Abstract
Methane formation in methanogenic Archaea is catalyzed by methyl-coenzyme M reductase (MCR) and takes place via the reduction of methyl-coenzyme M (CH3-S-CoM) with coenzyme B (HS-CoB) to methane and the heterodisulfide CoM-S–S-CoB. MCR harbors the nickel porphyrinoid coenzyme F430 as a prosthetic group, which has to be in the Ni(I) oxidation state for the enzyme to be active. To date no intermediates in the catalytic cycle of MCRred1 (red for reduced Ni) have been identified. Here, we report a detailed characterization of MCRred1m (“m” for methyl-coenzyme M), which is the complex of MCRred1a (“a” for absence of substrate) with CH3-S-CoM. Using continuous-wave and pulse electron paramagnetic resonance spectroscopy in combination with selective isotope labeling (13C and 2H) of CH3-S-CoM, it is shown that CH3-S-CoM binds in the active site of MCR such that its thioether sulfur is weakly coordinated to the Ni(I) of F430. The complex is stable until the addition of the second substrate, HS-CoB. Results from EPR spectroscopy, along with quantum mechanical calculations, are used to characterize the electronic and geometric structure of this complex, which can be regarded as the first intermediate in the catalytic mechanism.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Methyl-coenzyme M reductase (MCR) catalyzes the key step of methanogenesis in Archaea, namely the reduction of methyl-coenzyme M (CH3-S-CoM) with coenzyme B (HS-CoB) to methane and the heterodisulfide CoM-S–S-CoB (Eq. 1, Structure 1):
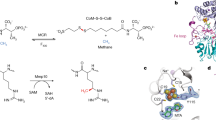
Left: schematic representation of coenzyme F430, CH3-S-CoM, Glnα′147 from the alpha subunit of MCR [3], and the orientation of the g 3-axis. Right: coenzyme M (HS-CoM), methyl-coenzyme M (CH 3 -S-CoM), and coenzyme B (HS-CoB)
Methanogenic Archaea are found in strictly anoxic habitats such as wetlands, sewage sites or the rumens and guts of animals, and they gain the energy necessary for ATP synthesis by producing methane from substrates such as H2/CO2, acetate, formate or methanol [1, 2]. This process is responsible for the largest part of the annual emission (estimated 5 × 108 tons per year) of this very effective greenhouse gas into the atmosphere. All methanogens contain MCR, and X-ray crystallography of inactive Ni(II) forms shows that the enzyme has two identical active sites, each containing one molecule of coenzyme F430 (Structure 1) [3]. In the active state, designated MCRred1, the central metal is in the Ni(I) valence state and its EPR spectrum is characteristic of a d 9 S = 1/2 species with the unpaired electron in a molecular orbital of predominantly nickel \( d_{{x^{2} - y^{2} }} \) character [4]. The active site of MCR is structured such that, of the two substrates, CH3-S-CoM has to bind first. The MCR states of relevance for this paper are summarized in Chart 1.
Chart showing different EPR-active forms of MCR and their interconversion. Catalysis by MCR is believed to start with MCRred1a, where the cofactor F430 has the Ni(I) oxidation state and the absence of a proximal substrate is denoted by “a.” Methyl-coenzyme M (CH3-S-CoM) is added to form the MCRred1m state, which is highlighted in red and is the main subject of this report. From this state, methane is produced by the addition of HS-CoB
The catalytic mechanism of the reduction (Eq. 1) at the nickel center is widely disputed [5–13]. In essence, two mechanisms mainly differing in the nature of the initial cleavage of the sulfur–carbon bond of CH3-S-CoM are currently discussed. In mechanism “A,” proposed by Pelmenschikov et al. [5, 6] on the basis of DFT calculations, the Ni(I) center is assumed to attack the thioether sulfur of CH3-S-CoM, generating ·CH3 and the thiolate complex CoM-S–Ni(II)F430 as intermediates (see Fig. 8).
According to mechanism “B,” the Ni(I) center initially acts as a nucleophile, attacking methyl-coenzyme M at the carbon of the CH3-S group, generating a CH3–Ni(III)F430 intermediate and HS-CoM [1, 9–12]. Recent findings indicate that such a species can exist: 3-bromopropane sulfonate was shown to react with the active enzyme MCRred1 to give −O3S(CH2)3–Ni(III)F430 + and Br−, while CH3Br and CH3I react with MCRred1 to form CH3–Ni(III)F430 + in the active site [14–17], with Br−/I− as the leaving group. Additional evidence is provided by the reaction of free Ni(I)F430 derivatives with electrophilic methyl donors such as methyl-dialkylsulfonium ions and methyl halides [16, 17]. These mechanistic studies, as well as the fact that methane formation proceeds under inversion of stereoconfiguration, are consistent with mechanism “B,” and CH3–Ni(III)L species have been postulated as intermediates on the basis of pulse radiolytic studies with simpler nickel model complexes, while a model close to the postulated CH3–Ni(III)F430 was found in a theoretical study to be a minimum on the hypersurface with a \( (d_{xy} )^{2} (d_{xz} )^{2} (d_{yz} )^{2} (d_{{x^{2} - y^{2} }} )^{1} \) ground-state configuration [18, 19]. In addition, CH3–Ni(II)F430 derivatives have been generated in situ and characterized spectroscopically [20].
Recently, a third type of reaction mechanism has been proposed by Duin and McKee [21] based on DFT calculations. In their study, the initial step is a protonation of Ni(I)F430, which is followed by the anchoring of CH3-S-CoM to the nickel via sulfur, and the rate-determining step is a subsequent oxidative addition. The transition state consists of a complex in which the methyl group and the sulfur of S-CoM are bound to the central nickel at the same time. For the sake of brevity we call this mechanism, which has features from both mechanisms “A” and “B,” mechanism “C.”
To date no chemical intermediates for Eq. 1 have been isolated and characterized, for example by freeze-quench experiments. It is known that the addition of the substrate CH3-S-CoM to active MCRred1a (“a” for absence of substrates) results in the formation of a species with a distinguishable CW EPR spectrum. This species, which has been named MCRred1m (“m” for methyl-coenzyme M as substrate) according to \( {\text{MCR}}_{{{\text{red}}1{\text{a}}}} \underset{{ -{\rm CH}_{3}{\text{-}} {\rm SCoM}}}{\overset{{ + {\rm CH}_{3}{\text{-}} {\rm SCoM}}}{\rightleftarrows}}{\text{ MCR}}_{{{\text{red}}1{\text{m}}}}, \) is stable until the addition of the second substrate HS-CoB [22].
Here, we report a detailed characterization of MCRred1m which shows that this state can be viewed as the first intermediate formed in the catalytic cycle of methyl-coenzyme M reductase. Using pulse and continuous-wave (CW) electron paramagnetic resonance (EPR) spectroscopy in combination with selective isotope labeling (13C and 2H) of CH3-S-CoM, and aided by quantum mechanical calculations, we find that there is weak coordination between Ni(I) and the thioether S of CH3-S-CoM. This surprising result is contrary to chemical intuition and experimental evidence showing that free Ni(I)F430M (pentamethylester of F430) in acetonitrile neither reacts with nor coordinates to CH3-S-CoM. Also, an X-ray absorption spectroscopy study of MCRred1m has shown that simulation of the spectra does not require the presence of a sulfur ligand, suggesting that CH3-S-CoM is not coordinated via its thioether sulfur atom to nickel in detectable amounts [23].
