Abstract
Guanine-rich quadruplex DNA (G-quadruplex) is of interest both in cell biology and nanotechnology. Its biological functions necessitate a G-quadruplex to be stabilized against escape of the monovalent metal cations. The potassium ion (\({{\varvec{K}}}^{+}\)) is particularly important as it experiences a potential energy barrier while it enters and exits the G-quadruplex systems which are normally found in human telomere. In the present work, we analyzed the time it takes for the \({{\varvec{K}}}^{+}\) cations to get in and out of the G-quadruplex. Our time estimate is based on entropic tunneling time—a time formula which gave biologically relevant results for DNA point mutation by proton tunneling. The potential energy barrier experienced by \({{\varvec{K}}}^{+}\) ions is determined from a quantum mechanical simulation study, Schrodinger equation is solved using MATLAB, and the computed eigenfunctions and eigenenergies are used in the entropic tunneling time formula to compute the time delay and charge accumulation rate during the tunneling of \({{\varvec{K}}}^{+}\) in G-quadruplex. The computations have shown that ion tunneling takes picosecond times. In addition, average \({{\varvec{K}}}^{+}\) accumulation rate is found to be in the picoampere range. Our results show that time delay during the \({{\varvec{K}}}^{+}\) ion tunneling is in the ballpark of the conformational transition times in biological systems, and it could be an important parameter for understanding its biological role in human DNA as well as for the possible applications in biotechnology. To our knowledge, for the first time in the literature, time delay during the ion tunneling from and into G-quadruplexes is computed.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Recent times have been witnessing a growing interest in the guanine-rich quadruplex DNAs (G-quadruplexes). They are found in the telomere and oncogene promoter regions of the genome and are involved in various studies in cell biology and nanotechnology.
The functions of G-quadruplexes vary with potentially protective or harmful effects in viral, neurological, and cancer diseases [1]. Their functions are associated with modulator roles in biological processes at each step of the central dogma in cellular life. Previously, it has been shown that G-quadruplexes act as pathogenic drivers linking a diverse range of neurologically important genes, non-coding RNAs, and molecular pathways. Furthermore, G-quadruplexes are enriched in the regulatory regions of microbes including bacteria, fungi, and viruses. In viral diseases, G-quadruplexes are considered as a disease facilitator by modulating translation and providing immune evasion of viruses such as hepatitis B virus (HBV) and hepatitis C virus (HCV), Epstein-Barr virus (EBV), and SARS-CoV-2. A very recent paper has shown that the G-quartet structure represents an attractive approach for potentially inhibiting the SARS-CoV-2 virus [2]. Moreover, G-quadruplexes in microbial genomes have been associated with some unique features such as radioresistance, antigenic variation, recombination, and latency [3]. On the other hand, more than 40% of promoter regions of human genes have at least one G-quartet formation. Different types of proteins can bind, unwind and stabilize G-quadruplexes, where they regulate responsible oncogenes. Thus, imbalance in quartet dynamics is believed to contribute to diseases, including cancer, and, in fact, targeting the G-quartet is suggested for cancer therapy in several publications [4, 5]. As a result, stabilization of G-quadruplexes is gaining priority in pathogenesis of a variety of diseases.
The structural importance of G-quadruplexes is of concern in not only the cell biology but also the nanotechnological applications. For example, it has been suggested that G-quadruplexes can be used as an alternative recognition element in disease-related target sensing [6]. In addition, they are utilized for pore formation, because G-quadruplexes have lipophilic and specific ion-dependent features which allow them to make highly selective artificial transmembrane channels [7]. (See the recent review Ref. [8] as well as Ref. [9] for an in-depth discussion of the quantum treatment of ion channels.) The G-quadruplex channels demonstrate high potential for applications in the fields of molecular diagnostics, logic biocomputing, ionophore construction, and single-molecule biosensing. The structure of G-quadruplex is preferred for nanomachine research because of controlling its reversible folding and extension based on external stimuli such as temperature, pH value, molecular recognition, and light irradiation. In a recent paper, Azobenzene-incorporated lipophilic G-quadruplex channels are used for the selective transport of \({{\varvec{K}}}^{+}\) ions across the lipid membrane [10]. In another study, transporting \({{\varvec{K}}}^{+}\) ion ionophore was constructed from a non-covalent assembly of a G-quadruplex [11]. Besides \({{\varvec{K}}}^{+}\) ion transport, Kaucher et al. have used a strategy that combines non-covalent synthesis and covalent capture to prepare a functional unimolecular G-quadruplex to move \({{\varvec{N}}{\varvec{a}}}^{+}\) across a phospholipid bilayer [12]. On the other hand, molecular targeting of G-quadruplexes is another important approach in nanotechnological applications. Rivera-Sanchez et al. designed self-assembled ligands (SALs) to overcome the challenge of biomolecular surface recognition. They reported that SALs made from 8-aryl-2′-deoxyguanosine derivatives can form precise hydrophilic supramolecular G-quadruplexes (SGQs) with excellent size, shape, and charge complementarity to G-quadruplex DNA [13]. Another research revealed bis-NI1, bis-NI4, and bis-NI6 as possible anticancer therapeutic candidates to target G-quadruplexes on the DNA [14]. The targeting approach for the G-quadruplexes at the specific regions of the genome often is aimed at stabilization of their structure.
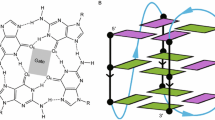
Metal ions are the most important factor for the stabilization of G-quadruplexes. Many species of cations, such as \({{\varvec{K}}}^{+}\), \({{\varvec{N}}{\varvec{a}}}^{+}\), \({{\varvec{N}}{\varvec{H}}4}^{+}\) and the divalent cations (\({{\varvec{S}}{\varvec{r}}}^{++}\), \({{\varvec{Z}}{\varvec{n}}}^{++}\), \({{\varvec{P}}{\varvec{b}}}^{++}\), etc.), are able to stabilize the whole structure of G-quadruplexes. Basically, four guanine bases are bonded together to form a square planar structure in which each of the guanine bases shares two hydrogen bonds. The final form of G-quadruplexes comprises two or more stacked guanine tetrads and binding one type of metal cations that dwell between plenary tetrads (Fig. 1). \({{\varvec{K}}}^{+}\) ion is considered to be the most physiologically relevant cation for G-quadruplexes. The lifetimes of cation intermediates could influence the biological function of G-quadruplexes such as arrest or death of cancer cells through a telomere-related mechanism [15]. Although monovalent cations (especially \({{\varvec{K}}}^{+}\)) have been accepted as a major factor in the stability of G-quadruplex, little is known about ion binding to the G4 motif and ion transition within the molecule. The role of environmental effects of water molecules in \({{\varvec{K}}}^{+}\) binding has not been completely revealed yet but quantum mechanical (QM) simulations help to predict the ion binding.
