Abstract
Native mass spectrometry is a potent technique to study and characterize biomacromolecules in their native state. Here, we have applied this method to explore the solution chemistry of human carbonic anhydrase I (hCA I) and its interactions with four different inhibitors, namely three sulfonamide inhibitors (AAZ, MZA, SLC-0111) and the dithiocarbamate derivative of morpholine (DTC). Through high-resolution ESI-Q-TOF measurements, the native state of hCA I and the binding of the above inhibitors were characterized in the molecular detail. Native mass spectrometry was also exploited to assess the direct competition in solution among the various inhibitors in relation to their affinity constants. Additional studies were conducted on the interaction of hCA I with the metallodrug auranofin, under various solution and instrumental conditions. Auranofin is a selective reagent for solvent-accessible free cysteine residues, and its reactivity was analyzed also in the presence of CA inhibitors. Overall, our investigation reveals that native mass spectrometry represents an excellent tool to characterize the solution behavior of carbonic anhydrase.
Graphic abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Carbonic anhydrase (CA, EC 4.2.1.1) is a zinc metalloenzyme that catalyzes the reversible hydration of carbon dioxide to bicarbonate according to the following equation: CO2 + H2O ⇆ HCO3− + H+ [1, 2]. The HCO3−/CO2 equilibrium is critical for human health, and its inhibition has been a goal of therapeutic intervention for several decades [3,4,5,6]. The majority of known CA inhibitors contain a primary sulfonamide group [7, 8]. The sulfonamide anion, RSO2NH−, coordinates to the active-site zinc and may form hydrogen bonds with active-site amino acid residues located in the immediate vicinity further stabilizing the enzyme/inhibitor complex [9].
Owing to the importance and variety of therapeutic applications, the search for inhibitors of carbonic anhydrase is still very intensely pursued. This search is greatly assisted by a precise knowledge of the binding mode of the inhibitors to the enzyme at the atomic level. To this end, a large number of biophysical methods including NMR, X-ray crystallography, surface plasmon resonance (SPR), etc. have been exploited during the last few decades. A detailed description of these methods and of the information that can be derived may be found in a few reviews [10,11,12].
Among the various biophysical techniques, mass spectrometry turned out to be a potent method to describe drug–target interactions [13,14,15,16]. Specifically, native mass spectrometry is a powerful technique to study and characterize biomolecules in their native state, as the gentle ionization method preserves in the gas phase the supramolecular interactions, the conformational features, and the non-covalent association with ligands that are present in solution. In native MS, a strict control of pH, temperature, presence of non-denaturing co-solvents, and of the instrumental parameters is required to guarantee not only the retention of the tertiary structure of the biomolecule, but also the biological function, in the case of an enzyme the catalytic activity [17, 18]. Native MS started in the 90s’ with the pioneering works of Ganem and Katta, who first independently demonstrated that non-covalent interactions between biomolecules can be preserved and transferred from solution to the gas phase, allowing their detection via ESI–MS. Ganem et al. successfully detected via ESI–MS the complex between the receptor FKBP and its ligand, the macrolide FK506 [19]. Katta et al. reported that the non-covalent heme–globin complex of myoglobin is preserved in the gas phase generated via electrospray [20]. Since then, ESI–MS has been recognized as an election tool to investigate and characterize many ligand–biomolecule interactions: protein–cofactors, protein–DNA, protein–metal–drug, enzyme–substrate, enzyme–inhibitors, and antigen–antibody, and a plethora of papers and reviews has been published about this issue [16, 21, 22].
Native-MS has many analytical advantages. The identification of the fragment bound to the biomolecule and the binding stoichiometry can be directly inferred simply from the inspection of the mass spectrum. In fact, once detected the target biomolecule signal, any shift toward greater mass values is a sign of the binding with a ligand whose mass is equal to the mass shift detected. Moreover, the high sensitivity of mass spectrometry requires just a very small sample quantity for analysis, a few micrograms compared to the larger quantities required by other methods, such as NMR and crystallography. These significant features make native-MS a compelling screening method for the fragment-based drug discovery (FBDD) allowing the identification of chemotypes that bind to a protein, even through weak interactions [13]. In a recent work of Woods et al., native MS has been successfully applied in a fragment screening analysis toward CA II, to disclose new potential inhibitors of the enzyme [23]. Native MS has been successfully proven to be a valid alternative to the traditional screening methods, such as SPR and X-ray crystallography, offering unique advantages over them, as no sample manipulation and a very small sample concentration are required.
Another significant biological application of native-MS concerns the structural investigation of proteins. Certainly, since the pioneering work of Chowdhury et al. in 1990, where the conformational changes of Cytochrome c have been monitored for the first time by ESI–MS, the ability of the native-MS to probe and characterize the conformational state of proteins is well known [24].
Indeed, during the soft ionization process occurring in the ESI source, the protein can take multiple charges in accordance with how many protonable (or deprotonable) residues it exposes to the source, giving rise to multicharged species’ signals in the mass/charge spectrum. This signals ensemble represents the charge state distribution (CSD) of the protein. The average charge state that a protein takes on depends on its tertiary structure and its solvent-accessible surface area: the more the residues are buried in the structure, fewer charges the protein can take [25, 26]. Among the many factors that influence the CDS (i.e., solvent, parameters of the instrument, etc.), the protein conformation is the most important [27, 28]. Indeed, it is widely documented that the unfolding of a protein in denaturing conditions causes the shift of its CDS toward higher charges (low m/z values) due to a greater accessibility of basic or acidic residues that can accommodate charges [28, 29]. In a study of Nabuchi et al., the unfolding and refolding processes of CA triggered via pH modulation were monitored by ESI mass spectrometry. These authors followed the conformational changes through the monitoring of the mass shift associated with zinc release in the unfolded state, and through the variation of the charge-state distribution of the protein signal [30].
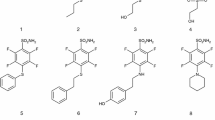
In this manuscript, we exploit native mass spectrometry to characterize the solution chemistry of hCA I and investigate its interactions with a few selected inhibitors. Specifically, four distinct inhibitors were chosen (Scheme 1) of which three are sulfonamide inhibitors, namely acetazolamide (AAZ), methazolamide (MZA), and SLC-0111, and the fourth one is a dithiocarbamate (potassium morpholine-4-carbodithioate, DTC). All these four inhibitors are known to produce their effect through direct binding to the Zn(II) ion, as the X-ray crystal structures of their adducts with various CAs have been reported [31,32,33,34]. AAZ and MZA are the first-generation CAIs used as systemic drugs for the management of glaucoma and as standard CAIs in many pharmacological investigations [5]. SLC-0111 is the first-in-class selective CA IX/XII inhibitor progressing to clinical trials [32]. It successfully completed and passed Phase I, and entered in Phase Ib/II clinical trials in 2017 for the treatment of advanced, metastatic solid tumors [35]. DTC is a main representative of dithiocarbamates, a class of potent zinc-binder CAIs, second only to sulfonamide-like derivatives [31]. In Table 1, the hCA I inhibition constants of the four inhibitors are reported. In addition, further studies are conducted concerning the reaction of hCA and its inhibitor complexes with the gold(I) drug auranofin that is known to bind selectively the free cysteine residue (Cys212). Overall, a quite detailed and satisfactory description of the occurring processes is achieved.