Materials and methods
Methanothermobacter marburgensis
Methanothermobacter marburgensis is the strain deposited under DSM 2133 in the Deutsche Sammlung von Mikroorganismen und Zellkulturen (Braunschweig). Coenzyme M (2-mercaptoethanesulfonate) was obtained from Fluka (Buchs, Switzerland); 2-(methylsulfanyl)ethanesulfonate (methyl-coenzyme M) was synthesized from coenzyme M by methylation with methyl iodide (Fluka). According to the same protocol, 2-([2H3]-methylsulfanyl)ethanesulfonate, 1, and 2-([13C]-methylsulfanyl)ethanesulfonate, 2, were synthesized from their corresponding iodides. Sodium methanolate (0.335 M) was prepared by dissolving 1.58 g of Na in 100 mL MeOH (distilled over Mg, I2) under nitrogen. [2H3]-methyl iodide was obtained from Dr. Glaser AG (Basel, Switzerland), and [13C]-methyl iodide from CIL (Cambridge Isotope Laboratories, Andover, MA, USA). All other routine chemicals were obtained from Fluka, Aldrich (Milwaukee, WI, USA), JT Baker (Phillipsburg, NJ, USA) and Merck (Darmstadt, Germany), and were used without further purification.
Purification of active MCR
M. marburgensis was grown at 65 °C in a 13 L glass fermenter (New Brunswick, Edison, NJ, USA) containing 10 L mineral medium stirred at 1,200 rpm and gassed with 80% H2/20% CO2/0.1% H2S at a rate of 1,200 mL/min [22]. When an ΔOD578 of 4.5 was reached, the gas supply was switched to 100% H2 for 30 min. Under these conditions, the intracellular methyl-coenzyme M reductase was reduced, as revealed by measuring the EPR signals of MCRred1 and MCRred2 shown by the intact cells [24]. After 30 min, the cells were cooled to 10 °C within 10 min under continuous gassing with 100% H2 and harvested anaerobically by centrifugation using a flow-through centrifuge (Hettich, Kirchlengern, Germany; centrifuge 17 RS). Approximately 70 g of wet cells were obtained. Only the MCR isoenzyme I was purified from these cells [25, 26]. All steps of the purification were performed in the presence of 10 mM coenzyme M and in an anaerobic chamber (Coy Instruments, Ann Arbor, MI, USA) filled with 95% N2/5% H2 as described previously [22]. During purification the enzyme lost its red2-type signal due to the removal of coenzyme B. In one purification, 150 mg of active MCR in the red1c state (in 3–4 ml) were generally obtained. The spin concentration per mol F430 was higher than 0.8. To obtain MCRred1a, the purified MCRred1c was washed free of coenzyme M with 50 mM Tris/HCl pH 7.6 by ultrafiltration with Amicon Ultra centrifugal filter devices with a 100 kDa cut-off (Millipore, Bedford, MA). The spin concentration per mole F430 generally decreased to values of between 0.6 and 0.8 during the washing procedure. MCRred1a was converted to MCRred1m through the addition of methyl-coenzyme M (13CH3-CoM or CD3-CoM) to give a final concentration of 10 mM. The protein concentration was determined by using the method of Bradford [27] with bovine serum albumin (Serva, Heidelberg, Germany) used as standard or by measuring the absorbance difference for the oxidized enzyme (MCRsilent) at 420 nm using ε = 44,000 M−1 cm−1 for a molecular mass of 280,000 Da. Both methods yielded almost the same results. The final concentration of protein obtained in our experiments was 0.6 mM.
Synthesis of 2-([2H3]-methylsulfanyl)ethanesulfonate (1) [22] [[2H3]-methyl-coenzyme M (NH4 + form)]
About 580 μL of [2H3]-methyl iodide were added to a solution of 1 g of 2-mercaptoethanesulfonate (sodium salt; 6.09 mmol) in 20 mL of 0.335 M sodium methanolate (6.09 mmol) at 0 °C (ice bath) and stirred at room temperature for 14 h. The dried residue was dissolved in a small amount of water, acidified with Amberlite IR-120 to pH 1, treated with concentrated NH3(aq) to get pH 12, and lyophilized to give 1.01 g of pure (>99% 2H-labeled according to 1H NMR) 1 (5.67 mmol; 97%). 1H NMR (D2O, 400 MHz) δ 3.20–3.24 (m, 2H), 2.88–2.92 (m, 2H). 13C NMR (D2O, 400 MHz) δ 16.41 (septet, J = 21.3), 30.27, 53.40. ESI-MS: m/z 95.3 (11), 91.1 (26), 113.0 (38), 156.1 (46), 62.3 (48), 45.3 (84), 157.9 (100).
Synthesis of 2-([13C]-methylsulfanyl)ethanesulfonate (2) [28] [[13C]-methyl-coenzyme M (NH4 + form)]
About 1 g of [13C]-methyl iodide (7.0 mmol) was added to a solution of 1.148 g of 2-mercaptoethanesulfonate (sodium salt; 7.0 mmol) in 23 mL of 0.335 M sodium methanolate (6.09 mmol) at 0 °C (ice bath) and stirred at room temperature for 14 h. The dried residue was dissolved in a small amount of water, acidified with Amberlite IR-120 to pH 1, treated with concentrated NH3(aq) to give pH 12, and lyophilized to give 1.10 g of pure (>99% 13C-labeled according to 1H NMR) 2 (6.31 mmol; 90%). 1H NMR (D2O, 400 MHz) δ 1.98–2.32 (d, J = 139.4, 3H), 2.83–2.88 (m, 2H), 3.15–3.19 (m, 2H). 13C NMR (D2O, 400 MHz) δ 17.00 (100), 30.36 (1), 53.33 (1). ESI-MS: m/z 206.1 (8), 46.4 (9), 334.8 (13), 125.1 (15), 62.3 (67), 156.0 (100).
Preparation of an EPR sample of Ni(I)F430 pentamethyl ester in tetrahydrofuran/2-methyl-tetrahydrofuran
Ni(II)F430 pentamethyl ester (ClO4 − salt) [F430M] was prepared via exhaustive methylation of crude Ni(II)F430 pentacarboxylic acid in methanol/p-TsOH and purified by thin-layer chromatography as described [29]. To achieve the removal of all of the methylene chloride remaining from the isolation procedure, the pentamethyl ester was dissolved in THF/toluene 1:1 (v/v) and the solvent mixture was evaporated three times. A solution of 0.5 μmol F430M in 400 μL THF/2-methyl-THF 2:8 (v/v) was placed in one side arm of a glass apparatus with two side arms as well as a sealed-on UV/VIS cell (d = 0.024 cm) and an X-band quartz EPR tube. Both organic solvents were freshly distilled from metallic potassium. Two hundred microliters of liquid 0.025% (w/w) Na–Hg were placed in the other side arm, which was separated from the main compartment by a fritted glass disk (porosity P3). Both the F430M solution and the amalgam were degassed by performing three freeze–thaw cycles at 10−5 mbar, and the whole apparatus was sealed under high vacuum. Quantitative reduction of Ni(II)F430M to the Ni(I) form was brought about by controlled contact with the amalgam while following the progress by UV/VIS spectroscopy. The final solution of Ni(I)F430M was then allowed to flow into the EPR tube, which was sealed off under vacuum after freezing the solution at 77 K to form a glass.