Structures of G-quadruplex system found in human telomere. A Lines representation of G-quadruplex, view from the side looking into the medium groove, B Hoogsteen hydrogen bonding between four guanines held in a square planar configuration, C cartoon representation of G-quadruplex, D schematic representation of a stable basket-type human telomere G-quadruplex (PDB code: 2KF8) in \({{\varvec{K}}}^{+}\) solution
QM simulations provide a leading-edge tool for quantum biology field. In quantum biology, the term ‘quantum’ does not simply mean quantization of electrons, while discretization of electron energies is used to mention for chemical stability, reactivity, binding, and structure of biomolecules. Although quantum biology is generally defined as a new field, it has a deep-rooted history to the quantum pioneers of the early twentieth century [16]. Nowadays, a number of specific mechanisms within living cells are known to make use of the non-trivial features of quantum mechanics such as long-lived coherence observed in the energy transport in photosynthesis [17, 18], quantum tunneling in the enzyme catalysis [19, 20], olfaction [21], and DNA mutations [22], quantum entanglement in avian navigation [23], and quantum coherence in consciousness [24]. These quantum features were not previously thought to be relevant to complex biological systems because quantum mechanistic features are observed mostly at the level of isolated molecular, atomic and subatomic systems or at temperatures near absolute zero. On the other hand, a microscopic biological system was defined as an open quantum system that must be continuously supplied with energy from its environment. Therefore, it was expected that any delicate quantum effects will very rapidly dissipate resulting in the suppression of any well-controlled quantum dynamics. However, recent studies showed that quantum phenomena would be likely to play a significant role in these biological systems [25, 26]. There are two statements for enabling quantum mechanics in complex biological systems: (i) quantum phenomena can take place over very short time scales before wash away by its surroundings and (ii) quantum dynamics can be enhanced by a finely tuned and constructive interplay with its surroundings [16]. In the last decades, the growing number of theoretical and experimental studies supported quantum tunneling and quantum coherence effects on biology [22, 27, 28], however, other areas of quantum biology remain speculative such as magnetoreception [29] and olfaction [30, 31]. Even though its fast progression, the main limitation of studies in quantum biology is the difficulty of monitoring experimental systems for precise physical measurement. Another limitation is that most simulations rest on zero-temperature calculations due to computational difficulties [8].
The QM approach can provide a more accurate description of the nature of the G-quadruplexes because it can explain the effects of polarization, hydrogen-bonding, and stacking interactions [32]. QM/MM simulations, specifically the DFT-D methods have been used to describe explicitly the electronic structure of a molecule for revealing the details of the structure [33]. First, potential energy surfaces of metal cations for three G-tetrad and one-ion systems have been obtained using QM/MM simulations and DFT-D3 methods by Gkionis et al. 2014. In this paper, the transition of the ion through the quartet plane seems to occur with quantum tunneling. The quantum effects prove relevant even for atoms like \({{\varvec{K}}}^{+}\) because even the heavier 87Rb atom has been observed to obey quantum tunneling [34]. Actually, quantum effects in large-mass atoms/molecules are not restricted to tunneling. They are a general feature, and best revealed by interference experiments. In fact, quantum behavior has been repeatedly observed in large-mass organic molecules [35, 36] much heavier than potassium, and seems to have strong future prospects [37]. For the potassium ion channel under consideration, quantum effects are essential concerning especially the tunneling, charge transfer and other features [8, 9].
The influence of quantum tunneling on \({{\varvec{K}}}^{+}\) ion transition within the G-quadruplex system has been studied in detail in [38] using M06-2X functional for the DFT describing both the hydrogen-bond interaction between the bases and the π–π interactions between the G-quadruplexes. They obtained the potential energy surface in ion transfer in all studied forms of G-quadruplex systems with and without hydration using the polarizable continuum model (PCM) approach.
In view of the growing biological and nanotechnological relevance of the G-quadruplex, and in light of the experimental results and simulation studies, we study in the present work \({{\varvec{K}}}^{+}\) ion transfer through the G-quadruplexes as a tunneling-enabled process. We analyze this process from both a biological and physical point of view. Our end goal is to estimate the duration of the \({{\varvec{K}}}^{+}\) transfer, namely time it takes for the ion to make a tunneling transition. Our estimates are based on the time formulae used for other quantum-biological processes [28] and will be found to give biologically relevant time scales also in \({{\varvec{K}}}^{+}\) transfer in G-quadruplexes.
Quantum tunneling time
In quantum mechanics, time is fundamentally different from position, energy, and momentum. Energy, position, and momentum vary from particle to particle. They are thus observables represented by the corresponding Hermitian operators. Time, on the other hand, does not belong to a particle, instead, it is a common ground for all. There is thus no method to calculate the traversal time in a given region of space. The problem becomes particularly striking when it comes to potential barriers—the regions in which the potential energy of a particle exceeds its total energy. The uncertainty principle governing quantum behavior ensures that quantum particles (electrons, protons, atoms) can traverse potential energy barriers [34]. Due to the unavailability of any first principle calculation method, over the decades various time formulas have been constructed. It proves complementary to emphasize certain time formulae in the literature (though most of them have been sidelined by the tunnel ionization times of noble gases [39]). Dwell time [40], unable to distinguish between transmission and reflection processes, gives the mean interaction duration. Phase times [41,42,43] follow the phase of the wavefunction though the structure of the phase is not clear in the barrier region. Local Larmor time [44] set by spin-precession about \(y\)-direction is not consistent with the dwell time. Local Larmor time [44] (\(z\)-direction) and Büttiker-Landauer time (1983) are in tension with the rule that an incident particle is either transmitted or reflected [45]. This rule is obeyed by the dwell time, as it equals the transmission time multiplied by the transmission probability plus reflection time multiplied by the reflection probability. Lastly, Sokolovski and Baskin (1987) introduced a complex traversal time the meaning of which has been much discussed [45]. These time formulae seem to be in tension with the measurements of the tunnel ionization times of the atoms [39].
In the following, we will focus on two tunneling time formulas: the dwell time [40] and the entropic tunneling time [28, 46].
Dwell time
Smith’s dwell time [40] was defined as the ratio of the number of particles under the barrier to the incoming flux. It gives the average duration of collision [28]. The formula is well known and uncontroversial [47] and proves useful for potential profiles like Fig. 3. It does not distinguish between transmitted and reflected particles [45]. Its definition assumes that the particles are asymptotic states outside the barrier. This does not hold for the \({{\varvec{K}}}^{+}\) ion transition potential in Fig. 2 but we will still include dwell time in our analysis as a reference time formula. It is defined as
Potential energy of the particle. Here, \({\zeta }_{nL}\) and \({\zeta }_{nR}\) are the turning points, and \({\zeta }_{m}\) is the point corresponding to the maximum barrier height \({V}_{max}\). (In Sec. 2, we prefer to use the generic symbols \(E\), \({\zeta }_{L}\) and \({\zeta }_{R}\) by dropping the energy level index \(n\).)
in which the incident particle is taken nearly free to have the momentum \(\hslash k=\sqrt{2{m}_{p}E}\), (assuming that potential energy is small enough). In Eq. (1), \(\psi {\left(\zeta \right)}^{*}\) is the complex conjugate of the wave function \(\psi (\zeta ).\) The particle wavefunction \(\psi (\zeta )\) in Eq. (2) is obtained by solving the Schrodinger equation:
where V \(\left(\zeta \right)\) is the potential energy profile seen by the particle, \({m}_{p}={m}_{{K}^{+}}\) is the potassium ion mass with the molar value of \({m}_{K+}=39.0983 \mathrm{g}/\mathrm{mol}\), and a = 1 Å is the normalization scale of the reaction coordinate. In the next section, we will determine the particle energy \(E\) in the barrier region for which ζL \(\le\) ζ \(\le\) ζR and in which \(E < V(\zeta )\) (see Fig. 2) by solving Eq. (2) numerically.