Materials and methods
Materials
Lyophilized human carbonic anhydrase (hCA I) was purchased from Sigma-Aldrich and used without further purification or manipulation. Sulfonamide inhibitors of hCA I were synthetized by one of our groups [31, 32] or are commercially available from Sigma-Aldrich (Milan, Italy). Auranofin (AF) was purchased from Enzo Life Sciences (Farmingdale, New York). Water, methanol, and ammonium acetate were of LC–MS grade and were purchased from Sigma-Aldrich.
Sample preparation
The stock solution of hCA I 10–4 M was prepared dissolving the protein in H2O LC–MS grade. Stock solutions (10–2 M) of the inhibitors were prepared dissolving the samples in DMSO. AF was freshly prepared in LC–MS grade water and methanol (50:50 v/v) to a final concentration of 3 × 10–3 M.
For the ESI–MS experiments with hCA I, aliquots of the protein stock solution were diluted to 7 × 10–7 M with ammonium acetate solution 2 × 10–3 M, pH 6.8.
For the experiments with the inhibitors, solutions of hCA I 10–5 M and each inhibitor at fixed protein-to-inhibitor ratio (1:1, 1:3, 1:10) were prepared diluting with ammonium acetate buffer 2 × 10–3 M, pH 6.8. The mixtures were then incubated at 37 °C for 5 min.
For the experiment with AF, a solution of hCA I 10–5 M and AF at 1:3 protein-to-metal complex ratio was prepared and diluted with ammonium acetate solution 2 × 10–3 M, pH 6.8. The solutions were then incubated at 37 °C for 2 h and then diluted to a final protein concentration of 7 × 10–7 M using ammonium acetate solution 2 × 10–3 M, pH 6.8.
ESI–MS analysis: final dilutions
After the incubation time, all solutions were sampled and diluted to a final protein concentration of 7 × 10–7 M using ammonium acetate solution 2 × 10–3 M, pH 6.8
In the non-native-like experiments, the final solutions were also added with 0.1% v/v of formic acid just before the infusion in the mass spectrometer.
Instrumental parameters
The ESI mass study was performed using a TripleTOF® 5600+ high-resolution mass spectrometer (Sciex, Framingham, MA, USA), equipped with a DuoSpray® interface operating with an ESI probe. Respective ESI mass spectra were acquired through a direct infusion at 5 μL/min flow rate.
The general ESI source parameters optimized for hCA I analysis were as follows:
-
Positive polarity, Ionspray Voltage Floating 5500 V, Temperature 0, Ion source Gas 1 (GS1) 40 L/min; Ion source Gas 2 (GS2) 0; Curtain Gas (CUR) 10 L/min, and Collision Energy (CE) 10 V.
-
Selective variations of some parameters were applied for DP value adjustment: for native hCA I and milder DP, a value of 200 V was used and 2500–5000 m/z mass range; for native hCA I and harder DP, a value of 300 V was used and 2500–5000 m/z mass range; for denatured hCA I positive polarity, Ionspray Voltage Floating 5500 V, Temperature 0, Ion source Gas 1 (GS1) 50 L/min; Ion source Gas 2 (GS2) 0; Curtain Gas (CUR) 20 L/min, Declustering Potential (DP) 50 V, Collision Energy (CE) 10 V, range 760–990 m/z.
-
For acquisition, Analyst TF software 1.7.1 (Sciex) was used and deconvoluted spectra were obtained by using the Bio Tool Kit micro-application v.2.2 embedded in PeakView™ software v.2.2 (Sciex).
Results and discussion
Native-MS analysis of hCA I
Before investigating the protein-binding properties of the selected inhibitors, it was necessary to assess the best conditions for the ESI–MS experiment to observe the protein in its native-like state. In accordance with the definition of “native mass spectrometry” proposed by Heck [17], we aimed to preserve the protein tertiary structure and the binding of the Zn ion in the enzyme active site, as most inhibitors like sulfonamides and dithiocarbamates directly bind this metal ion [36]. The Zn ion is located in a cone-shaped pocket and is coordinated to three His residues and a water molecule (or OH− ion) in a roughly tetrahedral geometry. The His residues are invariant in the whole α-CA family: they are His94, His96, and His119. Other neighboring residues complete the coordination shell establishing hydrogen bonds [37].
In a recent work of ours, CA I was investigated in depth through an established protocol of protein ESI–MS analysis [38]. This methodology includes the direct injection into the ESI-Q-TOF mass spectrometer of a protein sample dissolved in ammonium-acetate solution (see “Sample preparation”). Normally, a small percentage (0.1% v/v) of formic acid is added just before the injection into the mass spectrometer, to enhance the ionization process in the ESI source. Unfortunately, under these experimental conditions, the ESI–MS spectrum shown in Fig. 1 reveals that only apo-CA I is detectable. Notably, the signal at 28,780 Da is assigned to the apo protein and corresponds to the molecular weight of the hCA I amino acid sequence (Uniprot P00915) with the loss of Met1 and the presence of one acetylation on Ala2 residue.
The addition of 0.1% v/v of formic acid to the ammonium acetate solution induces a lowering of the pH from 6.8 to 2.9 with the rapid release of the zinc ion from the protein. Indeed, the His residues that coordinate the metal ion in the active site of the enzyme possess a pKa < 6, and in an acidic solution, they may undergo facile protonation, causing the loss of the zinc ion [39,40,41,42]. As a confirmation, the study of Coleman demonstrated, through optical rotatory dispersion, that hCA B (the old name for hCA I) loses irreversibly its native conformation under pH 4, with the consequent release of the Zn ion and the loss of the catalytic activity [43].
Most of the proteins analyzed in our previous experience with ESI–MS retained their native conformation under physiological-like conditions (i.e., with ammonium acetate solution, ammonium hydrogen carbonate, or water with the addition of 0.1% v/v of formic acid for the mass spectrometry analysis), as proved by our extensive works published during the last years [38, 44,45,46,48,49]. However, the present case can be extremely instructive given the particular sensitivity of hCA I for acidic conditions. It is noteworthy to remind that the so-called “native conditions” for mass spectrometry analysis of proteins in their biologically active conformation cannot be generalized in a standard method, but rather carefully set for each protein [17, 50, 51].
For this reason, the first experiments were devoted to the identification of the best experimental conditions to avoid protein unfolding and denaturation, besides the search of the best ionization performances. First of all, in the case of hCA I, the absence of acid in the ammonium acetate solution is one of the key point for ESI–MS experiments in native-like conditions. Moreover, in this case, also an adequate ionization is reached, leading to a well-resolved spectrum of holo-hCA I displayed in Fig. 2. The main signal at 28,843 Da is attributable to holo hCA I with the Zn(II) ion retained in its coordination site. Besides, signals of protein adducts with sodium (+ 23 Da) and oxidized forms of the protein (+ 16 Da) are also observed.
Therefore, non-denaturing conditions preserve the zinc binding to the active site of the enzyme, while the acidification induces the release of the metal center [52, 53].
In this regard, by comparison between the multicharged spectra of hCA I obtained in acidic and neutral conditions, some interesting observations can be proposed about the different protein conformation.
