2 in many structures seems to have lower electron density (leading to an assignment as OH−) or even two distinct positions that were then interpreted as bound water. Further work will be required to confirm this hypothesis and deliberately populate the two conformational states for in-depth analysis. Also, in many cases, one or several of the histidine ligands to the copper ions are not ideally oriented toward the metal, indicating that the observed structures may be partially demetallated. This is another typical effect of radiation damage, and the structural uniformity of CuZ in the present analysis at low X-ray dose underscores the value of this approach for the analysis of sensitive metal sites in general and the intrinsically flexible CuZ in particular.
Data availability
The atomic coordinates and experimental structure factor amplitudes for the low-dose structure of M. nauticus N2OR have been deposited with the Protein Data Bank at http://www.pdb.org with the accession number 9F8X.
Abbreviations
- N2OR:
-
Nitrous oxide reductase
- EPR:
-
Electron paramagnetic resonance
- MR:
-
Molecular replacement
References
Zumft WG (1997) Microbiol Mol Biol Rev 61:533–616
Einsle O, Kroneck PMH (2004) Biol Chem 385:875–883
Pauleta SR, Dell’Acqua S, Moura I (2013) Coord Chem Rev 257:332–349
Wüst A, Schneider L, Pomowski A, Zumft WG, Kroneck PMH, Einsle O (2012) Biol Chem 393:1067–1077
Zumft WG, Kroneck PMH (2007) Adv Microb Physiol 52:107–225
Canfield DE, Glazer AN, Falkowski PG (2010) Science 330:192–196
Schmittner A, Galbraith ED (2008) Nature 456:373–376
Ravishankara AR, Daniel JS, Portmann RW (2009) Science 326:123–125
Sproer C, Lang E, Hobeck P, Burghardt J, Stackebrandt E, Tindall BJ (1998) Int J Syst Bacteriol 48:1445–1448
Gauthier MJ, Lafay B, Christen R, Fernandez L, Acquaviva M, Bonin P, Bertrand JC (1992) Int J Syst Bacteriol 42:568–576
Prudêncio M, Pereira AS, Tavares P, Besson S, Moura I (1999) J Inorg Biochem 74:267–267
Carreira C, dos Santos MMC, Pauleta SR, Moura I (2020) Bioelectrochemistry 133:107483
Iwata S, Ostermeier C, Ludwig B, Michel H (1995) Nature 376:660–669
Brown K, Tegoni M, Prudêncio M, Pereira AS, Besson S, Moura JJ, Moura I, Cambillau C (2000) Nat Struct Biol 7:191–195
Brown K, Djinovic-Carugo K, Haltia T, Cabrito I, Saraste M, Moura JJG, Moura I, Tegoni M, Cambillau C (2000) J Biol Chem 275:41133–41136
Prudêncio M, Pereira AS, Tavares P, Besson S, Cabrito I, Brown K, Samyn B, Devreese B, Van Beeumen J, Rusnak F, Fauque G, Moura JJG, Tegoni M, Cambillau C, Moura I (2000) Biochemistry 39:3899–3907
Haltia T, Brown K, Tegoni M, Cambillau C, Saraste M, Mattila K, Djinovic-Carugo K (2003) Biochem J 369:77–88
Paraskevopoulos K, Antonyuk SV, Sawers RG, Eady RR, Hasnain SS (2006) J Mol Biol 362:55–65
Solomon EI, Heppner DE, Johnston EM, Ginsbach JW, Cirera J, Qayyum M, Kieber-Emmons MT, Kjaergaard CH, Hadt RG, Tian L (2014) Chem Rev 114:3659–3853