EPR spectroscopy [30]
The Q-band (35 GHz) experiments were carried out on an instrument built in-house [31] and equipped with a helium gas-flow cryostat from Oxford Instruments, Inc. (Beckenham, UK). All X- and W-band (9.7/94 GHz) measurements were carried out on a Bruker (Karlsruhe, Germany) E680 spectrometer. The 13C X-band Davies ENDOR [30, 32] spectra were measured at 25 K with the mw pulse sequence π–T–π/2–τ–π–τ–echo, with mw pulses of length t π/2 = 400 ns and t π = 800 ns, τ = 1,600 ns, and a radiofrequency pulse of length 28.5 μs and with variable frequency ν rf applied for a time T = 30 μs. A low-pass filter with a cut-off frequency of 7 MHz was used to avoid artifacts from higher-rf harmonics. The W-band MIMS 13C ENDOR spectra were measured at 15 K with the mw pulse sequence π/2–τ–π/2–T–π/2–τ–echo, with mw pulses of length t π/2 = 24 ns, τ = 300–500 ns as indicated, and a radiofrequency pulse of length 48 μs and with variable frequency ν rf applied for a time T = 50 μs. The variable mixing time W-band MIMS ENDOR experiments [33, 34] (see Figure S1 in the Supplementary material) with the mw sequence π/2–τ–π/2–T–t mix–π/2–τ–echo were performed at 10 K with a repetition time of 20 ms and with mixing times t mix = 0 ms, 1.5 ms (all other parameters were the same as for the W-band MIMS ENDOR measurements). The asymmetry of the ENDOR peaks as the time t mix is increased can then be used to determine the absolute sign of the hyperfine interaction.
HYSCORE (hyperfine sublevel correlation spectroscopy) [35] is a two-dimensional (2D) experiment which correlates nuclear frequencies in one electron spin manifold with nuclear frequencies in the other electron spin manifold. X-band (9.7 GHz) HYSCORE spectra were measured at 25 K with a repetition time of 1.5 ms using the sequence π/2–τ–π/2–t 1–π–t 2–π/2–τ–echo. The mw pulse lengths were t π/2 = 20 ns and t π = 16 ns, starting times t 10 = t 20 = 96 ns, and a time increment of Δt = 20 ns (data matrix 350 × 350) was used. Q-band Matched-HYSCORE spectra [36, 37] were measured with a loop gap resonator [38] at 30 K using a repetition time of 1 ms and the sequence π/2–τ–t m–t 1–π–t 2–t m–τ–echo. The mw pulse lengths were t π = t π/2 = 16 ns, with matched pulses of length t m = 46 ns (mw field strength ω 1/2π = 83.3 MHz), starting times for t 1 and t 2 of 96 ns, a time increment of Δt = 12 ns (data matrix 256 × 256), and τ = 108 ns. All HYSCORE data were processed with MATLAB 7.0 (The MathWorks, Inc., Natick, MA, USA). The time traces were baseline-corrected with an exponential function, apodized with a Gaussian window, and zero-filled. After a 2D Fourier transform, absolute-value spectra were calculated. All measurements used an eight-step phase cycle to remove unwanted echoes. The field-swept EPR and Davies ENDOR spectra were simulated with the MATLAB-based program package EasySpin [39], and the HYSCORE spectra with a program written in-house [40].
Density functional theory calculations
Structural optimizations were carried out with the Turbomole program package [41] by employing two different density functionals, namely the pure functional BP86 [42, 43] (in combination with the resolution-of-the-identity density fitting technique with Karlsruhe auxiliary basis sets [44]) and the hybrid functional B3LYP (i.e., Becke’s three-parameter functional in combination with the LYP correlation functional [45, 46], as implemented in Turbomole). For all calculations, the valence-triple-zeta plus polarization basis set TZVP by Schäfer et al. [47] was applied. Coordination energies are given below for the structure optimized with the functional BP86. EPR-spectroscopic parameters were obtained by ADF calculations only for the BP86/TZVP structure (see Tables S1, S2, Figure S6). It is well known that BP86 structures of transition metal complexes are in general more reliable than B3LYP structures when compared to X-ray structural data. The (exothermic) coordination energy of the thioether to the nickel ion at 0 K amounts to −16.9 kJ/mol. This energy has been obtained for relaxed isolated fragments, i.e., for CH3-S-CoM and the metal fragment, respectively.
The hyperfine interactions were calculated with the Amsterdam Density Functional package (ADF 2005.01) [48]. The functional RPBE [49] with the relativistic scalar zeroth-order regular approximation (ZORA [50]) was employed. The calculation was spin-unrestricted, with a Slater-type basis set of triple-ζ quality with two polarization functions (TZ2P) with no frozen cores, as implemented in ADF.
Results and discussion
Figure 1 displays echo-detected W-band (94.2587 GHz) EPR spectra of MCRred1a (containing Ni(I)F430 in the “absence” of a substrate) and of MCRred1m, the species formed when CH3-S-CoM is added to a MCRred1a preparation. MCRred1a displays an EPR spectrum that can be simulated by two species labeled c1 (33%) and c2 (67%). c1 is most likely the species MCRred1c (\( {\text{MCR}}_{{{\text{red}}1{\text{a}}}} \underset{{ - {\rm H} {\text{-}}{\rm SCoM}}}{\overset{{ + {\rm H} {\text{-}}{\rm SCoM}}}{\rightleftarrows}}{\text{ MCR}}_{{{\text{red}}1{\text{c}}}} , \) with “c” denoting the presence of coenzyme M), since c1 has similar g values to those of a MCRred1c preparation (see the bottom row of Table 1) and HS-CoM is known to be present in the preparations in small quantities and to be difficult to remove completely. Species c2 is thus assigned to the “true” MCRred1a complex without a bound substrate and has an axial EPR spectrum (g 1 ≈ g 2 ≠ g 3). The addition of the substrate CH3-S-CoM causes a small but clearly discernable change in the EPR spectrum, with a well-resolved rhombic splitting (g 1 ≠ g 2 ≠ g 3), and the spectrum can be simulated by a single species with a rhombic g-matrix.