Since folded protein molecules can accommodate fewer charges in comparison to the unfolded counterpart, the analysis of the charge-state distribution (CSD) in the multicharged spectra can give some clear information on the conformational state of the protein [28]. A drastic variation of CDS in the spectrum of CA, has been highlighted among both experimental conditions. Reasonably, the acidic conditions also cause a considerable alteration of the protein tertiary structure, giving rise to a partial unfolding and, thus, to a higher degree of protonation in the ESI source [54,55,56]. As a result, with the addition of formic acid, the CDS becomes broader and shifted to highly charged ions, with a maximum at + 35 protonation state, as depicted in Fig. 3.
Contrariwise, the spectrum in Fig. 4 has been performed on the CA solution in non-acidic conditions and shows a narrower CDS with a maximum charge state of + 10. Therefore, it is evident that the acidification induces a partial loss of the tertiary structure of the protein, exposing more amino acid residues to the solvent and then leading to a larger protonation in the electrospray compared to the folded state [27].
Another important consideration that may help to better characterize the protein and its interactions via mass spectrometry is related to the greater instrumental resolution achievable when dealing with lower CDS spectra. In fact, a lower charge state corresponds to a greater separation (being ∆m = 1/z) between consecutive signals in the multicharged isotopic cluster increasing, in case of a Tof instrument, the FWHM resolution [57,58,59]. As a result, the mass spectrum of CA in native-like conditions reported in Fig. 2 is more isotopically resolved and informative with respect to the spectrum in Fig. 1 recorded for a solution of CA under slightly acidic conditions (0.1% v/v formic acid).
The binding of the inhibitors
As reported in the literature, sulfonamide and dithiocarbamate inhibitors bind directly to the Zn ion of the CA [36]. Through high-resolution ESI–MS analysis, we succeeded in providing a clear demonstration of such a binding. Indeed, the interaction between hCA and a small panel of its inhibitors (Scheme 1) can be investigated in detail and the adducts formed can be easily observed just in native conditions, in which the Zn ion binding to the three histidine residues in the active site of the enzyme is preserved.
First, we investigated the adduct formation of CA with the inhibitors in native-like conditions, varying properly the instrumental parameters to optimize the adduct signal.
Different aliquots of a 10–5 M protein solution were prepared in the presence of the selected inhibitor (at 1:10 protein-to-inhibitor ratio) in ammonium acetate solution. Each mixture was incubated at 37 °C for five minutes and then diluted to the final concentration with the same ammonium acetate solution (pH 6.8). The ESI–MS spectra were then acquired, and adduct formation was assessed for each of the four inhibitors. From the literature, it is known that the sulfonamide inhibitors bind the Zn ion through their R-SO2-NH− moiety [9, 60, 61], while the dithiocarbamate through the R2NCS2− moiety [31, 62]. For example, Fig. 5 reports the deconvoluted mass spectrum of MZA incubated with hCA I.
Notably, the unbound protein signal at 28,843 Da is no longer observed, indicating that CA reacts completely with the inhibitor. The main signal detected at 29,080 Da perfectly matches the mass of the MZA/CA adduct formed through the binding of the R-SO2-NH− moiety to the Zn ion. Moreover, a signal corresponding to a bis adduct is also detected at 29,316 Da with a relative intensity of about 20% respect to the main peak. The formation of bis adducts between sulfonamide inhibitors and CA, although apparently surprising, is in perfect agreement with some NMR and MS studies already reported in the literature [63, 64]. In fact, as described by Whitesides and co-workers, CA retains the tertiary structure of its binding pocket in the gas phase on the time scale (seconds to minutes) of the ESI–MS measurements [63]. Therefore, although the binding stoichiometry between CA and its inhibitors was 1:1 in solution, these authors postulated that the second equivalent of inhibitor was likely condensed on the surface of the hCA I (and then at non-Zn(II) site) during the ESI desolvation process [63].
By comparing the multicharged spectrum of unreacted CA (Fig. 4) and that with a bound inhibitor, some conformational considerations can be proposed. As an example, in Fig. 6, the charge-state distribution for the CA/MZA adduct is reported. Interestingly, no relevant changes in the CDS can be observed in comparison to the unreacted protein. Indeed, the most abundant charge state shifts from + 10 to + 8, and no broadening of CDS can be observed. This experimental evidence leads us to reasonably exclude any significant variation of the overall protein conformation due to inhibitor binding [55]. Probably, the small variation in the CDS can be truly attributed to a slight and localized conformational variation of the enzyme binding pocket following the inhibitor binding. Likewise, the insertion of the inhibitor molecule inside the enzymatic pocket may be of hindrance to the amino acid protonation of pocket itself.
Nevertheless, when formic acid was later added to the mixture of the inhibitor and CA, the protein lost the Zn ion and, as expected, the inhibitor too. Indeed, in these slightly acidic conditions, only the apo-CA signal was still detectable, with the total disappearance of adduct-related peaks. To monitor this loss, we first prepared the solution of the enzyme with inhibitor in “native-like” condition as previously determined, and then we added the 0.1% v/v of formic acid just before the injection in the mass spectrometer. As a proof of concept, in the Supporting Information, the mass spectrum of CA reacted with another sulfonamide inhibitor, i.e., SLC-0111, is reported: the only signal detected at 28,780 Da refers to the apo-CA peak. Again, when the acid was added to the protein solution, the overall resolution rapidly dropped down and the relative CDS was at the higher values, confirming the pH-related behavior discussed in the previous paragraph.
Therefore, as we assessed from the comparison of the charge-state distribution of unreacted CA in both native and acidified condition, the acidification probably induces a partial loss of the protein tertiary structure making the protonation of the His residues in the active site possible and causing the release of the Zn(II) ion from the enzyme’s pocket [40, 65, 66]. On the contrary, the adduct between the enzyme and the inhibitor is clearly detectable in the absence of any acid addition, when the protein preserves its folded state, as clearly demonstrated from the CDS analysis.
Another important equilibrium that takes place upon varying the pH conditions is the protonation and deprotonation of the sulfonamide and dithiocarbamate reactive moieties. It has already been reported in the literature that the inhibitor binding was suppressed at low pH values (< 2.5), presumably due to neutralization of the negative charge on the zinc-reactive moiety [67].
Definitely, the ESI–MS data obtained for the hCA/inhibitor system perfectly agree with the knowledge on the CA behavior already gathered with other complementary techniques (i.e. XRD), pointing out that the inhibitor binds directly to the Zn ion, when the enzyme is in its native state with the Zn ion tightly bound to its active site [62].
The declustering potential modulation: comparison between the Zn binding to the enzyme and the inhibitor binding to the Zn
Another experimental parameter that deeply influences the obtained results in native ESI–MS analysis of proteins, and particularly metalloproteins, is the declustering potential (DP) [68, 69]. Briefly, the DP is an electric potential difference applied between the orifice and the following lens of the mass spectrometer (in the region of rapid gas expansion from atmosphere into vacuum) [18]. This potential difference establishes an electric field accelerating the ions through the low-density gas and is commonly adjusted to provide the optimum signal-to-noise ratio for the compound of interest. In particular, protonated solvent clusters are collisionally stripped from clustered protein as the potential is increased. Again, low charge-state ions can be more effectively declustered, while high charge-state ions are normally fragmented. Since we demonstrated that the CDS is closely related to the solution pH, this latter parameter is not only responsible for the native conformation retention, but it also plays a pivotal role in the efficacy of the DP. Then, the control of DP is another fundamental instrumental parameter that can improve the instrumental resolution and, therefore, deserves to be carefully evaluated.