Entropic time
The tunneling region (\({\zeta }_{L}\le \zeta \le {\zeta }_{R}\) in which \(E<V(\zeta )\)) in Fig. 2 is characterized by two prime facts [45]: (i) particle’s kinetic energy is negative, which means that time flows in imaginary direction under the barrier, and (ii) there is actually no real propagation under the barrier so that wavefunction evanescences from \(\zeta ={\zeta }_{L}\) towards \(\zeta ={\zeta }_{R}\) with exponential suppression. These two facts guide us to a statistical formulation of the tunneling time. The problem is to derive tunneling delay time as a function of particle properties (potential energy). This is achieved by considering probability of finding the particle under barrier in defining the dwell time in Sect. 2. It is a well-known fact that quantum mechanics with imaginary time \(i\tau\) (\(i=\sqrt{-1}\) is the imaginary unit) corresponds to the equilibrium statistical mechanics with temperature \(T=\frac{\hslash }{{k}_{B}\tau }\) (where \({k}_{B}\) is the Boltzmann constant). This can be taken to conclude that tunneling process can be approached from a thermodynamical perspective. Actually, this kind of time model has already been constructed in Ref. [46], with good agreement with experimental data on the He ionization under strong laser fields [39, 48]. Here, we simplify the formalism of Demir and Güner [46].
It proves useful to recall that, in thermodynamics, the energy of the random motions of constituents equals \(TS\), where \(T\) is the temperature and \(S\) is the entropy. This energy is useless in that it does no volume work. This feature remains parallel to the absence of any real propagation in the barrier region. This parallelism can be taken to imply that the energy \(TS\) can actually be the energy to be associated with the tunneling ion. In other words, \(TS\) can be the energy conjugate to time-lapse in the barrier region in the sense of uncertainty principle. With this reasoning, tunneling time can be formulated via the following steps:
-
(A)
As mentioned in item (a) above, if classical equations of motion are continued into the barrier region particle is seen to get from the left-turning point \({\zeta }_{L}\) to the right-turning point \({\zeta }_{R}\) (under the barrier in Fig. 2 or Fig. 3) in a purely imaginary time \(i\tau\) such that
$$\tau \left({\zeta }_{R}, {\zeta }_{L}\right)={\int }_{{\zeta }_{L}}^{{\zeta }_{R}} \frac{{m}_{p}ad\zeta }{\sqrt{2{m}_{p}(V\left(\zeta \right)-E)}}$$(3)is the classical time delay [49]. Needless to say, \(i\tau\) is a purely imaginary time; no clock can read it. It has clearly nothing to do with the tunneling duration as a temporal parameter (though it has the units of time). In fact, as discussed in item (b) above, it sets the effective temperature [46]
$$T=\frac{\hslash }{{k}_{B}\tau \left({\zeta }_{R}, {\zeta }_{L}\right)}$$(4)of the corresponding statistical system. This effective temperature is an indicator of the statistical nature of the particle dynamics under the barrier. This is in accordance with the quantum probabilistic behavior encoded in the probability density [28, 46].
-
B.
Quantum behavior is inherently random. The energy contained in this randomness is \(TS\) and it holds for a single tunneling particle (not an ensemble of particles in the sense of classical thermodynamics). In fact, the von Neumann entropy [50]
$$S=-{k}_{B}P\left({\zeta }_{R},{\zeta }_{L}\right)log P({\zeta }_{R},{\zeta }_{L})$$(5)gives the entropy of a single tunneling particle, where \(P\left({\zeta }_{R},{\zeta }_{L}\right)\) defined as
$$P\left({\zeta }_{R},{\zeta }_{L}\right)={\int }_{{\zeta }_{L}}^{{\zeta }_{R}} d\zeta \psi {\left(\zeta \right)}^{*}\psi (\zeta )$$(6)gives the probability to find the particle under the barrier (between the turning points \({\zeta }_{L}\le \zeta \le {\zeta }_{R}\)). The entropy in Eq. (5) is a measure of the increase in disorder (of the quantum uncertainty) in the course of tunneling [28].
-
(C)
Now, collecting the temperature \(T\) from Eq. (4) and entropy \(S\) from Eq. (5) one can define then the entropic tunneling time [28]
$$\Delta {t}_{E}\left({\zeta }_{R}, {\zeta }_{L}\right)=\frac{\hslash }{TS}=-\frac{\tau \left({\zeta }_{R},{\zeta }_{L}\right)}{P\left({\zeta }_{R},{\zeta }_{L}\right)log P\left({\zeta }_{R},{\zeta }_{L}\right)}$$(7)by interpreting the energy-time uncertainty in quantum theory in terms of the energy \(TS\) of the quantal random motion. Energy-time uncertainty with the entropic thermal energy \(TS\) makes sense because \(TS\) varies with the probability distribution as well as the system parameters. In this sense, saturation of the uncertainty relation \(\Delta {t}_{E}\left({\zeta }_{R},{\zeta }_{L}\right) TS\ge \hslash\) leads to the entropic tunneling time \(\Delta {t}_{E}\left({\zeta }_{R},{\zeta }_{L}\right)\) in Eq. (7).
Numerical analyses of the dwell and entropic tunneling times
Computation of the tunneling time
-
(a)
It is convenient (though not necessary) to work with dimensionless quantities especially when we deal with numerical computations. To this end, one defines
$$\zeta =\frac{\mathrm{reaction coordinate}}{a}$$(8)where the distance “\(a\)” is about the characteristic size of the system. In biological structures, one can take a =1 Å, where 1 Å=10−10 m. The Schrodinger equation in Eq. (2) is already written in terms of this dimensionless coordinate.
-
(b)
The potential energy profile seen by the \({{\varvec{K}}}^{+}\) ion in G-quadruplex is obtained from Villani’s study [38]. We perform a regression analysis on the results and determine the potential energy surface in eV (with the conversion 1 cm−1 = 1.24 10–4 eV). In regression analysis, we prefer to use polynomial regression in modeling the relationship between potential energy (\(V)\) and dimensionless coordinate \((\zeta\)) mainly because of its effectiveness and low computational cost.