Dell’Acqua S, Pauleta SR, Moura I, Moura JJG (2011) J Biol Inorg Chem 16:183–194
Riester J, Zumft WG, Kroneck PMH (1989) Eur J Biochem 178:751–762
Fujita K, Chan JM, Bollinger JA, Alvarez ML, Dooley DM (2007) J Inorg Biochem 101:1836–1844
Dell’Acqua S, Pauleta SR, Moura JJ, Moura I (2012) Philos Trans R Soc Lond B Biol Sci 367:1204–1212
Pomowski A, Zumft WG, Kroneck PMH, Einsle O (2010) Acta Crystallogr Sect F Cryst Comm 66:1541–1543
Pomowski A, Zumft WG, Kroneck PMH, Einsle O (2011) Nature 477:234–237
Zhang L, Wüst A, Prasser B, Müller C, Einsle O (2019) Proc Natl Acad Sci USA 116:12822–12827
Zhang L, Bill E, Kroneck PMH, Einsle O (2021) Chem Sci 12:3239–3244
Johnston EM, Dell’Acqua S, Pauleta SR, Moura I, Solomon EI (2015) Chem Sci 6:5670–5679
Ghosh S, Gorelsky SI, Chen P, Cabrito I, Moura JJG, Moura I, Solomon EI (2003) J Am Chem Soc 125:15708–15709
Gorelsky SI, Ghosh S, Solomon EI (2006) J Am Chem Soc 128:278–290
Johnston EM, Dell’Acqua S, Ramos S, Pauleta SR, Moura I, Solomon EI (2014) J Am Chem Soc 136:614–617
Smith PK, Krohn RI, Hermanson GT, Mallia AK, Gartner FH, Provenzano MD, Fujimoto EK, Goeke NM, Olson BJ, Klenk DC (1985) Anal Biochem 150:76–85
Kabsch W (2010) Acta Crystallogr Sect D 66:125–132
Evans PR, Murshudov GN (2013) Acta Crystallogr Sect D 69:1204–1214
Winn MD, Ballard CC, Cowtan KD, Dodson EJ, Emsley P, Evans PR, Keegan RM, Krissinel EB, Leslie AGW, McCoy A, McNicholas SJ, Murshudov GN, Pannu NS, Potterton EA, Powell HR, Read RJ, Vagin A, Wilson KS (2011) Acta Crystallogr Sect D 67:235–242
Vagin A, Teplyakov A (2010) Acta Crystallogr Sect D 66:22–25
Emsley P, Lohkamp B, Scott WG, Cowtan K (2010) Acta Crystallogr D Biol Crystallogr 66:486–501
Murshudov GN, Skubak P, Lebedev AA, Pannu NS, Steiner RA, Nicholls RA, Winn MD, Long F, Vagin AA (2011) Acta Crystallogr D Biol Crystallogr 67:355–367
Chen VB, Arendall WB 3rd, Headd JJ, Keedy DA, Immormino RM, Kapral GJ, Murray LW, Richardson JS, Richardson DC (2010) Acta Crystallogr D Biol Crystallogr 66:12–21
Beinert H (1988) Ann N Y Acad Sci 550:374–379
Beinert H (1997) Eur J Biochem 245:521–532
Kroneck PMH, Antholine WA, Riester J, Zumft WG (1989) FEBS Lett 248:212–213
Li PM, Malmström BG, Chan SI (1989) FEBS Lett 248:210–211
Kroneck PMH, Antholine WE, Kastrau DHW, Buse G, Steffens GCM, Zumft WG (1990) FEBS Lett 268:274–276
Farrar JA, Neese F, Lappalainen P, Kroneck PMH, Saraste M, Zumft WG, Thomson AJ (1996) J Am Chem Soc 118:11501–11514
Kroneck PMH (2018) J Biol Inorg Chem 23:27–39
Williams PA, Blackburn NJ, Sanders D, Bellamy H, Stura EA, Fee JA, McRee DE (1999) Nat Struct Biol 6:509–516
Liu J, Chakraborty S, Hosseinzadeh P, Yu Y, Tian SL, Petrik I, Bhagi A, Lu Y (2014) Chem Rev 114:4366–4469
Hwang HJ, Berry SM, Nilges MJ, Lu Y (2005) J Am Chem Soc 127:7274–7275
Karlsson BG, Nordling M, Pascher T, Tsai LC, Sjolin L, Lundberg LG (1991) Protein Eng 4:343–349