Some papers show that the DP can be modulated during the MS analysis, to probe the nature of protein–ligand association. The raising of DP can induce the dissociation of the weaker interactions between a protein and a ligand, while the covalent ones are retained [70, 71].
On the other hand, a high DP value can induce, yet, the dissociation of the weaker ligand–protein adducts [72]. Therefore, we started the systematic study of the interactions between hCA I and its inhibitors, once fixed the best pH conditions, varying the DP value until any significant variation in the multicharged spectrum (in terms of signals intensities, appearance/disappearance of signals at greater masses, resolution, and overall quality of the spectrum) is observed.
First, a solution of CA alone has been prepared and analyzed, as previously described, avoiding any acid addition. At low DP values (i.e., 100 V), as reported in Supporting Information, the signal of the holo-protein is followed by many signals of adducts with salts from the solution, typically observed in protein’s MS analysis, which considerably complicates the mass spectrum. Specifically, the very intense peak at 28,904 Da (shift = + 60 Da with respect to hCA signal) can be reasonably attributed to the addition of an acetate ion from the buffer solution, while the second very intense peak at 29,100 Da has not yet been clearly assigned but probably dues to some ionic clusters from the buffer solution that weakly interact with the protein surface.
By raising the DP value up to 300 V (see Fig. 2), those adducts with salts are destroyed and the mass spectrum becomes cleaner with the protein signal that emerges as the main one. Significantly, even with this high voltage applied, in this case, the Zn ion is retained on its binding site and no significant protein unfolding occurs. A clear and well-resolved holo-protein signal is then observed.
However, when the ESI–MS experiments have been performed with the CA and inhibitor mixed together in these latter conditions, completely different results have been gathered. Indeed, at 300 V, we observed a drastic reduction of intensity of the adducts signals. In the Supporting Information, are displayed the deconvoluted mass spectra for each of the four inhibitors. Notably, in all cases, the main signal is now the one of the unreacted holo-proteins at 28,843 Da. The intensity of the adduct signals drastically collapses at about 30–50% intensity compared to the unreacted protein signal. Interestingly, bis adduct signals are no longer detected. At the moment, we cannot provide a plausible explanation and this aspect will be the object of further investigations.
Then, the DP value was decreased to the optimal value of 200 V and the recorded spectra are shown in Fig. 7. Now the peak corresponding to the unreacted protein is no longer detected in case of the three sulfonamides, but it is still present in case of DTC. Again, for the sulfonamide inhibitors, the main signal detected corresponds to the mono adducts with CA and further bis adducts are also revealed with a relative abundance of 30–40% respect to the first ones. A different behavior is observed for DTC; in this case, in addition to the partial adduct formation with CA, there is no trace about the bis adduct formation.
Thereafter, from the comparative analysis of the spectra obtained with different DP values, we can assert that 200 V can be considered the best value for ESI–MS analysis of CA/inhibitors adducts in native-like conditions, allowing to gather highly informative spectra with a clear and precise rendering of the protein reactivity towards those studied inhibitors.
Clearly, upon increasing the value of the applied voltage, the coordination bond between CA and the inhibitors is no longer preserved. Therefore, the observed residual mono-adduct signal at DP 300 V is due to the presence of a large excess of the inhibitor. Indeed, repeating the analysis on a solution with a 1:1 protein-to-inhibitor ratio, no adduct signal is observable for MZA, AAZ, and DTC, and only the protein signal is detected (see SI). The residual SLC-0111/CA adduct is probably due to the larger number of hydrogen bonds formed with other amino acid residues in the enzymatic pocket respect to the other inhibitors, making this adduct more resistant also to the higher DP values [9, 73].
Notably, although the DP set to the maximum value possible with our instrument (i.e., 300 V), it is clear that the protein preserves the Zn(II) ion into the active site, despite the high energy involved. Indeed, the Zn ion is tightly bound to the enzyme through various direct and indirect interactions with amino acid residues that stabilize the metal ion into the enzyme’s pocket [66].
Differently, the high DP values cause the fragmentation of the CA/inhibitors adducts with the loss of their respective signals. In this case, the kinetic energy applied to the system results too high to keep intact the inhibitors bound to the protein. Since the main signal in all the spectra at 300 V belongs to the holo-CA (see SI), this is suggestive for a different binding energy between Zn-protein and inhibitor-Zn adduct, resulting this latter one less stable with higher acceleration energy [74,75,76]. This is perfectly consistent with the great stabilization that the Zn ion receives from the coordination to the three histidine residues, while the inhibitor establishes only one bond with the metal ion [76].
Competition between inhibitors
For a given inhibitor, the inhibition constant KI represents the inhibitor concentration required to decrease the catalytic activity of the enzyme by 50%. Thus, KI provides an estimate of the inhibitor’s affinity for the enzyme: more tightly the inhibitor binds to the enzyme active site, the smaller the amount of inhibitor required for inhibition [77].
To test the validity of this concept also with mass spectrometry, we carried out a competition experiment between two inhibitors with different KI. Specifically, the two sulfonamides AAZ and MZA were chosen, whose inhibition constants are KI 250 nM and KI 50 nM, respectively [78, 79]. Therefore, AAZ is our weaker inhibitor, while MZA is the stronger one. The protein was mixed with the two sulfonamide inhibitors in a 1:10:10 protein/inhibitor 1/inhibitor 2 ratio. We started the protein incubation with the less tight-binding inhibitor (AAZ), then we added the stronger one (MZA). After a further 5 min of incubation at 37 °C, the spectrum is acquired. Later, we also carried out the same experiment by reversing the order of inhibitor addition.
In the first experiment, AAZ is incubated with hCA I. The ESI mass spectrum (see SI) displays the intense signal of the CA/AAZ adduct. After the addition of MZA, the CA/AAZ signal was no longer detectable. In its place, the peak of the CA/MZA adduct appeared very clearly (Fig. 8). A less intense peak attributable to the holo-hCA I adduct with both inhibitors was also revealed. Also, this behavior can be reasonably explained as described above, considering the particular reaction in the gas phase [63]. As expected, the stronger inhibitor causes the displacement of the weaker one, binding in turn to the protein.
In the opposite experiment, MZA was first incubated with hCA I, and then, AAZ was added. Both spectra showed only the signals due to the holo-CA and to the CA/MZA adduct (see SI), reflecting the tendency of MZA to preferentially bind the protein compared to the less strong inhibitor AAZ. Notably, no adduct with both inhibitors has been observed in this case.
The case of auranofin
We finally compared the reactivity of the studied inhibitors and CA with a well-known system investigated in the previous works: the binding of the gold(I)–drug auranofin (AF) [38, 44, 80, 81]. A recent paper of ours has clearly demonstrated that AF binds selectively to the free and solvent-accessible cysteine residues of proteins [38]. In the presence of an accessible thiol, AF loses its thiosugar ligand and binds covalently through the gold(I) center directly to the sulfur atom of the thiol. Through our well-consolidated protocol for the ESI–MS analysis, adducts with AuPEt3+ fragment were observed for some representative proteins containing solvent-accessible free Cys residues; the exclusive binding on this amino acid residue has been clearly confirmed by competition studies with ebselen, an organoselenium compound that binds selectively and covalently to thiols [82,83,84,85,86].