The general \(m\)-th order polynomial regression model is
$${V}_{i}={\beta }_{0}+{\beta }_{1}{\zeta }_{i}+{\beta }_{2}{\zeta }_{i}^{2}+\cdots +{\beta }_{m}{\zeta }_{i}^{m} (i=\mathrm{1,2},\cdots ,k)$$(9)where the potential \(V\) is the dependent variable, \(\zeta\) is the independent variable, \({\beta }_{0},{\beta }_{1}\cdots {\beta }_{m}\) are the fit parameters, and \(k\) is the total number of sample points. Equation (9) can be written as a system of linear equations:
$$\underbrace {{\left[ {\begin{array}{*{20}c} {V_{1} } \\ {V_{2} } \\ {\begin{array}{*{20}c} {V_{3} } \\ \vdots \\ {V_{k} } \\ \end{array} } \\ \end{array} } \right]}}_{{\varvec{V}}} = \underbrace {{\left[ {\begin{array}{*{20}c} {\begin{array}{*{20}c} 1 & {\zeta_{1} } & {\zeta_{1}^{2} } \\ 1 & {\zeta_{2} } & {\zeta_{2}^{2} } \\ 1 & {\zeta_{3} } & {\zeta_{3}^{2} } \\ \end{array} } & \cdots & {\begin{array}{*{20}c} {\zeta_{1}^{m} } \\ {\zeta_{2}^{m} } \\ {\zeta_{3}^{m} } \\ \end{array} } \\ \vdots & \ddots & \vdots \\ {\begin{array}{*{20}c} 1 & {\zeta_{k} } & {\zeta_{k}^{2} } \\ \end{array} } & \cdots & {\zeta_{k}^{m} } \\ \end{array} } \right]}}_{{\varvec{X}}}\underbrace {{\left[ {\begin{array}{*{20}c} {\beta_{0} } \\ {\beta_{1} } \\ {\begin{array}{*{20}c} {\beta_{2} } \\ \vdots \\ {\beta_{m} } \\ \end{array} } \\ \end{array} } \right]}}_{{\varvec{\beta}}}$$(10)whose vector–matrix form is
$${\varvec{V}}={\varvec{X}}{\varvec{\beta}}$$(11)With this form, the vector of estimated polynomial regression parameters is obtained using the least squares estimation as
$${\varvec{\beta}}={({{\varvec{X}}}^{{\varvec{T}}}{\varvec{X}})}^{-1}{\varvec{V}}$$(12)where the matrix \({\varvec{X}}\) and the vector \({\varvec{V}}\) are read off from Villani’s simulation plots. In this study, the \({\varvec{\beta}}\) vector is found to have the value
$${\varvec{\beta}}{=\left[0.0143, -0.3479, 0.9498, -0.2640,-0.1535, 0.0756, -0.0083\right]}^{T}$$(13)at the order \(m=6\). As a result, the potential energy profile takes the form
$$\mathrm{V}\left(\zeta \right)=-0.0083{\zeta }^{6}+0.0756{\upzeta }^{5}-0.1535{\zeta }^{4}-0.2640{\zeta }^{3}+0.9498{\zeta }^{2}-0.3479\upzeta +0.0143$$(14) -
(c)
Next, we solve the Schrodinger equation in Eq. (2) numerically in the MATLAB computing environment using the Chebfun toolkit [51]. The MATLAB gives an eigensystem of the form
$$\left({E}_{n}, {\psi }_{n}\right)$$(15)where \(n=\mathrm{1,2},3,\dots\) is an index showing the energy eigenvalue \({E}_{n}\) (measured in \(eV\)) and energy eigenfunction \({\psi }_{n}\) (dimensionless in the dimensionless coordinate \(\zeta\)). (This eigensystem characterizes all confining potentials but for the purpose of analyzing \({{\varvec{K}}}^{+}\) tunneling in G-quadruplex it suffices to keep in mind the potential energy surface in Fig. 2.)
-
(d)
Having determined the energy eigenvalues \({E}_{n}\), we turn to the determination of the classical left-turning point \({\zeta }_{nL}\) and the classical right-turning point \({\zeta }_{nR}\) at the energy level \({E}_{n}\) (See Fig. 2). Basically, \({\zeta =\zeta }_{nL}\) (\({\zeta =\zeta }_{nR}\)) is the point at which a particle (potassium ion) incident from the left (right) scatters and turns back to the region of incidence. In one dimension, turning back requires velocity to vanish at a point and the turning points \({\zeta }_{nL}\) and \({\zeta }_{nR}\) are the points at which kinetic energy vanishes. This means that at these points, potential energy is equal to the total energy. Thus, we solve the algebraic equation:
$$V\left(\zeta \right)={E}_{n}$$(16)which returns \(\zeta ={\zeta }_{nL}\) and \(\zeta ={\zeta }_{nR}\) as the two non-trivial roots corresponding to the energy level \({E}_{n}\). These points are depicted in Fig. 2.
-
(e)
Let us now start with the computation of the Dwell Time. For potassium ion, it takes the following half-numerical form:
$$\Delta {t}_{D}\left({\zeta }_{nL}, {\zeta }_{nR}\right)= \frac{44.904\times {10}^{-15}sec}{\sqrt{{E}_{n}}} {\int }_{{\zeta }_{nL}}^{{\zeta }_{nR}} \mathrm{d\zeta }\psi {\left(\zeta \right)}^{*}{\psi }_{n}(\zeta )$$(17)in which \({E}_{n}\) is purely numerical (measured in \(eV\)).
-
(f)
To compute the Entropic Time, we need to compute first the time \(\tau\) in Eq. (3). Its half-numerical formula takes the form
$$\tau \left({\zeta }_{nR}, {\zeta }_{nL}\right)=44.904\times {10}^{-15}sec {\int }_{{\zeta }_{nL}}^{{\zeta }_{nR}} \frac{d\zeta }{\sqrt{V\left(\zeta \right)-{E}_{n}}}$$(18)where \(V\left(\zeta \right)-{E}_{n}\) is purely numerical (measured in \(eV\)). Having now \(\tau \left({\zeta }_{nR}, {\zeta }_{nL}\right)\) at hand, we can compute the entropic tunneling time as
$$\Delta {t}_{E}\left({\zeta }_{nR}, {\zeta }_{nL}\right)=-\frac{\tau \left({\zeta }_{nR},{\zeta }_{nL}\right)}{P\left({\zeta }_{nR},{\zeta }_{nL}\right)log P\left({\zeta }_{nR},{\zeta }_{nL}\right)}$$(19)where
$$P\left({\zeta }_{nR},{\zeta }_{nL}\right)={\int }_{{\zeta }_{nL}}^{{\zeta }_{nR}} {\psi }_{n}{\left(\zeta \right)}^{*}{\psi }_{n}(\zeta )$$(20)is the probability to find the particle (the \({{\varvec{K}}}^{+}\) ion) in the barrier region.
Computation of the ion entrance rate
With every single \({{\varvec{K}}}^{+}\) tunneling, there occurs a net electric charge increase/decrease in the G-quadruplex. The rate at which this process occurs at the \(n\)-th energy level is given by
in which \(X=D\) for the dwell time, and \(X=E\) for the entropic time. The potassium ions are singly charged and, therefore, net charge is equal to the proton charge. The dwell and entropic times lead to different rates. The quantity \({I}_{X}\), having the units of amperes, indicates at what rate the \({{\varvec{K}}}^{+}\) ions are populating (leaving) the G-quadruplex.