W-band (94.2587 GHz) EPR spectra of (A) MCRred1a and (B, C) MCRred1m. A and B show the echo-detected field-swept EPR spectrum measured at 20 K. The numerically calculated first derivative is shown in each case. C shows the CW EPR spectrum recorded at 235 K. The MCRred1a sample was simulated by two components, c1 (assigned to MCRred1c) and c2 (assigned to MCRred1a), and MCRred1m was simulated by a single component with a rhombic g-matrix
The induction of rhombicity in the g-matrix upon the addition of substrates has also been observed when HS-CoM is added to MCRred1a to form MCRred1c, and is an indication that a small change in the electronic structure takes place such that the electronic ground state \( (d_{{x^{2} - y^{2} }} ) \) has a larger \( d_{{z^{2} }} {\text{-component}} \) admixture through interaction with a proximal ligand.
A comparison between the echo-detected spectrum recorded at 20 K and the CW spectrum recorded at 235 K shows that the complex MCRred1m is stable in this temperature range, and provides an indication that the complex could well be stable at the physiological temperature of the enzyme at approximately 60 °C.
Hydropyrrolic nitrogens of F430
To characterize the electronic and geometric structure of MCRred1m, we studied the interactions of the Ni-centered electron spin with the four hydropyrrolic nitrogens of F430. The nitrogen nuclei have large hyperfine couplings, resulting in well-resolved splittings in the X-band CW EPR spectrum, as shown in Fig. 2a. To improve the accuracy of the determined EPR parameters, a matched HYSCORE spectrum (Fig. 2b) and Q-band Davies ENDOR spectra (Fig. 2c) were also recorded. The matched HYSCORE spectrum was used to supplement the data recorded at the observer positions representing g 1/g 2 (close to the CW EPR spectral maximum), where the signals are most complicated due to many contributing orientations. In this spectrum, the positions of the double-quantum cross-peaks, which are, to first order, free from the nuclear quadrupole interaction (NQI) and are centered at the hyperfine coupling A and split by 4ν I (ν I is the nuclear Larmor frequency), show that the hyperfine couplings A 1/A 2 range from 23–33 MHz. The 14N signals in the CW EPR and Q-band ENDOR spectra were then simulated with this constraint to give the best fit to the data, which resulted in the hyperfine and nuclear quadrupole parameters given in Table 2. To reduce the number of fitted parameters, the Euler angles were fixed in line with single-crystal metalloporphyrin studies [51] where the axis of the largest principal hyperfine and nuclear quadrupole value point at the nickel ion [52]. Given the large number of adjustable parameters (five coupling parameters for each nitrogen), the error in the parameters is comparable to the relatively small difference between the nitrogen couplings, in particular in the g 1–g 2 plane. At the “single-crystal” observer position corresponding to g 3 (Fig. 2c, 1,098 mT), however, the resolution is significantly better and the simulation shows that there are three nitrogens with similar couplings [|A| = (24.0–26.8) MHz], and one with a slightly larger coupling [|A| = 30.4 MHz]. The optimized values are comparable with those given for the species MCRred1c [53], MCRBPS [14, 15], and MCRox1 [54], but vastly different from those of MCRred2r [53], where the four F430 hydropyrrolic nitrogen hyperfine interactions span a large range, indicating a significant asymmetry in the spin-density distribution (this species has a strong proximal sulfur ligand from HS-CoM). The outcome for MCRred1m is that the spin-density distribution on F430 is not significantly perturbed by the weak axial ligand.
a X-band CW EPR spectra of MCRred1m derived from experiment (upper trace in black) and a simulation (lower trace in red) calculated with the principal g values determined from the W-band EPR spectrum and including the four hydropyrrolic 14N hyperfine interactions. b Q-band (34.3678 GHz) matched HYSCORE spectrum measured at the intensity maximum of the EPR spectrum showing nitrogen double-quantum cross-peaks which indicate hyperfine couplings in the range from 23 to 33 MHz. c Q-band (34.5038 GHz) spectra. The field-swept EPR spectrum (20 K) is shown at the top; below this are Davies ENDOR spectra measured at 20 K at the observer positions indicated. The upper traces show the experimental spectra; the simulations consisting of the sum of signals from the four hydropyrrolic nitrogens are slightly shifted downwards from these. For the field position of 1,098 mT, the four nitrogen components are shown as dashed lines
Addition of isotope-labeled (13C/1H/2H) CH3-CoM
Using selectively isotope-labeled CH3-S-CoM, we have studied the hyperfine (and nuclear quadrupole) interactions of the substrate added to Ni(I)F430 when the MCRred1m state is formed.
13CH3-S-CoM
Carbon signals from 13CH3-S-CoM could be observed at X-band with Davies ENDOR (Fig. 3a) using a very selective inversion π mw pulse of length 800 ns, or at W-band using Mims ENDOR (Fig. 3c) with a long τ value of 300–500 ns. Data at both mw frequencies show a major component with 13C signals in the range ±(0.3–0.8) MHz, and much weaker signals in the range ±(0–0.5) MHz from a minor component. This can be best appreciated by inspecting Fig. 3b, which was measured with a long τ value (500 ns) to enhance the smaller splittings. The figure displays a main peak at ca. ±0.6 MHz, and a weaker one marked by diamonds at ca. ±0.2 MHz; the control experiment using unlabeled CH3-S-CoM showed no detectable carbon signals (green trace in Fig. 3b). A variable mixing time ENDOR experiment [33, 34] (see Fig. S1) performed at 10 K shows that the hyperfine interaction from the major component is negative. Note that the sign and the strength of the 13C hyperfine coupling is much smaller than those observed in compounds that feature a direct Ni–alkyl bond such as MCRBPS prepared from 13C-enriched 3-bromopropane sulfonate, MCRBrMe (prepared from methyl bromide), and MCRIMe (prepared from methyl iodide; note that MCRBrMe and MCRIMe denote the same species and are only named differently based on their preparation) [14–17].
13C ENDOR spectra from MCRred1m with 13CH3-S-CoM. a X-band (9.7249 GHz) Davies ENDOR spectra measured at 25 K at the observer positions indicated. The inversion π pulse in the mw sequence was 800 ns long (excitation width, fwhh ~0.7 MHz); simulations are shown as red lines. b, c W-band (94.274 GHz) Mims ENDOR spectra measured at 15 K with a τ of 300 ns unless indicated otherwise at the observer positions indicated. b is recorded at the observer position of the intensity maximum of the field-swept EPR spectrum and shows a trace from a 12CH3-S-CoM (green) and a 13CH3-S-CoM (blue) MCRred1m sample. Asterisks highlight methyl 13C signals which imply that a second distinct configuration of MCRred1m exists. In c, the red lines are the simulations for the methyl 13C nucleus, corrected for the τ-dependent blind spots by I(ν rf) = I sin(2π(ν rf − ν 1H)τ)2
A model containing two carbons with axial hyperfine interactions was investigated and found to give a satisfactory fit to the data (Table 2). Component 1: >90%, A 1 = −[1.3, 1.3, 0.8] MHz = −1.1 + [−0.16, −0.16, 0.33] MHz, linewidth 0.2 MHz; component 2: not well defined but has couplings in the range |A 2| = (0.0–0.5) MHz. The relatively large ENDOR linewidth of the major component (A 1) indicates a distribution of hyperfine interactions and thus that the methyl group has a certain degree of structural freedom. The hyperfine coupling A 1 has an isotropic part a iso 1 = −1.1 MHz which indicates that there is a Fermi contact interaction \( a_{\text{iso}} = \frac{{2\mu_{0} }}{3\hbar }g_{e} \beta_{e} g_{n} \beta_{n} \left| {\psi_{0} (0)} \right|^{2} , \) and thus that CH3-S-CoM is coordinated to NiIF430. Note that the contribution of the pseudo-isotropic interaction
is far too small to explain the observed a iso value [30]. Using [Δg 1 Δg 2 Δg 2] = [0.249, 0.069, 0.059] and the 13C principal values to give the dipolar part [T 1, T 2, T 3] = [−0.16, −0.16, 0.33] MHz gives an upper limit of |a Fiso | < 0.02 MHz. This value is far too small to explain the experimental value.