Especially for the case of CA I, the binding with AF takes place in acidic, denatured, conditions (0.1% v/v of formic acid, pH 2.9). The spectrum in Fig. 9 displays the deconvoluted mass spectrum of apo-CA with AF (1:3 protein to AF ratio) in the presence of acid. The signal of the apo protein is accompanied by two other peaks at 29,095 and 29,409 Da corresponding to the mono and bis adduct of the apo protein with AuPEt3+ fragment, respectively. The hCA I owns just one Cys residue (Cys212) not involved in disulphide bridges, therefore, potentially available for the binding with AF. However, the bis adduct most likely forms upon binding of two AuPEt3+ fragments to the thiol of Cys212, giving rise to the kinetically favored thiolated-bridged digold complex, as already observed by F. Shaw III [87].
Nevertheless, in the native-like conditions identified for hCA I, adduct formation has not been observed between AF and hCA I but only the holo-protein signal is detected (see SI). From this evidence, we inferred that, just after the partial unfolding of the protein, the free Cys residue becomes available for the binding to the gold center. Indeed, the Cys212 residue is not completely solvent-accessible in the CA I native state [88, 89], as it can be observed from the enzyme crystal structure reported in Fig. 10.
Only when some partial unfolding of the protein takes place (due for example to acidification), the Cys residue becomes spatially accessible for the binding with AF. Contrariwise, in the absence of a potential and accessible binding site, AF retains the thiosugar ligand and no interaction occurs.
Two additional experiments were conducted mixing both AF and MZA with CA in a solution of 1:3:3 protein/inhibitor/metallodrug ratio. First, we analyzed the sample in native-like conditions and just the mono-adduct between the holo-CA and MZA is detected (see SI). Again, the protein reacts completely with the inhibitor, but there is no AuPEt3+ biding. Adding to the same solution a 0.1% v/v of formic acid and repeating the ESI-MS analysis, the new mass spectrum is perfectly superimposable with the one displayed in Fig. 9 (see SI). In this case, the reveled adducts are only between AF and apo-CA; the Zn ion and the inhibitor are no more bound to the protein.
Conclusions
Native mass spectrometry has been recognized as a rapid, sensitive, high throughput, and label-free method to directly investigate protein–ligand interactions, preserving all the biological functions of the macromolecule. However, it is worth reminding that the so-called “native conditions” are strictly related to the nature of the analyzed protein and to its chemico-physical properties [28]. In this paper, we investigated in depth the parameters that are involved in the ESI–MS analysis of human carbonic anhydrase, starting from the solution pH value up to the critical instrumental parameters. We found that the best conditions for preserving the CA tertiary structure, jointly to its biological activity, require a solution pH value near to neutrality; therefore, the conventional addition of formic acid must be avoided. In fact, the resulting acidic conditions cause the release of the Zn ion and prevent the possibility to bind the inhibitor.
Once found the best conditions for CA analysis, we extended the MS study to the protein binding of three sulfonamide and one dithiocarbamate inhibitors. In studying these protein/inhibitor adducts, the declustering potential turned out to be another fundamental parameter to be considered to avoid an artificial alteration of the formed adduct.
Then, through this powerful and reliable methodology, we proved that the four inhibitors react rapidly and almost completely with the protein, forming mono and bis adducts in the case of sulfonamides and only mono adducts with dithiocarbamate; the adducts were characterized in detail.
As completion of this study, some competition experiments between inhibitors with different affinity for CA were performed. Also, in this case, the technique turned out to be extremely reliable, highlighting a good correlation between adduct formation and the relative inhibitors’ affinities for the protein itself.
Finally, some further experiments were carried out with the gold drug auranofin. This latter compound can be considered as a site-specific ligand, reacting only with free and solvent-accessible cysteine residues. Although auranofin does not hinder at all the reaction of CA with its inhibitors, it reacts only when the protein is partially unfolded, thus making the free cysteine residue accessible. At the same time, with a partial loss of the protein tertiary structure, the Zn ion and the inhibitor are released once again.
References
Fegler J (1944) Function of carbonic anhydrase in blood. Nature 153:137–138
Lindskog S (1997) Structure and mechanism of carbonic anhydrase. Pharmacol Ther 74:1–20
Pocker Y, Tanaka N (1978) Inhibition of carbonic anhydrase by anions in the carbon dioxide-bicarbonate system. Science (80-) 199:907–909. https://doi.org/10.1126/science.199.4331.907
Mann T, Keilin D (1940) Sulphanilamide as a specific inhibitor of carbonic anhydrase. Nature 146:164–165
Supuran CT (2008) Carbonic anhydrases: Novel therapeutic applications for inhibitors and activators. Nat Rev Drug Discov 7:168–181
Supuran CT (2016) Carbonic anhydrase inhibition and the management of neuropathic pain. Expert Rev Neurother 16:961–968
Supuran CT, Scozzafava A, Casini A (2003) Carbonic anhydrase inhibitors. Med Res Rev 23:146–189. https://doi.org/10.1002/med.10025
Supuran CT, Scozzafava A (2000) Carbonic anhydrase inhibitors: aromatic sulfonamides and disulfonamides act as efficient tumor growth inhibitors. J Enzyme Inhib 15:597–610. https://doi.org/10.3109/14756360009040713
Ferraroni M, Cornelio B, Sapi J, Supuran CT, Scozzafava A (2018) Sulfonamide carbonic anhydrase inhibitors: zinc coordination and tail effects influence inhibitory efficacy and selectivity for different isoforms. Inorganica Chim Acta 470:128–132. https://doi.org/10.1016/j.ica.2017.03.038
Degn H, Kristensen B (1986) Stopped flow mass spectrometry: applications to the carbonic anhydrase reaction. J Biochem Biophys Methods 12:305–310. https://doi.org/10.1016/0165-022X(86)90067-9
Day YSN, Baird CL, Rich RL, Myszka DG (2002) Direct comparison of binding equilibrium, thermodynamic, and rate constants determined by surface- and solution-based biophysical methods. Protein Sci 11:1017–1025. https://doi.org/10.1110/ps.4330102