Results and discussion
In this work, we assess time delay with QM descriptions of a potassium (\({{\varvec{K}}}^{+}\)) ion transition within basket-type G-quadruplex which is normally found in the human telomere. We have a potential energy surface (PES) scan for a single ion with hydration in a basket-style G-quadruplex system [38]. The \({{\varvec{K}}}^{+}\) ion faces a barrier while passing through the second quartet and is found in multiple diabatic states during its transition. In Villani’s pioneering work, two kinds of minima are given: the first concerns the curves of potential energy for \({{\varvec{K}}}^{+}\) ion when an explicit molecule of water is present. The second pertains to a case where there is a \({{\varvec{K}}}^{+}\) ion inside the cavity and there is another one outside the cavity. We study only the PES scan of \({{\varvec{K}}}^{+}\) ion + H2O because the one with \({{\varvec{K}}}^{+}\) ion alone has practically no barrier (and hence no time delay to investigate). In this PES scan of \({{\varvec{K}}}^{+}\) ion with hydration, a water molecule is present in the cavity while the \({{\varvec{K}}}^{+}\) ion arrives, and this makes the simulation study of Villani as realistic as possible. To understand the delay time during \({{\varvec{K}}}^{+}\) transition from the cavity of the G-tetrad system to outside the system (2KF8 code, Fig. 1), we have studied the entropic time (in comparison to the dwell time).
Our goal is to determine the time delay occurring in the course of the tunneling of an ion from the deep minimum (in the cavity of the system) to the shallow minimum in Fig. 2 (outside the cavity). Using open-source chebfun toolkit in MATLAB computing environment (licensed to Sabancı University), we have solved the Schrodinger equation. Solution of the Schrodinger equation in Eq. (2) yields the energy eigenvalues and eigenstates plotted in Fig. 3 (energy eigenfunctions and the corresponding eigenvalues are depicted with the same color). Figure 3 shows it manifestly that \({{\varvec{K}}}^{+}\) wavefunction exhibits an evanescent pattern in the barrier region, and this means that there is actually no propagation at all in the barrier region and their wavefunction is described by a purely imaginary momentum. This is a characteristic of the tunneling transition.
Now, we use the energy eigenfunctions and eigenvalues in Fig. 3 to compute the tunneling time delay for \({{\varvec{K}}}^{+}\) ion. In calculations, we use the dwell time in Eq. (1) and the entropic time in Eq. (7). For convenience, we introduce the notion of “barrier height”, which is equal to \({V}_{max}-{E}_{n}\), where \({V}_{max}\) is the maximum of the potential barrier, depicted in Fig. 2. For instance, \({V}_{max}\approx 0.47 \mathrm{eV}\) at \({\zeta }_{m}\approx 1.65\) in Fig. 3 above. Our results are tabulated in Table 1. The first column of the table shows eigenenergies \({E}_{n}\) (\(n=1,\cdots ,5\) as depicted in Fig. 3). The second column corresponds to the barrier height and the third column to the dwell time in attoseconds (1 as = 10–18 s). The fourth column, on the other hand, gives the entropic time in picoseconds (1 ps = 10–12 s). The entropic time is roughly 106 times longer than the dwell time. By its physical definition [44] the dwell time is not expected to give accurate results for the PES in Figs. 2 and 3. But the entropic time is expected to do so. In fact, the entropic time values in Table 1, which remain around tenth of picoseconds, turn out to be in the ballpark of the conformational transition times in biological structures [28, 52]. The entropic time can therefore be a relevant time formula for biological transitions, including the tunneling time delays of \({{\varvec{K}}}^{+}\) ions in G-quadruplex.
The shallow minimum in Figs. 2 and 3 corresponds to the configuration where the potassium ion is outside the G-quadruplex. The question is this: given the tunneling time delays tabulated in Table 1, what is the rate at which the \({{\varvec{K}}}^{+}\) ions enter the G-quadruplex? This we defined in Eq. (21). We compute this rate, the entrance rate, energy level by energy level and tabulate the results in Table 2. Our results show that the \({{\varvec{K}}}^{+}\) ions return to G-quadruplex at a rate tenth of pico-amperes. This finding can be tested with future measurements on G-quadruplex.
Computational predictions and whole-genome resequencing showed that there are over 10,000 potential G-tetrads within the human transcriptome and a number of 736,689 G-tetrad structure within the human genome [53, 54]. These G-tetrads are stabilized through interactions with monovalent cations according to the following order: \({{\varvec{K}}}^{+}\) > \({{\varvec{N}}{\varvec{a}}}^{+}\) > Li, where \({{\varvec{K}}}^{+}\) ion elicits the greatest stabilization [55]. In addition, it has been previously shown that the structure of G-tetrads is a key driver of subcellular localization of several messenger RNAs, thus controlling the regulation of the expressions of particular proteins such as potassium leak channels localized in the cellular membrane [56]. Their unique features of G-tetrads gained attention for the development of ion-dependent smart biomimetic transmembrane channels. It has been presented that transmembrane channels forming from “lipophilic G-quadruplex isomers” are suitable candidates for applications in molecular diagnostics, logic computing, selective separation, and single-molecule biosensing, including ion sensing on membranes or characterization of ion–DNA interactions [7]. Interestingly, \({{\varvec{K}}}^{+}\)-stabilized parallel G-quadruplexes facilitated the highly selective transport of \({{\varvec{K}}}^{+}\) ions [57]. Recently, a successful selective \({{\varvec{K}}}^{+}\) ion transporter also called an “ionophore” was constructed from a non-covalent assembly of a G-quadruplex [11]. Ionophores made of G-quadruplex may have great potential to replace transient potassium channels which are important therapeutic targets for brain disorders such as epilepsy, fragile X syndrome, Alzheimer’s disease, and Parkinson’s disease, normally found in most brain areas [58]. Therefore, it is important to know the physical features of G-quadruplex and ion transition in the complex to design transporters, drug targets, and imaging tools for biotechnological applications. On the other hand, several studies provided compelling evidence that the stability of the G-quadruplex structure is quite important for disease development [59, 60]. In our genome, it has been shown that G-rich strands could form specific G-quadruplex structures in the presence of potassium ions, not only in telomeric regions but also in immunoglobulin switch regions [61], insulin gene, the control region of the retinoblastoma susceptibility gene, the promotor region of the c-Myc gene [62], fragile X syndrome triplet repeats, and muscle-specific protein [63]. It will not be a coincidence that the disruption of the G-quadruplex structure in these important genomic regions results in genes suppressing or activating. For example, a specific G-quadruplex structure formed in the c-Myc gene promoter, a proto-oncogene on the path to cancer, serves as a transcriptional repressor element [64]. According to our study, G-quadruplex stability with \({{\varvec{K}}}^{+}\) ions becomes more important than common knowledge because potassium ions could pass potential energy barriers by quantum tunneling in the picoseconds disrupting the stability of the G-quadruplex structure and activate transcription of the oncogene. This type of quantum–mechanical instability may well play a far more important role in the development of cancer and other diseases than has hitherto been suggested.