The 13C hyperfine interactions indicate that CH3-S-CoM can coordinate to F430 in most probably two different configurations with large degrees of freedom. The structural freedom leads to a relatively broad distribution in the couplings and thus broad ENDOR lines, modeled by a large linewidth. The structural freedom of the major component (>90%) may be the angle of the methyl group around the S–Cβ bond relative to F430 (i.e., the CH3 group pointing into and out of the plane of the paper in Structure 1). The minor component (<10%) could be due to a small percentage of CH3-S-CoM that is close to but not chemically bound to the nickel ion.
CH3-S-CoM and CD3-S-CoM
Deuterium signals from the methyl nuclei can be unambiguously identified with X-band HYSCORE with the substrate CD3-S-CoM, or with Q-band ENDOR by comparing the signals from samples with those from the substrates CH3-S-CoM and CD3-S-CoM. Figure 4a shows the low-frequency region of a HYSCORE spectrum measured at X-band, where a clear deuterium signal that is split by the nuclear quadrupole interaction is observed. The full set of HYSCORE spectra measured at five field positions is shown in Fig. S2. Simulations of these data show that the methyl proton hyperfine interactions are in the range |A(1H)| = (−0.5, −0.5, +1.0) MHz [A(1H) = A(2H) × 6.5144].
Methyl signals from CD3-S-CoM/CH3-S-CoM in MCRred1m. a 2H X-band (9.73 GHz) HYSCORE spectrum of the CD3-S-CoM sample measured at 12.5 K at the observer position near to the field value corresponding to g 3 (low-field edge) of the field-swept EPR spectrum (see inset). b 1H Q-band (34.515 GHz) Davies ENDOR spectra measured at 25 K at the observer positions of the intensity maximum (near g 1/g 2) and at the low-field position (near g 3) of the field-swept EPR spectrum. The proton frequency regions from the CH3-S-CoM sample (solid blue line), the CD3-S-CoM sample (dashed black line), and the difference between those two (dotted green line, not to scale), which highlights signals from the methyl protons of methyl-coenzyme M, are shown
This small deuterium hyperfine coupling determined by HYSCORE is consistent with the 1H Davies ENDOR data (Q-band, 34.515 GHz, 25 K) shown in Fig. 4b for observer positions corresponding to g 3 (low-field end) and g 1/g 2 (intensity maximum of the EPR spectrum). The spectra from the CH3-S-CoM sample (solid blue line) and the CD3-S-CoM sample (dashed black line) are shown at both field positions. The difference spectrum (dotted green line), which highlights signals from the methyl 1H nuclei, is also displayed. At the observer position corresponding to g 1/g 2, |A(1H)| ≈ 0.4 MHz, and at g 3, |A(1H)| ≈ 0.8 MHz.
An estimate of the nickel–methyl proton (electron–proton) distance can be obtained using the point–dipole model [55]
with
where r k is the distance between the unpaired electron and the kth nucleus with spin population ρ k , and R(α,β,γ) is the rotation matrix transforming the kth point–dipole interaction into the g-matrix principal axis system. The following spin populations were assumed: Ni 88%, each hydropyrrolic nitrogen 3% (this was estimated from the corresponding hyperfine couplings). If we assume that the methyl nuclei have a iso(1H) ≈ 0, then a lower limit for the shortest Ni–Hmethyl distance of r(Ni–Hmethyl) > 0.5 nm is obtained from Eqs. 3a and 3b with T = 0.6 MHz (A(1H) ≈ [−0.6, −0.6, 1.2] MHz). The nickel–methyl-carbon distance cannot be accurately estimated using just the point–dipole model without knowing the hyperfine anisotropy due to carbon p-orbital contributions (see the DFT data later). For a 13C distance of r(Ni–Cmethyl) > 0.5 nm, Eq. 2 predicts a small coupling of T < 0.16 MHz, in the range of the experimental coupling. These data imply that the methyl group points away from the nickel, as depicted in Structure 1. This methyl group orientation puts the largest principal axis of the hyperfine interaction (HI) of each methyl proton close to the orientation of the g 3 axis, consistent with the experimental data, which exhibit the largest splitting approximately along the g 3 direction. Note that, given the limited resolution of these 1H/2H data, it is not possible to determine whether one or more sets of methyl couplings contribute to the signals, as was the case for the 13C methyl data.
Glutamine and lactam 14N signals
Figure 5 shows X-band (9.7263 GHz) HYSCORE spectra recorded at the echo maximum of the field-swept EPR spectrum for (A) MCRred1a, (B) MCRred1m, and for comparison (C) free factor Ni(I)F430M [9] in THF/2-MeTHF 1:1. The intense cross-peaks at ca. (2.7, 3.3) MHz and (3.3, 2.7) MHz represent the two double-quantum (dq) signals from a weakly coupled 14N nucleus, and a comparison of the three spectra demonstrates that the weakly coupled nitrogen is very similar in MCRred1a and MCRred1m but nonexistent in free F430M. Simulation of the data (HYSCORE spectra recorded at middle- and low-field positions in the EPR spectrum are given in Fig. S3) resulted in the hyperfine coupling |A(14N)| ≈ [0.5, 0.5, 0.6] MHz and the nuclear quadrupole parameters |e 2 qQ/h| ≈ 2.6 MHz, η = 0.2. Since there is an isotropic contribution to the hyperfine interaction (HI), there is delocalization of the electron spin density onto this nitrogen, implying coordination of the structure to the paramagnetic center. There are two possible assignments: the NH nitrogen of the lactam ring of F430, or the NH2 nitrogen bound to the oxygen of Glnα′147, which is coordinated to the nickel from the distal face in the Ni(II) crystal structure. The measurement of the methylated free factor Ni(I)F430M can help to distinguish between the two possible assignments. Unlike MCR in the red1a or red1m states, the HYSCORE spectrum of F430M does not show the intense nitrogen peaks; instead there is a weak peak along the diagonal (indicating A ≈ 0 MHz) that is assigned to the lactam nitrogen. Simulations of this feature yield |A| < 0.2 MHz, |e 2 qQ/h| ≈ 2.8 MHz, η ≈ 0.7 (see also Fig. S4). The fact that the F430M sample does not contain Glnα′147 and that the lactam ring nitrogen signals are observed along the diagonal indicates that the intense 14N peaks in MCRred1a and MCRred1m should be assigned to the NH2 of the Glnα′147 residue. A comparison of the NQI data with those for model compounds delivers additional indications that this assignment is the correct one [56]. Glutamine and asparagine NH2 nitrogens have |e 2 qQ/h| ≈ 2.6–2.8 MHz, η ≈ 0.3–0.4 [57–59], whereas the NH nitrogens of histidine and proline have |e 2 qQ/h| ≈ 1.4–1.7 MHz, η = 0.6–1.0 [60], and the NH nitrogen of guanine has |e 2 qQ/ħ| = 2.63 MHz, η = 0.60. Our parameters most closely resemble those of a glutamine [61]. Additionally, the Glnα′147 residue coordinated to the nickel ion via the oxygen would be expected to have a small isotropic nitrogen HI. We thus assign this coupling to the NH2 of Glnα′147.