Krishnamurthy VM, Kaufman GK, Urbach AR, Gitlin I, Gudiksen KL, Weibel DB, Whitesides GM (2008) Carbonic anhydrase as a model for biophysical and physical-organic studies of proteins and protein-ligand binding. Chem Rev 108:946–1051
Pacholarz KJ, Garlish RA, Taylor RJ, Barran PE (2012) Mass spectrometry based tools to investigate protein–ligand interactions for drug discovery. Chem Soc Rev 41:4335. https://doi.org/10.1039/c2cs35035a
Siegel M (2002) Early discovery drug screening using mass spectrometry. Curr Top Med Chem 2:13–33
Lee M (2005) Integrated strategies for drug discovery using mass spectrometry. Wiley, Hoboken
Hofstadler SA, Sannes-Lowery KA (2006) Applications of ESI-MS in drug discovery: interrogation of noncovalent complexes. Nat Rev Drug Discov 5:585–595
Leney AC, Heck AJR (2017) Native mass spectrometry: what is in the Name? J Am Soc Mass Spectrom 28:5–13. https://doi.org/10.1007/s13361-016-1545-3
Breuker K, McLafferty FW (2008) Stepwise evolution of protein native structure with electrospray into the gas phase, 10–12 to 102 s. Proc Natl Acad Sci USA 105:18145–18152
Ganem B, Li YT, Henion JD (1991) Detection of noncovalent receptor-ligand complexes by mass spectrometry. J Am Chem Soc 113:6294–6296. https://doi.org/10.1021/ja00016a069
Katta V, Chait BT (1991) Observation of the heme–globin complex in native myoglobin by electrospray-ionization mass spectrometry. J Am Chem Soc 113:8534–8535. https://doi.org/10.1021/ja00022a058
Loo JA (1997) Studying noncovalent protein complexes by electrospray ionization mass spectrometry. Mass Spectrom Rev 16:1–23. https://doi.org/10.1002/(SICI)1098-2787(1997)16:1<1:AID-MAS1>3.0.CO;2-L
Hofstadler SA, Griffey RH (2001) Analysis of noncovalent complexes of DNA and RNA by mass spectrometry. Chem Rev 101:377–390. https://doi.org/10.1021/cr990105o
Woods LA, Dolezal O, Ren B, Ryan JH, Peat TS, Poulsen SA (2016) Native state mass spectrometry, surface plasmon resonance, and X-ray crystallography correlate strongly as a fragment screening combination. J Med Chem 59:2192–2204. https://doi.org/10.1021/acs.jmedchem.5b01940
Chowdhury SK, Katta V, Chait BT (1990) Probing conformational changes in proteins by mass spectrometry. J Am Chem Soc 112:9012–9013. https://doi.org/10.1021/ja00180a074
Kaltashov IA, Zhang M, Eyles SJ, Abzalimov RR (2006) Investigation of structure, dynamics and function of metalloproteins with electrospray ionization mass spectrometry. Anal Bioanal Chem 386:472–481
Kaltashov AI, Mohimen A (2005) Estimates of protein surface areas in solution by electrospray ionization mass spectrometry. Anal Chem 77:5370–5379. https://doi.org/10.1021/ac050511+
Grandori R, Santambrogio C, Brocca S, Invernizzi G, Lotti M (2009) Electrospray-ionization mass spectrometry as a tool for fast screening of protein structural properties. Biotechnol J 4:73–87. https://doi.org/10.1002/biot.200800250
Li J, Santambrogio C, Brocca S, Rossetti G, Carloni P, Grandori R (2016) Conformational effects in protein electrospray-ionization mass spectrometry. Mass Spectrom Rev 35:111–122. https://doi.org/10.1002/mas.21465
Konermann L, Douglas DJ (1997) Acid-induced unfolding of cytochrome c at different methanol concentrations: electrospray ionization mass spectrometry specifically monitors changes in the tertiary structure. Biochemistry 36:12296–12302. https://doi.org/10.1021/bi971266u
Nabuchi Y, Murao N, Asoh Y, Takayama M (2007) Probing the unfolding and refolding processes of carbonic anhydrase 2 using electrospray ionization mass spectrometry combined with pH jump. Anal Chem 79:8342–8349. https://doi.org/10.1021/ac071130u
Carta F, Aggarwal M, Maresca A, Scozzafava A, McKenna R, Masini E, Supuran CT (2012) Dithiocarbamates strongly inhibit carbonic anhydrases and show antiglaucoma action in vivo. J Med Chem 55:1721–1730. https://doi.org/10.1021/jm300031j
Pacchiano F, Aggarwal M, Avvaru BS, Robbins AH, Scozzafava A, McKenna R, Supuran CT (2010) Selective hydrophobic pocket binding observed within the carbonic anhydrase II active site accommodate different 4-substituted-ureido-benzenesulfonamides and correlate to inhibitor potency. Chem Commun 46:8371–8373. https://doi.org/10.1039/c0cc02707c
Aggarwal M, Kovalevsky AY, Velazquez H, Fisher SZ, Smith JC, McKenna R (2016) Neutron structure of human carbonic anhydrase II in complex with methazolamide: mapping the solvent and hydrogen-bonding patterns of an effective clinical drug. IUCrJ 3:319–325. https://doi.org/10.1107/S2052252516010514
Sippel KH, Robbins AH, Domsic J, Genis C, Agbandje-Mckenna M, McKenna R (2009) High-resolution structure of human carbonic anhydrase II complexed with acetazolamide reveals insights into inhibitor drug design. Acta Crystallogr Sect F Struct Biol Cryst Commun 65:992–995. https://doi.org/10.1107/S1744309109036665
McDonald PC, Chia S, Bedard PL, Chu Q, Lyle M, Tang L, Singh M, Zhang Z, Supuran CT, Renouf DJ, Dedhar S (2020) A phase 1 study of SLC-0111, a novel inhibitor of carbonic anhydrase IX, in patients with advanced solid tumors. Am J Clin Oncol Cancer Clin Trials 43:484–490. https://doi.org/10.1097/COC.0000000000000691
Nocentini A, Supuran CT (2019) Advances in the structural annotation of human carbonic anhydrases and impact on future drug discovery. Expert Opin Drug Discov 14:1175–1197
Avvaru BS, Kim CU, Sippel KH, Gruner SM, Agbandje-McKenna M, Silverman DN, McKenna R (2010) A short, strong hydrogen bond in the active site of human carbonic anhydrase II. Biochemistry 49:249–251. https://doi.org/10.1021/bi902007b
Zoppi C, Messori L, Pratesi A (2020) ESI MS studies highlight the selective interaction of Auranofin with protein free thiols. Dalt Trans 49:5906–5913. https://doi.org/10.1039/d0dt00283f
King RW, Roberts GCK (1971) Nuclear magnetic resonance studies of human carbonic anhydrase B. Histidine residues. Biochemistry 10:558–565. https://doi.org/10.1021/bi00780a003
Song H, Wilson DL, Farquhar ER, Lewis EA, Emerson JP (2012) Revisiting zinc coordination in human carbonic anhydrase II. Inorg Chem 51:11098–11105. https://doi.org/10.1021/ic301645j
Brasuń J, Matera-Witkiewicz A, Ołdziej S, Pratesi A, Ginanneschi M, Messori L (2009) Impact of ring size on the copper(II) coordination abilities of cyclic tetrapeptides. J Inorg Biochem 103:813–817. https://doi.org/10.1016/j.jinorgbio.2009.02.003