In Fig. 3, we give the sample eigenfunctions and eigenvectors of the \({{\varvec{K}}}^{+}\) ion tunneling process derived from the PES scan of \({{\varvec{K}}}^{+}\) ion + H2O. The time delays in ion tunneling through the G-tetrads, as tabulated in Table 1, take attoseconds as the dwell time and picoseconds as the entropic time whereas entrance rate of milliamperes for the dwell time and average current of picoampere for the entropic time, as tabulated in Table 2. Here, we considered the transition of the one \({{\varvec{K}}}^{+}\) ion through the G-tetrads to calculate the average ion current. The entrance rate we discuss here should not be confused with the tunneling current, which arises with a certain potential bias to form a preferred electric charge flow direction [65]. The said tunneling current has the form
in which \(e\) is the proton charge, \(\Delta U\) is the applied voltage difference across the barrier, and \({T}_{{K}^{+}}\) is the tunneling transmission rate of the potassium ion. This electric current arises when a potential difference \(\Delta U\) is applied across the barrier. In contrast to this, the G-quadruplex energetics analyzed in the present work has \(\Delta U=0\), which means that there is no current flow. Nevertheless, single ion tunneling changes the potassium ion number in the quadruplex and we parametrize this change with the potassium ion entrance (or exit) rate \({I}_{ X}\) in Eq. (21) and tabulate its numerical values in Table 2. In certain applications, however, one can apply a certain voltage difference \(\Delta U\) to generate the current in Eq. (22) with the energetics given in Fig. 3. Actually, it turns out that the current in Eq. (22) nears the ion entrance rate in Eq. (21) when \(e\Delta U\approx 4{\pi }^{2}\hslash /\tau\) after considering the entropic tunneling time with \({T}_{{K}^{+}}\approx P\mathrm{log}\frac{1}{P}\), where \(P\) was defined in Eq. (20). (Here, we focus on the entropic tunneling time as it is the actual time we analyze.) Using the explicit expression of the classical time \(\tau\) given in Eq. (3), one obtains the relation \(e\Delta U\approx \sqrt{\frac{32{\pi }^{4}}{m}} \frac{\hslash }{L}{\left(V-{E}_{n}\right)}^\frac{1}{2}\) after assuming a nearly constant potential barrier \(V\) with an effective width \(L\) (as a simple illustration). This solution for the voltage difference \(\Delta U\) ensures that the potassium ion entrance rate in Eq. (21) involves the Nernst potential \(V-{E}_{n}\), if one identifies \({E}_{n}\) with the \({K}^{+}\) reversal potential in the Nernst equation [66, 67]. In the G-quadruplex, Nernst potential of the potassium ion \({K}^{+}\) is the potential difference at which no current flow occurs, which is what is implied by the ion entrance (or exit) rate in Eq. (21).
In the case of ionophore made of G-quadruplex applied in brain areas as a therapeutic, current density calculation becomes important. In addition, ion tunneling in G-quadruplex should be considered during the development of the biotechnological tools with the help of G-tetrads.
Before closing, we should add some words about tunneling time calculations. For the PES in Figs. 2 and 3, the problem is actually not a scattering problem. Namely, the \({{\varvec{K}}}^{+}\) ions are coming in as asymptotically free states, and, therefore, the definition of the dwell time becomes somewhat subtle. The dwell time in our calculations is therefore not expected to return a realistic tunneling delay time but it sets a reference time interval for comparing the entropic tunneling time. It should be noted, therefore, that entropic tunneling time is a viable time formula for a PES such as Figs. 2 and 3, and its values can be taken more seriously. In addition, indeed the entropic tunneling time turns out to be close to the conformational times of biological molecules [52, 66]. This agreement might be taken as a confirmation of the entropic tunneling time formula [28, 46]. In general, a quantum approach to ion channel is justified from different aspects [8, 9]. As was also with tunnel ionization of atoms [46], potential barrier is usually taken static on time scales during which the tunneling process occurs. (Typically analyzed with \(T=0 K\) calculations.) Actually, it is this static nature of the potential barrier that allows for the stationary-state analysis of the tunneling process, and the entropic time is formulated in this stationary-state. In our analysis, potassium transport properties remain constant in time due to the stationary nature of the process. Our goal is to determine the time delay during the potassium ion tunneling from the G-quadruplex in the framework of the Schrodinger dynamics. Our analysis of the tunneling delay time (like tunneling delay time analyses in the literature) pertains to a regime where quantum effects are dominant. One may wonder if potassium transport can be approached by some other method. Actually, tunneling is a quantum process for which DFT-based potentials are more natural. (MD, for instance, is classical in nature [8].) In fact, entropic tunneling time formula we use here was applied before to inter-base and intra-base proton tunnelings in DNA (whose potentials were calculated with DFT), and tunneling delay was found to be in picoseconds range [27, 28], which is seen to agree with recent experimental results [52, 68]. It is in this sense that entropic time remains in the ballpark of conformational transition times in biological systems, and we expect it to be so also for potassium ion tunneling in G-quadruplex. Interestingly enough, according to MD simulations [69] G-quadruplex is unavoidably unstable. In contrast, QM simulations give a realistic description of the potassium tunneling in G-quadruplex because, quoting from Villani [38], “it is more accurate in the treatment of hydrogen-bonding and stacking interactions.” It is in light of this fact [38] and it is with our experience from our earlier studies on proton tunneling in DNA [27, 28] that our analysis of the G-quadruplex is based on quantum simulations [38].
Conclusion
In conclusion, the results of this work can shed light on the results of potential applications of G-quadruplexes in the quantum mechanical regime. Therefore, time delay and average current from ion tunneling were underlined to understand the behavior of the G-tetrads. In the present work, we demonstrate time delay due to ion tunneling for the first time by modeling the tunneling time (the entropic time), computing it for ion transition (solution of the Schrodinger equation and concomitant time calculation), and comparing it with a known time model (the dwell time). Furthermore, application of quantum mechanical methods to the G-quadruplex system is testable, and experimental results could provide further evidence on the validity of the quantum tunneling of the ions in the biological context.
Data availability
The authors confirm that the data supporting the findings of this study are available within the article.