1H signals
The proton region of the X-band HYSCORE spectra shown in Fig. 6 allows the hyperfine interaction(s) from the proton(s) with the largest anisotropy to be estimated by making use of the frequency shift from the antidiagonal line marking the 1H resonance frequency. The maximum frequency shift Δν max behind the antidiagonal can be used to calculate the anisotropy of the hyperfine interaction,
where T is the dipolar term of an axial hyperfine matrix with principal values −T, −T, 2T. In Fig. 6, the signal is shifted a maximum of Δν max = 0.25 MHz behind the antidiagonal line, allowing an initial estimate of T ≈ 3.5 MHz. Further refinement by simulation allowed the principal values |A(1H)| ≈ [−3.2, −3.2, 6.4] MHz to be obtained, and the direction of the main axis (z-axis) with respect to g 3 to be estimated as β ≈ 35°. Using the point–dipolar model given in Eqs. 3a and 3b, the electron–proton distance is estimated to be approximately 0.3 nm. The orientation of the g 3-axis and the distance are in good agreement with what is expected for either the closest β proton of CH3-S-CoM or the closest proton from the amine nitrogen Glnα′147. In the Ni(II) crystal structure with HS-CoM and Glnα′147 coordinated, these two protons are 0.3 nm from the nickel, and their nickel–proton vectors are both inclined at β ≈ 45° to the plane of F430. In MCRred1a (Fig. 6a), these signals are assigned to the closest proton from Glnα′147, since there is no proximal ligand. In MCRred1m (Fig. 6b), the observed 1H signal probably has two overlapping contributions: the closest β proton of CH3-S-CoM and Glnα′147.
The strength of the HI with the β protons of methyl-coenzyme M can be compared to the corresponding interaction with the β proton of HS-CoM closest to the nickel in the two MCR species that have been shown to have the sulfur of CoM strongly coordinated to the nickel, namely MCRox1 {A(1H) = −2.0 + [−3.5, −4.6, 8.1] MHz} and MCRred2r {A(1H) = −7.8 + [−3.8, −3.8, 7.6] MHz} [54, 62]. In all three cases, the dipolar part of the HI is similar, indicating similar distances of the β-CH2 group from the nickel. In contrast to MCRox1 and MCRred2r, however, in MCRred1m the isotropic part of the HI is a iso ≈ 0. This can be explained by different spin densities on the coordinated sulfur. In MCRox1 and MCRred2r, these were estimated from the experimental 33S hyperfine coupling to be ρ = 6 ± 3% and ρ = 7–17%, respectively, resulting in a significant a iso on the β protons due to spin polarization. In the case of the MCRred1m presented here, the very weak coordination of the sulfur atom (and consequently the small electron spin density on the sulfur) leads to a negligibly small a iso on the β protons.
Coordination geometry of CH3-S-CoM in MCRred1m
From the set of measured hyperfine and nuclear quadrupole couplings, a picture of the coordination geometry of CH3-S-CoM in MCRred1m can be constructed. This picture is aided by density functional theory (DFT) calculations, which yield the isotropic and the dipolar parts of the HI between 13C on the labeled substrate and the Ni(I) center. These calculations were performed on the model system (in vacuum) shown in Fig. 7, which contains F430 (with hydrogen instead of side chains in positions 2, 4, 7, 8, 12, 13, 18) and CH3-S-CoM. Optimizing the geometry of this structure shows that there is a weak but positive interaction between the Ni(I) ion and the thioether sulfur amounting to −16.5 kJ/mol, with a long bond length of 0.394 nm. Figure 7 reveals that this interaction is electronic in nature since CH3-S-CoM has nonzero unpaired electron spin populations on sulfur (ρ = +0.16%) and the methyl carbon (ρ = −0.12%). The 13C-methyl group hyperfine coupling from DFT is A cal(13C) = −0.93 + [−0.08, −0.03, 0.10] MHz, [α,β,γ] = [22, 25, 141]°, which is in general agreement with the experimentally obtained couplings of A(13C) = −1.10 + [−0.16, −0.16, 0.33] MHz. According to DFT, the methyl-carbon spin density comprises an s-orbital contribution of −0.02%, and a −0.10% p-orbital contribution which corresponds to a 13C hyperfine interaction contribution of T p-orbital = [0.1, 0.1, −0.2] MHz (orientated approximately along the Cmethyl–S bond). The remaining anisotropy comes from interaction between the 13C nuclear spin moment and the spin density (distributed mainly on F430), given by T ρ = [−0.13, −0.13, 0.26] MHz.Footnote 1 Both contributions have different axis orientations and (significantly) an opposite sign, so, when summed together, the largest principal axis no longer points closely along the g 3 axis direction (see the Euler angles in Table 2). This is in contrast to the methyl protons described below with no p-orbital contribution. A close inspection of Fig. 3c shows that the largest 13C methyl splitting is not along a g value principal axis [which corresponds to the low (g ||) and high (g ⊥) ENDOR field positions].
Spin density plots (blue, positive; red, negative) of the DFT model for MCRred1m at the level of (a) 0.001 and (b) 0.0001. Note the small negative spin density of the methyl carbon of CH3-S-CoM. Selected bond lengths: Ni–S = 3.94 Å, Ni–NA = 2.13 Å, Ni–NB = 2.01 Å, Ni–NC = 2.07 Å, Ni–ND = 2.01 Å; unpaired electron spin populations: Ni = 90.5%, NA = 4.2%, NB = 4.1%, NC = 5.8%, ND = 4.2%, Sthioether = 0.16%, Cmethyl = −0.12%. Note that the spin population total sums to 100% with both positive and negative contributions due to spin polarization, as can be appreciated from the blue and red densities on the figure
There is a broad agreement between the largest methyl proton hyperfine couplings determined experimentally (Table 2, A ⊥ = −0.5 to −0.6, A || = 1.0–1.2 MHz) and by DFT (A(1H)max = [−0.4, −0.3, 0.82] MHz). The larger experimental value would suggest a slightly shorter distance than given by the ADF calculations. A point–dipole calculation using Eqs. 3a and 3b with the ADF model structure and total spin density distributed over the remaining 76 nuclei (see Table S2 for this input data) gives a good match to the experiment when the Ni–S bond is reduced from 0.394 nm to 0.345–0.375 nm (by moving CH3-S-CoM toward F430 along the Ni–S vector). The corresponding shortest methyl Ni–H bond distance is 0.535–0.565 nm, in agreement with the previous lower limit obtained using Eqs. 3a and 3b with experimental estimates of the spin populations [r(Ni–Hmethyl) > 0.5 nm].