Matera-Witkiewicz A, Brasuń J, Światek-Kozłowska J, Pratesi A, Ginanneschi M, Messori L (2009) Short-chain oligopeptides with copper(II) binding properties: the impact of specific structural modifications on the copper(II) coordination abilities. J Inorg Biochem 103:678–688. https://doi.org/10.1016/j.jinorgbio.2009.01.001
Coleman JE (1965) Human carbonic anhydrase. Protein conformation and metal ion binding. Biochemistry 4:2644–2655. https://doi.org/10.1021/bi00888a014
Pratesi A, Cirri D, Ciofi L, Messori L (2018) Reactions of auranofin and its pseudohalide derivatives with serum albumin investigated through ESI-Q-TOF MS. Inorg Chem 57:10507–10510. https://doi.org/10.1021/acs.inorgchem.8b02177
Michelucci E, Pieraccini G, Moneti G, Gabbiani C, Pratesi A, Messori L (2017) Mass spectrometry and metallomics: a general protocol to assess stability of metallodrug-protein adducts in bottom-up MS experiments. Talanta 167:30–38. https://doi.org/10.1016/j.talanta.2017.01.074
Marzo T, De Pascali SA, Gabbiani C, Fanizzi FP, Messori L, Pratesi A (2017) ESI–MS studies of the reactions of novel platinum(II) complexes containing O,O′-chelated acetylacetonate and sulfur ligands with selected model proteins. Biometals 30:609–614. https://doi.org/10.1007/s10534-017-0031-0
Pratesi A, Cirri D, Đurović MD, Pillozzi S, Petroni G, Bugarčić ŽD, Messori L (2016) New gold carbene complexes as candidate anticancer agents. Biometals 29:905–911. https://doi.org/10.1007/s10534-016-9962-0
Massai L, Pratesi A, Bogojeski J, Banchini M, Pillozzi S, Messori L, Bugarčić ŽD (2016) Antiproliferative properties and biomolecular interactions of three Pd(II) and Pt(II) complexes. J Inorg Biochem 165:1–6. https://doi.org/10.1016/j.jinorgbio.2016.09.016
Tamasi G, Carpini A, Valensin D, Messori L, Pratesi A, Scaletti F, Jakupec M, Keppler B, Cini R (2014) {Ru(CO)x}-core complexes with selected azoles: synthesis, X-ray structure, spectroscopy, DFT analysis and evaluation of cytotoxic activity against human cancer cells. Polyhedron 81:227–237
Heck AJR (2008) Native mass spectrometry: a bridge between interactomics and structural biology. Nat Methods 5:927–933. https://doi.org/10.1038/nmeth.1265
Donnelly DP, Rawlins CM, DeHart CJ, Fornelli L, Schachner LF, Lin Z, Lippens JL, Aluri KC, Sarin R, Chen B, Lantz C, Jung W, Johnson KR, Koller A, Wolff JJ, Campuzano IDG, Auclair JR, Ivanov AR, Whitelegge JP et al (2019) Best practices and benchmarks for intact protein analysis for top-down mass spectrometry. Nat Methods 16:587–594. https://doi.org/10.1038/s41592-019-0457-0
Yin S, Loo JA (2011) Top-down mass spectrometry of supercharged native protein-ligand complexes. Int J Mass Spectrom 300:118–122. https://doi.org/10.1016/j.ijms.2010.06.032
Lomeli SH, Yin S, Ogorzalek Loo RR, Loo JA (2009) Increasing charge while preserving noncovalent protein complexes for ESI-MS. J Am Soc Mass Spectrom 20:593–596. https://doi.org/10.1016/j.jasms.2008.11.013
Natalello A, Santambrogio C, Grandori R (2017) Are charge-state distributions a reliable tool describing molecular ensembles of intrinsically disordered proteins by native MS? J Am Soc Mass Spectrom 28:21–28. https://doi.org/10.1007/s13361-016-1490-1
Hamdy OM, Julian RR (2012) Reflections on charge state distributions, protein structure, and the mystical mechanism of electrospray ionization. J Am Soc Mass Spectrom 23:1–6. https://doi.org/10.1007/s13361-011-0284-8
Kafader JO, Melani RD, Schachner LF, Ives AN, Patrie SM, Kelleher NL, Compton PD (2020) Native vs denatured: an in depth investigation of charge state and isotope distributions. J Am Soc Mass Spectrom 31:574–581. https://doi.org/10.1021/jasms.9b00040
Heck AJR, van den Heuvel RHH (2004) Investigation of intact protein complexes by mass spectrometry. Mass Spectrom Rev 23:368–389. https://doi.org/10.1002/mas.10081
Yan X, Watson J, Ho PS, Deinzer ML (2004) Mass spectrometric approaches using electrospray ionization charge states and hydrogen-deuterium exchange for determining protein structures and their conformational changes. Mol Cell Proteom 3:10–23
Krusemark CJ, Frey BL, Belshaw PJ, Smith LM (2009) Modifying the charge state distribution of proteins in electrospray ionization mass spectrometry by chemical derivatization. J Am Soc Mass Spectrom 20:1617–1625. https://doi.org/10.1016/j.jasms.2009.04.017
Menchise V, De SG, Di FA, Scozzafava A, Supuran CT (2006) Carbonic anhydrase inhibitors: X-ray crystallographic studies for the binding of 5-amino-1,3,4-thiadiazole-2-sulfonamide and 5-(4-amino-3-chloro-5-fluorophenylsulfonamido)-1,3,4-thiadiazole-2-sulfonamide to human isoform II. Bioorganic Med Chem Lett 16:6204–6208. https://doi.org/10.1016/j.bmcl.2006.09.022
Alterio V, Di Fiore A, D’Ambrosio K, Supuran CT, De Simone G (2012) Multiple binding modes of inhibitors to carbonic anhydrases: How to design specific drugs targeting 15 different isoforms? Chem Rev 112:4421–4468
Winum JY, Supuran CT (2015) Recent advances in the discovery of zinc-binding motifs for the development of carbonic anhydrase inhibitors. J Enzyme Inhib Med Chem 30:321–324
Gao J, Wu Q, Carbeck J, Lei QP, Smith RD, Whitesides GM (1999) Probing the energetics of dissociation of carbonic anhydrase-ligand complexes in the gas phase. Biophys J 76:3253–3260. https://doi.org/10.1016/S0006-3495(99)77477-1
Dugad LB, Cooley CR, Gerig JT (1989) NMR studies of carbonic anhydrase-fluorinated benzenesulfonamide complexes. Biochemistry 28:3955–3960. https://doi.org/10.1021/bi00435a049
Kovalevsky A, Aggarwal M, Velazquez H, Cuneo MJ, Blakeley MP, Weiss KL, Smith JC, Fisher SZ, McKenna R (2018) “To be or not to be” protonated: atomic details of human carbonic anhydrase-clinical drug complexes by neutron crystallography and simulation. Structure 26:383–390.e3. https://doi.org/10.1016/j.str.2018.01.006
Avvaru BS, Busby SA, Chalmers MJ, Griffin PR, Venkatakrishnan B, Agbandje-McKenna M, Silverman DN, McKenna R (2009) Apo-human carbonic anhydrase II revisited: implications of the loss of a metal in protein structure, stability, and solvent network. Biochemistry 48:7365–7372. https://doi.org/10.1021/bi9007512
Ray SS, Singh SK, Balaram P (2001) An electrospray ionization mass spectrometry investigation of 1-anilino-8-naphthalene-sulfonate (ANS) binding to proteins. J Am Soc Mass Spectrom 12:428–438. https://doi.org/10.1016/S1044-0305(01)00206-9
Thomson BA (1997) Declustering and fragmentation of protein ions from an electrospray ion source. J Am Soc Mass Spectrom 8:1053–1058. https://doi.org/10.1016/S1044-0305(97)00129-3