Abbreviations
- G-quadruplex:
-
Guanine-rich quadruplex DNA
- HBV:
-
Hepatitis B virus
- HCV:
-
Hepatitis C virus
- EPV:
-
Epstein–Barr virus
- SAL:
-
Self-assembled ligands
- SGQ:
-
Supramolecular G-Quadruplex
- QM:
-
Quantum mechanics
- PCM:
-
Polarizable continuum model
- PES:
-
Potential energy surface
References
Wang E, Thombre R, Shah Y et al (2021) G-Quadruplexes as pathogenic drivers in neurodegenerative disorders. Nucleic Acids Res 49:4816–4830. https://doi.org/10.1093/NAR/GKAB164
Ji D, Juhas M, Tsang CM et al (2021) Discovery of G-quadruplex-forming sequences in SARS-CoV-2. Brief Bioinform 22:1150–1160. https://doi.org/10.1093/BIB/BBAA114
Saranathan N, Vivekanandan P (2019) G-quadruplexes: more than just a kink in microbial genomes. Trends Microbiol 27:148–163. https://doi.org/10.1016/J.TIM.2018.08.011
Nakanishi C, Seimiya H (2020) G-quadruplex in cancer biology and drug discovery. Biochem Biophys Res Commun 531:45–50. https://doi.org/10.1016/J.BBRC.2020.03.178
Kosiol N, Juranek S, Brossart P et al (2021) G-quadruplexes: a promising target for cancer therapy. Mol Cancer. https://doi.org/10.1186/S12943-021-01328-4
Ida J, Chan SK, Glökler J et al (2019) G-quadruplexes as an alternative recognition element in disease-related target sensing. Molecules 24:1079. https://doi.org/10.3390/MOLECULES24061079
Li C, Chen H, Chen Q et al (2020) Lipophilic G-quadruplex isomers as biomimetic ion channels for conformation-dependent selective transmembrane transport. Anal Chem 92:10169–10176. https://doi.org/10.1021/ACS.ANALCHEM.0C02222/SUPPL_FILE/AC0C02222_SI_001.PDF
Kariev AM, Green ME (2021) Quantum calculations on ion channels: why are they more useful than classical calculations, and for which processes are they essential? Symmetry (Basel) 13:655. https://doi.org/10.3390/sym13040655
Vaziri A, Plenio MB (2010) Quantum coherence in ion channels: resonances, transport and verification. New J Phys 12:085001. https://doi.org/10.1088/1367-2630/12/8/085001
Li C, Chen H, Yang X et al (2021) An ion transport switch based on light-responsive conformation-dependent G-quadruplex transmembrane channels. Chem Commun 57:8214–8217. https://doi.org/10.1039/D1CC03273A
Debnath M, Chakraborty S, Kumar YP et al (2020) Ionophore constructed from non-covalent assembly of a G-quadruplex and liponucleoside transports K+-ion across biological membranes. Nat Commun 111(11):1–12. https://doi.org/10.1038/s41467-019-13834-7
Kaucher MS, Harrell WA, Davis JT (2006) A unimolecular G-quadruplex that functions as a synthetic transmembrane Na + transporter. J Am Chem Soc 128:38–39. https://doi.org/10.1021/ja056888e
del Rivera-Sa MC, García-Arriaga M, Hobley G et al (2017) Small-molecule-based self-assembled ligands for G-quadruplex DNA surface recognition. ACS Omega. https://doi.org/10.1021/acsomega.7b01255
Kulsi G, Agnes A, Samson S et al (2020) Side-chain-dependent binding of bis-naphthalimide self-assembled nanoparticles to G-quadruplex DNA for potential anticancer therapy. ACS Appl Nano Mater 2020:1339–1353. https://doi.org/10.1021/acsanm.9b02187
Bryan TM (2020) G-quadruplexes at telomeres: friend or foe? Molecules. https://doi.org/10.3390/MOLECULES25163686
McFadden J, Al-Khalili J (2018) The origins of quantum biology. Proc R Soc A Math Phys Eng Sci 474:20180674. https://doi.org/10.1098/rspa.2018.0674
Engel GS, Calhoun TR, Read EL et al (2007) Evidence for wavelike energy transfer through quantum coherence in photosynthetic systems. Nature 446(7137):782–786. https://doi.org/10.1038/nature05678
Lee H, Cheng YC, Fleming GR (2007) Coherence dynamics in photosynthesis: protein protection of excitonic coherence. Science (80-) 316:1462–1465. https://doi.org/10.1126/SCIENCE.1142188/SUPPL_FILE/LEEH.SOM.PDF
Kohen A, Cannio R, Bartoluccl S, Klinman JP (1999) Enzyme dynamics and hydrogen tunnelling in a thermophilic alcohol dehydrogenase. Nature 399(6735):496–499. https://doi.org/10.1038/20981
Masgrau L, Roujeinikova A, Johannissen LO et al (2006) Atomic description of an enzyme reaction dominated by proton tunneling. Science (80-) 312:237–241. https://doi.org/10.1126/SCIENCE.1126002/SUPPL_FILE/MASGRAU.SOM.PDF
Turin L (1996) A spectroscopic mechanism for primary olfactory reception. Chem Senses 21:773–791. https://doi.org/10.1093/CHEMSE/21.6.773
Godbeer AD, Al-Khalili JS, Stevenson PD (2015) Modelling proton tunnelling in the adenine-thymine base pair. Phys Chem Chem Phys 17:13034–13044. https://doi.org/10.1039/c5cp00472a
Gauger EM, Rieper E, Morton JJL et al (2011) Sustained quantum coherence and entanglement in the avian compass. Phys Rev Lett 106:040503. https://doi.org/10.1103/PHYSREVLETT.106.040503/FIGURES/4/MEDIUM
Tuszynski J (2020) From quantum chemistry to quantum biology: a path toward consciousness. J Integr Neurosci 19:687. https://doi.org/10.31083/j.jin.2020.04.393
Huelga SF, Plenio MB (2013) Vibrations, quanta and biology. Contemp Phys 54:181–207. https://doi.org/10.1080/00405000.2013.829687
Rivas Á, Huelga SF, Plenio MB (2014) Quantum non-Markovianity: characterization, quantification and detection. Reports Prog Phys 77:094001. https://doi.org/10.1088/0034-4885/77/9/094001
Özçelik E, Akar DE, Zaman S, Demir D (2022) Time delay during intra-base proton tunneling in the guanine base of the single stranded DNA. Prog Biophys Mol Biol 173:4–10. https://doi.org/10.1016/J.PBIOMOLBIO.2022.05.009
Çelebi G, Özçelik E, Vardar E, Demir D (2021) Time delay during the proton tunneling in the base pairs of the DNA double helix. Prog Biophys Mol Biol 167:96–103. https://doi.org/10.1016/J.PBIOMOLBIO.2021.06.001
Ritz T, Adem S, Schulten K (2000) A model for photoreceptor-based magnetoreception in birds. Biophys J 78:707–718. https://doi.org/10.1016/S0006-3495(00)76629-X
Hoehn RD, Nichols DE, Neven H, Kais S (2018) Status of the vibrational theory of olfaction. Front Phys 6:25. https://doi.org/10.3389/FPHY.2018.00025/BIBTEX
Tirandaz A, Taher Ghahramani F, Salari V (2017) Validity examination of the dissipative quantum model of olfaction. Sci Reports 71(7):1–10. https://doi.org/10.1038/s41598-017-04846-8
Gkionis K, Kruse H, Platts JA et al (2014) Ion binding to quadruplex DNA stems. Comparison of MM and QM descriptions reveals sizable polarization effects not included in contemporary simulations. J Chem Theory Comput 10:1326–1340. https://doi.org/10.1021/CT4009969/SUPPL_FILE/CT4009969_SI_001.PDF
Ortiz De Luzuriaga I, Lopez X, Gil A (2021) Learning to model G-quadruplexes: current methods and perspectives. Annu Rev Biophys. https://doi.org/10.1146/annurev-biophys-060320
Ramos R, Spierings D, Racicot I, Steinberg AM (2020) Measurement of the time spent by a tunnelling atom within the barrier region. Nature 583:529–532. https://doi.org/10.1038/s41586-020-2490-7