The relatively weak bond and the long bond distance are consistent with solution studies showing that free Ni(I)F430M does not bind CH3-S-CoM, and the long distance explains why it could not be positively detected in XAS studies [23]. In the protein it would be expected that further complex stabilization occurs as a result of the binding of the negatively charged sulfonate group of CH3-S-CoM to a positively charged arginine side chain of the protein (as shown in the X-ray structures containing HS-CoM) and the steric constraints of the F430 environment, which help to place the substrate on the proximal side of F430 and directly above the Ni(I) ion.
How relevant is the weak Ni–S coordination in MCRred1m for the next steps in the catalytic cycle? The ordered two-substrate kinetics indicate that MCRred1m has a higher affinity for the binding of the second substrate HS-CoB than MCRred1a. Once HS-CoB is added to MCRred1m the catalytic process starts, and—so far—no further intermediates have been observed before the formation of the final products, methane and the heterodisulfide CoM-S–S-CoB. At first sight, the weak Ni–S coordination and the geometry of CH3-S-CoM with the methyl group pointing away from the Ni center reported here for MCRred1m would be consistent with the geometrical and stereoelectronic requirements of mechanism “A” or the newly proposed mechanism “C.” For an SN2-like attack of Ni(I) on the methyl carbon, as proposed in mechanism “B,” a rotation around the CH3S-CH2(β) bond of CH3-S-CoM, which turns the CH3-group towards the Ni with concomitant loss of the (weak) Ni–S coordination energy, would be required first.
Nevertheless, we hesitate to speculate about the implications of our findings on MCRred1m for the mechanism of the subsequent bond-breaking steps, because results (to be reported separately) we obtained for the MCRred2 species indicate that binding of the second substrate HS-CoB may induce a major structural change in the active site. The recent discovery of an MCR state that contains a Ni-hydride in its active site, which is formed upon the addition of HS-CoM and HS-CoB (called MCRred2a due to its almost axial g-tensor), and the existence of the MCRred2r state induced simultaneously (called MCRred2r due to its rhombic g-tensor) also suggest that larger structural rearrangements of the protein are possible when HS-CoB binds [62, 63].
Furthermore, the overall broad linewidths in the ENDOR spectra of Fig. 3 point toward a substantial degree of structural freedom on the proximal side of F430 in MCRred1m. Therefore, it cannot be excluded that the methyl group of CH3SCoM might reorient itself towards the nickel upon the binding of the second substrate.
Conclusions
As schematically shown in Fig. 8, the weak Ni–S coordination and the lower limit for the distance between Ni(I) and the S-methyl carbon determined in this study (along with the binding of the sulfonate group to the enzyme backbone [1]) determine the position of CH3-S-CoM in the active site of MCRred1m. The S-methyl group points away from the nickel and has a large degree of structural freedom, which can be deduced from the observation of broad linewidths in ENDOR spectra. Since the irreversible part of the catalytic reaction is triggered only after HS-CoB is added to the MCRred1m state (center of Fig. 8), this state can be considered the first and, so far, only observable intermediate in the cycle of MCR (right hand side of Fig. 8). In view of the indirect evidence for a major structural change in the active site upon the binding of HS-CoB and the substantial degree of freedom found for the S-methyl group in this study, the question of the binding geometry of CH3SCoM with respect to the nickel center in the transition state of the bond-breaking step remains open.
Schematic of the coordination geometry of CH3-S-CoM in MCRred1m as derived from our EPR spectroscopic and DFT results. Upon the addition of CH3-S-CoM to MCR in the MCRred1a state, a complex between the NiI-center of F430 in MCR and the thioether sulfur atom of the substrate is formed. Also shown are the two mechanisms “A” and “B” that have been proposed for the actual bond-breaking step which follows the addition of HS-CoB. Mechanism “C” proposed by Duin and McKee is not explicitly shown here
Notes
The dipolar part T ρ can be estimated by either performing a point–dipole calculation using Eqs. 3a and 3b and the 76 DFT spin populations given in Table S2, or by subtracting out the p-orbital contribution and assuming that it is orientated along the C–S bond. Both methods give essentially the same result.
Abbreviations
- CH3-S-CoM:
-
Methyl-coenzyme M
- ENDOR:
-
Electron nuclear double resonance
- EPR:
-
Electron paramagnetic resonance
- HS-CoB:
-
Coenzyme B
- HS-CoM:
-
Coenzyme M
- HYSCORE:
-
Hyperfine sublevel correlation spectroscopy
- MCR:
-
Methyl-coenzyme M reductase
- MCRBPS :
-
MCR after the addition of 3-bromopropane sulfonate
- MCRBrMe :
-
MCR after the addition of methyl bromide
- MCRIMe :
-
MCR after the addition of methyl iodide
- MCRox :
-
MCR exhibiting the ox1, ox2 or ox3 EPR spectra
- MCRred1 :
-
Active MCR exhibiting one of the red1 EPR spectra
- MCRred1a :
-
MCR-red1 in the absence of any substrates
- MCRred1c :
-
MCR-red1 in the presence of 10 mM coenzyme M
- MCRred1m :
-
MCR-red1 in the presence of 10 mM methyl-coenzyme M
- MCRred2 :
-
MCR exhibiting the two red2 EPR spectra
- MCRred2a :
-
MCR exhibiting the axial red2 EPR spectrum
- MCRred2r :
-
MCR exhibiting the rhombic red2 EPR spectrum
References