Susa AC, Xia Z, Tang HYH, Tainer JA, Williams ER (2017) Charging of proteins in native mass spectrometry. J Am Soc Mass Spectrom 28:332–340. https://doi.org/10.1007/s13361-016-1517-7
Huang EC, Pramanik BN, Tsarbopoulos A, Reichert P, Ganguly AK, Trotta PP, Nagabhushan TL, Covey TR (1993) Application of electrospray mass spectrometry in probing protein-protein and protein-ligand noncovalent interactions. J Am Soc Mass Spectrom 4:624–630. https://doi.org/10.1016/1044-0305(93)85026-T
Baca M, Kent SBH (1992) Direct observation of a ternary complex between the dimeric enzyme HIV-1 protease and a substrate-based inhibitor. J Am Chem Soc 114:3992–3993
Busemann A, Araman C, Flaspohler I, Pratesi A, Zhou X-Q, van Rixel HS, Siegler AM, Messori L, van Kasteren SI, Bonnet S (2020) Alkyne functionalization of a photoactivated ruthenium polypyridyl complex for click-enabled serum albumin interaction studies. Inorg Chem 59:7710–7720. https://doi.org/10.1021/acs.inorgchem.0c00742
Lomelino CL, Mahon BP, McKenna R, Carta F, Supuran CT (2016) Kinetic and X-ray crystallographic investigations on carbonic anhydrase isoforms I, II, IX and XII of a thioureido analog of SLC-0111. Bioorganic Med Chem 24:976–981. https://doi.org/10.1016/j.bmc.2016.01.019
Nguyen GTH, Tran TN, Podgorski MN, Bell SG, Supuran CT, Donald WA (2019) Nanoscale ion emitters in native mass spectrometry for measuring ligand-protein binding affinities. ACS Cent Sci 5:308–318. https://doi.org/10.1021/acscentsci.8b00787
Nguyen GTH, Nocentini A, Angeli A, Gratteri P, Supuran CT, Donald WA (2020) Perfluoroalkyl substances of significant environmental concern can strongly inhibit human carbonic anhydrase isozymes. Anal Chem 92:4614–4622. https://doi.org/10.1021/acs.analchem.0c00163
De Simone G, Langella E, Esposito D, Supuran CT, Monti SM, Winum J-Y, Alterio V (2017) Insights into the binding mode of sulphamates and sulphamides to hCA II: crystallographic studies and binding free energy calculations. J Enzyme Inhib Med Chem 32:1002–1011. https://doi.org/10.1080/14756366.2017.1349764
Bachmann KA, Lewis JD (2005) Predicting inhibitory drug—drug interactions and evaluating drug interaction reports using inhibition constants. Ann Pharmacother 39:1064–1072. https://doi.org/10.1345/aph.1E508
El-Azab AS, Abdel-Aziz AAM, Bua S, Nocentini A, AlSaif NA, Almehizia AA, Alanazi MM, Hefnawy MM, Supuran CT (2019) New anthranilic acid-incorporating N-benzenesulfonamidophthalimides as potent inhibitors of carbonic anhydrases I, II, IX, and XII: synthesis, in vitro testing, and in silico assessment. Eur J Med Chem 181:111573. https://doi.org/10.1016/j.ejmech.2019.111573
Brzozowski Z, Sławiński J, Vullo D, Supuran CT (2012) Carbonic anhydrase inhibitors. Regioselective synthesis of novel series 1-substituted 1,4-dihydro-4-oxo-3-pyridinesulfonamides and their inhibition of the human cytosolic isozymes I and II and transmembrane cancer-associated isozymes IX and XII. Eur J Med Chem 56:282–291. https://doi.org/10.1016/j.ejmech.2012.08.006
Marzo T, Massai L, Pratesi A, Stefanini M, Cirri D, Magherini F, Becatti M, Landini I, Nobili S, Mini E, Crociani O, Arcangeli A, Pillozzi S, Gamberi T, Messori L (2019) Replacement of the thiosugar of auranofin with iodide enhances the anticancer potency in a mouse model of ovarian cancer. ACS Med Chem Lett 10:656–660. https://doi.org/10.1021/acsmedchemlett.9b00007
Landini I, Lapucci A, Pratesi A, Massai L, Napoli C, Perrone G, Pinzani P, Messori L, Mini E, Nobili S (2017) Selection and characterization of a human ovarian cancer cell line resistant to auranofin. Oncotarget 8:96062–96078. https://doi.org/10.18632/oncotarget.21708
Maurais AJ, Weerapana E (2019) Reactive-cysteine profiling for drug discovery. Curr Opin Chem Biol 50:29–36
Xu K, Zhang Y, Tang B, Laskin J, Roach PJ, Chen H (2010) Study of highly selective and efficient thiol derivatization using selenium reagents by mass spectrometry. Anal Chem 82:6926–6932. https://doi.org/10.1021/ac1011602
Capper MJ, Wright GSA, Barbieri L, Luchinat E, Mercatelli E, McAlary L, Yerbury JJ, O’Neill PM, Antonyuk SV, Banci L, Hasnain SS (2018) The cysteine-reactive small molecule ebselen facilitates effective SOD1 maturation. Nat Commun. https://doi.org/10.1038/s41467-018-04114-x
Terentis AC, Freewan M, Sempértegui Plaza TS, Raftery MJ, Stocker R, Thomas SR (2010) The selenazal drug ebselen potently inhibits indoleamine 2,3-dioxygenase by targeting enzyme cysteine residues. Biochemistry 49:591–600. https://doi.org/10.1021/bi901546e
Müller A, Cadenas E, Graf P, Sies H (1984) A novel biologically active seleno-organic compound-1. Glutathione peroxidase-like activity in vitro and antioxidant capacity of PZ 51 (Ebselen). Biochem Pharmacol 33:3235–3239. https://doi.org/10.1016/0006-2952(84)90083-2
Isab AA, Frank Shaw C, Lockela J, Hoeschele JD (1988) Reactions of trimethylphosphine analogues of auranofin with bovine serum albumin. Inorg Chem 27:3588–3592. https://doi.org/10.1021/ic00293a029
Carlsson U, Aasa R, Jonsson B-H, Lindskog S, Henderson LE (1975) Paramagnetic and fluorescent probes attached to “buried” sulfhydryl groups in human carbonic anhydrases. Application to inhibitor binding, denaturation and refolding. Eur J Biochem 52:25–36. https://doi.org/10.1111/j.1432-1033.1975.tb03969.x
Rana TM, Meares CF (1991) Transfer of oxygen from an artificial protease to peptide carbon during proteolysis. Proc Natl Acad Sci USA 88:10578–10582
Acknowledgements
L. M. and A. P. acknowledge the Fondazione Italiana per la Ricerca sul Cancro (AIRC), Milan, and Fondazione Cassa Risparmio Firenze for funding the project “Advanced mass spectrometry tools for cancer research: novel applications in proteomics, metabolomics and nanomedicine” (Multi-user Equipment Program 2016, Ref. code 19650). CIRCMSB and Benefiecentia Stiftung (Vaduz) are also acknowledged. A. P. would like to thank University of Pisa (Rating Ateneo 2019-2020) for the financial support.
Funding
Open access funding provided by Università di Pisa within the CRUI-CARE Agreement.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zoppi, C., Nocentini, A., Supuran, C.T. et al. Native mass spectrometry of human carbonic anhydrase I and its inhibitor complexes. J Biol Inorg Chem 25, 979–993 (2020). https://doi.org/10.1007/s00775-020-01818-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-020-01818-8