Gerlich S, Eibenberger S, Tomandl M et al (2011) Quantum interference of large organic molecules. Nat Commun 21(2):1–5. https://doi.org/10.1038/ncomms1263
Juffmann T, Milic A, Müllneritsch M et al (2012) Real-time single-molecule imaging of quantum interference. Nat Nanotechnol 75(7):297–300. https://doi.org/10.1038/nnano.2012.34
Kiałka F, Fein YY, Pedalino S et al (2022) A roadmap for universal high-mass matter-wave interferometry. AVS Quantum Sci 4:020502. https://doi.org/10.1116/5.0080940
Villani G (2018) Quantum mechanical investigation of the G-quadruplex systems of human telomere. ACS Omega 3:9934–9944. https://doi.org/10.1021/acsomega.8b01678
Sainadh US, Xu H, Wang X et al (2019) Attosecond angular streaking and tunnelling time in atomic hydrogen. Nature 568:75–77. https://doi.org/10.1038/s41586-019-1028-3
Smith FT (1960) Lifetime matrix in collision theory. Phys Rev 118:349. https://doi.org/10.1103/PhysRev.118.349
Eisenbud L (1948) The Formal Properties of Nuclear Collisions
Bohm D (1951) Quantum theory. Prentice-Hall, New York
Wigner EP (1955) Lower limit for the energy derivative of the scattering phase shift. Phys Rev 98:145. https://doi.org/10.1103/PhysRev.98.145
Büttiker M (1983) Larmor precession and the traversal time for tunneling. Phys Rev B 27:6178–6188. https://doi.org/10.1103/PhysRevB.27.6178
Hauge EH, Støvneng JA (1989) Tunneling times: a critical review. Rev Mod Phys 61:917. https://doi.org/10.1103/RevModPhys.61.917
Demir D, Güner T (2017) Statistical approach to tunneling time in attosecond experiments. Ann Phys (N Y) 386:291–304. https://doi.org/10.1016/j.aop.2017.09.009
Grace Field (2020) On the status of quantum tunneling times. University of Cambridge
Landsman AS, Weger M, Maurer J et al (2014) Ultrafast resolution of tunneling delay time. Optica 1:343. https://doi.org/10.1364/optica.1.000343
Demir DA, Sargin O (2014) Tunneling in polymer quantization and the quantum zeno effect. Phys Lett A 378:3237–3243. https://doi.org/10.1016/J.PHYSLETA.2014.09.044
von Neumann J (2018) Mathematical foundations of quantum mechanics (John von Neumann ). Princeton University Press, Oxford
Driscoll TA, Hale N, Trefethen LN (2014) Chebfun guide: For Chebfun version 5. 212
Lépine F, Sansone G, Vrakking MJJ (2013) Molecular applications of attosecond laser pulses. Chem Phys Lett 578:1–14. https://doi.org/10.1016/j.cplett.2013.05.045
Tu J, Duan M, Liu W et al (2021) Direct genome-wide identification of G-quadruplex structures by whole-genome resequencing. Nat Commun 121(12):1–9. https://doi.org/10.1038/s41467-021-26312-w
Beaudoin JD, Perreault JP (2010) 5’-UTR G-quadruplex structures acting as translational repressors. Nucleic Acids Res 38:7022–7036. https://doi.org/10.1093/NAR/GKQ557
Guiset Miserachs H, Donghi D, Börner R et al (2016) Distinct differences in metal ion specificity of RNA and DNA G-quadruplexes. J Biol Inorg Chem 21:975–986. https://doi.org/10.1007/S00775-016-1393-4
Maltby CJ, Schofield JPR, Houghton SD et al (2020) A 5′ UTR GGN repeat controls localisation and translation of a potassium leak channel mRNA through G-quadruplex formation. Nucleic Acids Res 48:9822–9839. https://doi.org/10.1093/NAR/GKAA699
Forman SL, Fettinger JC, Pieraccini S et al (2000) Toward artificial ion channels: a lipophilic G-quadruplex. J Am Chem Soc 122:4060–4067. https://doi.org/10.1021/JA9925148/SUPPL_FILE/JA9925148_S.PDF
Noh W, Pak S, Choi G et al (2019) Transient potassium channels: therapeutic targets for brain disorders. Front Cell Neurosci 13:265. https://doi.org/10.3389/FNCEL.2019.00265/BIBTEX
Sun D, Guo K, Rusche JJ, Hurley LH (2005) Facilitation of a structural transition in the polypurine/polypyrimidine tract within the proximal promoter region of the human VEGF gene by the presence of potassium and G-quadruplex-interactive agents. Nucleic Acids Res 33:6070–6080. https://doi.org/10.1093/NAR/GKI917
Sun D, Hurley LH (2009) The importance of negative superhelicity in inducing the formation of G-quadruplex and i-motif structures in the c-Myc promoter: Implications for drug targeting and control of gene expression. J Med Chem 52:2863–2874. https://doi.org/10.1021/JM900055S/SUPPL_FILE/JM900055S_SI_001.PDF
Dalloul Z, Chenuet P, Dalloul I et al (2018) G-quadruplex DNA targeting alters class-switch recombination in B cells and attenuates allergic inflammation. J Allergy Clin Immunol 142:1352–1355. https://doi.org/10.1016/J.JACI.2018.06.011
Siddiqui-Jain A, Grand CL, Bearss DJ, Hurley LH (2002) Direct evidence for a G-quadruplex in a promoter region and its targeting with a small molecule to repress c-MYC transcription. Proc Natl Acad Sci USA 99:11593–11598. https://doi.org/10.1073/PNAS.182256799/SUPPL_FILE/2567SUPPTEXT.HTML
Yafe A, Etzioni S, Weisman-Shomer P, Fry M (2005) Formation and properties of hairpin and tetraplex structures of guanine-rich regulatory sequences of muscle-specific genes. Nucleic Acids Res 33:2887. https://doi.org/10.1093/NAR/GKI606
Ambrus A, Chen D, Dai J et al (2005) Solution structure of the biologically relevant G-quadruplex element in the human c-MYC promoter. Implications for G-quadruplex stabilization. Biochemistry 44:2048–2058. https://doi.org/10.1021/BI048242P
Qaswal AB (2019) Quantum tunneling of ions through the closed voltage-gated channels of the biological membrane: a mathematical model and implications. Quantum Reports 1:219–225. https://doi.org/10.3390/QUANTUM1020019
Alberts B, Johnson A, Lewis J et al (2002) Molecular biology of the cell - NCBI bookshelf. Garland Science, New York
Dayan P, Abbott LF (2001) Theoretical neuroscience computational and mathematical modeling of neural systems. The MIT Press, Cambridge
Muntean CM, Dina NE, Bratu I et al (2021) Effects of femtosecond UV laser pulses on the structure and surface dynamics of medicinal plants DNA, monitored by surface-enhanced Raman spectroscopy. J Mol Struct 1239:130482. https://doi.org/10.1016/J.MOLSTRUC.2021.130482
Wang Z, Liu JP (2017) Effects of the central potassium ions on the G-quadruplex and stabilizer binding. J Mol Graph Model 72:168–177. https://doi.org/10.1016/J.JMGM.2017.01.006
Acknowledgements
This work is supported by the IPS Project B.A.CF-20-02239 at Sabancı University. Graphical abstract and Fig. 1 were created with BioRender.com.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Celebi Torabfam, G., K. Demir, G. & Demir, D. Quantum tunneling time delay investigation of \({{\varvec{K}}}^{+}\) ion in human telomeric G-quadruplex systems. J Biol Inorg Chem 28, 213–224 (2023). https://doi.org/10.1007/s00775-022-01982-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-022-01982-z