Thauer RK (1998) Microbiology 144:2377–2406
Thauer RK, Jungermann K, Decker K (1977) Bacteriol Rev 41:100–180
Ermler U, Grabarse W, Shima S, Goubeaud M, Thauer RK (1997) Science 278:1457–1462
Goubeaud M, Schreiner G, Thauer RK (1997) Eur J Biochem 243:110–114
Pelmenschikov V, Blomberg MRA, Siegbahn PEM, Crabtree RH (2002) J Am Chem Soc 124:4039
Pelmenschikov V, Siegbahn PEM (2003) J Biol Inorg Chem 8:653
Horng YC, Becker DF, Ragsdale SW (2001) Biochemistry 40:12875
Ghosh A, Wondimagegn T, Ryeng H (2001) Curr Opin Chem Biol 5:744
Jaun B, Pfaltz A (1986) J Chem Soc Chem Commun 1327
Grabarse W, Mahlert F, Duin EC, Goubeaud M, Shima S, Thauer RK, Lamzin V, Ermler U (2001) J Mol Biol 309:315
Goenrich M, Duin EC, Mahlert F, Thauer RK (2005) J Biol Inorg Chem 10:333–342
Duin EC, Signor L, Piskorski R, Mahlert F, Clay MD, Goenrich M, Thauer RK, Jaun B, Johnson MK (2004) J Biol Inorg Chem 9:563–576
Ermler U (2005) Dalton Trans 3451–3458
Hinderberger D, Piskorski RP, Goenrich M, Thauer RK, Schweiger A, Harmer J, Jaun B (2006) Angew Chem Int Ed 45:3602–3607
Kunz RC, Horng YC, Ragsdale SW (2006) J Biol Chem 281:34663–34676
Yang N, Reiher M, Wang M, Harmer J, Duin EC (2007) J Am Chem Soc 129:11028–11029
Dey M, Telser J, Kunz RC, Lees NS, Ragsdale SW, Hoffman BM (2007) J Am Chem Soc 129:11030–11032
Sauer A, Cohen H, Meyerstein D (1988) Inorg Chem 27:4578
Wondimagegn T, Ghosh A (2001) J Am Chem Soc 123:1543
Lin SK, Jaun B (1991) Helv Chim Acta 74:1725
Duin EC, McKee ML (2008) J Phys Chem B 112:2466–2482
Mahlert F, Grabarse W, Kahnt J, Thauer RK, Duin EC (2002) J Biol Inorg Chem 7:101–112
Duin EC, Cosper NJ, Mahlert F, Thauer RK, Scott RA (2003) J Biol Inorg Chem 8:141–148
Rospert S, Boecher R, Albracht SPJ, Thauer RK (1991) FEBS 291:371–375
Rospert S, Linder D, Ellermann J, Thauer RK (1990) Eur J Biochem 194:871–877
Bonacker LG, Baudner S, Mörschel E, Böcher R, Thauer RK (1993) J Biochem 217:587–595
Bradford MM (1976) Anal Biochem 72:248–254
Gunsalus RP, Romesser JA, Wolfe RS (1978) Biochemistry 17:2374–2377
Pfaltz A, Jaun B, Faessler A, Eschenmoser A, Jaenchen R, Gilles HH, Diekert G, Thauer RK (1982) Helv Chim Acta 65:828
Schweiger A, Jeschke G (2001) Principles of pulse electron paramagnetic resonance. Oxford University Press, Oxford
Gromov I, Shane J, Forrer J, Rakhmatoullin R, Rozentzwaig Yu, Schweiger A (2001) J Magn Reson 149:196–203
Davies ER (1974) Phys Lett A 47:1–2
Bennebroek MT, Schmidt J (1997) J Magn Reson 128:199–206
Epel B, Poppl A, Manikandan P, Vega S, Goldfarb D (2001) J Magn Reson 148:388–397
Höfer P, Grupp A, Nebenführ G, Mehring M (1986) Chem Phys Lett 132:279–282
Jeschke G, Rakhmatoullin R, Schweiger A (1998) J Magn Reson 131:261–271
Jeschke G, Schweiger A (1996) Mol Phys 88:355–383
Forrer J, Garcia-Rubio I, Schuhmann R, Tschaggelar R, Harmer J (2008) J Magn Reson 190:280–291
Stoll S, Schweiger A (2006) J Magn Reson 178:42–55 (see http://www.esr.ethz.ch)
Madi Z, Van Doorslaer S, Schweiger A (2002) J Magn Reson 154:181–191
Ahlrichs R, Bär M, Häser M, Horn H, Kölmel C (1989) Chem Phys Lett 162:165–169
Becke AD (1988) Phys Rev A 38:3098–3100
Perdew JP (1986) Phys Rev B 33:8822–8824
Universität Karlsruhe (2008) FTP directory /pub/jbasen at ftp://ftp.chemie.uni-karlsruhe.de/pub/jbasen. Last accessed 8th August 2008
Becke AD (1993) J Chem Phys 98:5648–5652
Lee C, Yang W, Parr RG (1988) Phys Rev B 37:785–789
Schäfer A, Huber C, Ahlrichs R (1994) J Chem Phys 100:5829
te Velde G, Bickelhaupt FM, van Gisbergen SJA, Fonseca Guerra C, Baerends EJ, Snijders JG, Ziegler T (2001) J Comput Chem 22:931–967
Hammer B, Hansen LB, Norskov JK (1999) Phys Rev B 59:7413
van Lenthe E, Baerends EJ, Snijders JG (1993) J Chem Phys 99:4597
Brown TG, Hoffman BM (1980) Mol Phys 39:1073–1109
Telser J, Horng YC, Becker DF, Hoffman BM, Ragsdale SW (2000) J Am Chem Soc 122:182–183
Finazzo C, Harmer J, Jaun B, Duin EC, Mahlert F, Thauer RK, Van Doorslaer S, Schweiger A (2003) J Biol Inorg Chem 8:586–593
Harmer J, Finazzo C, Piskorski R, Bauer C, Jaun B, Duin EC, Goenrich M, Thauer RK, Van Doorslaer S, Schweiger A (2005) J Am Chem Soc 127:17744
Pöppl A, Kevan L (1996) J Phys Chem 100:3387–3394
Grimaldi S, MacMillan F, Ostermann T, Ludwig B, Michel H, Prisner T (2001) Biochemistry 40:1037–1043
Lendzian F, Rautter J, Käss H, Gardiner A, Lubitz W (1996) Ber Bunsen-Ges Phys Chem 100:2036–2040
Spoyalov AP, Hulsebosch RJ, Shochat S, Gast P, Hoff A (1996) Chem Phys Lett 263:715–720
Deligiannakis Y, Hanley J, Rutherford AW (1999) J Am Chem Soc 121:7653–7664
Hanley J, Deligiannakis Y, MacMillan F, Bottin H, Rutherford AW (1997) Biochemistry 36:11543–11549
Garcia MLS, Smith JAS (1983) J Chem Soc Perkin Trans II 1401–1408
Harmer J, Finazzo C, Piskorski R, Ebner S, Duin EC, Goenrich M, Thauer RK, Reiher M, Schweiger A, Hinderberger D, Jaun B (2008) J Am Chem Soc 130:10907–10920
Kern DI, Goenrich M, Jaun B, Thauer RK, Harmer J, Hinderberger D (2007) J Biol Inorg Chem 12:1097–1105
Acknowledgments
We thank the Swiss National Science Foundation (SNF), the Fonds der Chemischen Industrie, and the EPSRC and BBSRC for financial support. Dariush Hinderberger gratefully acknowledges a research scholarship (HI 1094/1-1) from the Deutsche Forschungsgemeinschaft (DFG).
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Hinderberger, D., Ebner, S., Mayr, S. et al. Coordination and binding geometry of methyl-coenzyme M in the red1m state of methyl-coenzyme M reductase. J Biol Inorg Chem 13, 1275–1289 (2008). https://doi.org/10.1007/s00775-008-0417-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-008-0417-0