Schneider LK, Wüst A, Pomowski A, Zhang L, Einsle O (2014) Metal Ions Life Sci 14:177–210
Zhang L, Bill E, Kroneck PMH, Einsle O (2021) J Am Chem Soc 143:830–838
Tsai ML, Hadt RG, Marshall NM, Wilson TD, Lu Y, Solomon EI (2013) Proc Natl Acad Sci USA 110:14658–14663
Morgada MN, Llases ME, Giannini E, Castro MA, Alzari PM, Murgida DH, Lisa MN, Vila AJ (2020) Chem Commun (Camb) 56:1223–1226
Sasaki S (1989) KEK Report. National Laboratory for High-Energy Physics, Ibaraki, Japan
Schneider LK, Einsle O (2016) Biochemistry 55:1433–1440
Müller C, Zhang L, Zipfel S, Topitsch A, Lutz M, Eckert J, Prasser B, Chami M, Lü W, Du J, Einsle O (2022) Nature 608:626–631
Ghosh S, Gorelsky SI, George SD, Chan JM, Cabrito I, Dooley DM, Moura JJG, Moura I, Solomon EI (2007) J Am Chem Soc 129:3955–3965
Bagherzadeh S, Mankad NP (2018) Chem Commun 54:1097–1100
Johnston EM, Carreira C, Dell’Acqua S, Dey SG, Pauleta SR, Moura I, Solomon EI (2017) J Am Chem Soc 139:4462–4476
Chen P, Gorelsky SI, Ghosh S, Solomon EI (2004) Angew Chem Int Edit 43:4132–4140
Weiss M, Hilgenfeld R (1997) J Appl Crystallogr 30:203–205
Karplus PA, Diederichs K (2012) Science 336:1030–1033
Cruickshank DWJ (1999) Acta Crystallogr Sect D 55:583–601
Laskowski RA, MacArthur MW, Moss DS, Thornton JM (1993) J Appl Crystallogr 26:283–291
Acknowledgements
This work was supported by the Deutsche Forschungsgemeinschaft (RTG 1976, Project No. 235777276, and PP 1927, Project No. 311061829 to O.E.) and the European Research Council (Grant No. 310656 to O.E.). The authors thank Lin Zhang for the helpful discussions. This work was also supported by national funds from Fundação para a Ciência e Tecnologia (FCT) under the scope of the project UIDP/04378/2020 and UIDB/04378/2020 of the Research Unit on Applied Molecular Biosciences—UCIBIO, the project LA/P/0140/2020 of the Associate Laboratory Institute for Health and Bioeconomy—i4HB, and the projects UIDB/50006/2020 and UIDP/50006/2020 of the Laboratório Associado de Química Verde–LAQV. FCT has also supported I.M. through the project 2022.01152.PTDC, and S.D. was supported by an EMBO Short-Term Fellowship (ref. ASTF 282.00-2010).
Funding
Open Access funding enabled and organized by Projekt DEAL. This work was funded by European Molecular Biology Organization, ASTF 282.00-2010, Deutsche Forschungsgemeinschaft, PP 1927, Project No. 311061829, RTG 1976, Project No. 235777276, FP7 Ideas: European Research Council, 310656, Fundação para a Ciência e a Tecnologia, 2022.01152.PTDC, UCIBIO, the project LA/P/0140/2020, UIDB/50006/2020, and UIDP/50006/2020.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pomowski, A., Dell’Acqua, S., Wüst, A. et al. Revisiting the metal sites of nitrous oxide reductase in a low-dose structure from Marinobacter nauticus. J Biol Inorg Chem 29, 279–290 (2024). https://doi.org/10.1007/s00775-024-02056-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-024-02056-y