Abstract
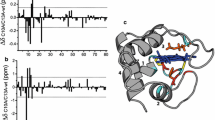
Hydrogen exchange rates for backbone amide protons of oxidized Pseudomonas aeruginosa cytochrome c-551 (P. aeruginosa cytochrome c) have been measured in the presence of low concentrations of the denaturant guanidine hydrochloride. Analysis of the data has allowed identification of submolecular unfolding units known as foldons. The highest-energy foldon bears similarity to the proposed folding intermediate for P. aeruginosa cytochrome c. Parallels are seen to the foldons of the structurally homologous horse cytochrome c, although the heme axial methionine-bearing loop has greater local stability in P. aeruginosa cytochrome c, in accord with previous folding studies. Regions of low local stability are observed to correspond with regions that interact with redox partners, providing a link between foldon properties and function.






Similar content being viewed by others
References
Winkler JR (2004) Curr Opin Chem Biol 8:169–174
Travaglini-Allocatelli C, Gianni S, Brunori M (2004) Trends Biochem Sci 29:535–541
Ptitsyn OB (1998) J Mol Biol 278:655–666
Russell BS, Melenkivitz R, Bren KL (2000) Proc Natl Acad Sci USA 97:8312–8317
Russell BS, Bren KL (2002) J Biol Inorg Chem 7:909–916
Hong XL, Dixon DW (1989) FEBS Lett 246:105–108
Rosell FI, Ferrer JC, Mauk AG (1998) J Am Chem Soc 120:11234–11245
Bartalesi L, Bertini I, Ghosh K, Rosato A, Turano P (2002) J Mol Biol 321:693–701
Yamamoto Y, Terui N, Tachiiri N, Minakawa K, Matsuo H, Kameda T, Hasegawa J, Sambongi Y, Uchiyama S, Kobayashi Y, Igarashi Y (2002) J Am Chem Soc 124:11574–11575
Baxter SM, Fetrow JS (1999) Biochemistry 38:4493–4503
Fetrow JS, Baxter SM (1999) Biochemistry 38:4480–4492
Bai YW, Sosnick TR, Mayne L, Englander SW (1995) Science 269:192–197
Russell BS, Zhong L, Bigotti MG, Cutruzzolà F, Bren KL (2003) J Biol Inorg Chem 8:156–166
Bartalesi I, Bertini I, Di Rocco G, Ranieri A, Rosato A, Vanarotti M, Vasos PR, Viezzoli MS (2004) J Biol Inorg Chem 9:600–608
Bartalesi I, Rosato A, Zhang W (2003) Biochemistry 42:10923–10930
Dobson CM, Sali A, Karplus M (1998) Angew Chem Int Ed Engl 37:868–893
Dill KA, Chan HS (1997) Nat Struct Biol 4:10–19
Dinner AR, Sali A, Smith LJ, Dobson CM, Karplus M (2000) Trends Biochem Sci 25:331–339
Matsuura Y, Takano T, Dickerson RE (1982) J Mol Biol 156:389–409
Bushnell GW, Louie GV, Brayer GD (1990) J Mol Biol 214:585–595
Kraulis PJ (1991) J Appl Crystallogr 24:946–950
Krishna MMG, Hoang L, Lin Y, Englander SW (2004) Methods 34:51–64
Weinkam P, Zong CH, Wolynes PG (2005) Proc Natl Acad Sci USA 102:12401–12406
Englander SW, Kallenbach NR (1983) Q Rev Biophys 16:521–655
Maity H, Lim WK, Rumbley JN, Englander SW (2003) Protein Sci 12:153–160
Maity H, Maity M, Englander SW (2004) J Mol Biol 343:223–233
Wen X, Bren KL (2005) Inorg Chem 44:8587–8593
Morar AS, Kakouras D, Young GB, Boyd J, Pielak GJ (1999) J Biol Inorg Chem 4:220–222
Pace CN, Scholtz JM (1997) In: Creighton TE (ed) Protein structure: a practical approach. IRL, Oxford, pp 299–321
Timkovich R, Cai ML (1993) Biochemistry 32:11516–11523
Hvidt A, Nielsen SO (1966) Adv Protein Chem 21:287–386
Bai YW, Milne JS, Mayne L, Englander SW (1993) Proteins 17:75–86
Zhang Y-Z (1995) University of Pennsylvania
Myers JK, Pace CN, Scholtz JM (1995) Protein Sci 4:2138–2148
Wen X, Patel KM, Russell BS, Bren KL (2007) Biochemistry 46:2537–2544
Bigotti MG, Allocatelli CT, Staniforth RA, Arese M, Cutruzzolà F, Brunori M (1998) FEBS Lett 425:385–390
Gianni S, Travaglini-Allocateli C, Cutruzzolà F, Brunori M, Shastry MCR, Roder H (2003) J Mol Biol 330:1145–1152
Borgia A, Bonivento D, Travaglini-Allocatelli C, Di Matteo A, Brunori M (2006) J Biol Chem 281:9331–9336
Gianni S, Brunori M, Travaglini-Allocatelli C (2001) Protein Sci 10:1685–1688
Gianni S, Travaglini-Allocatelli C, Cutruzzolà F, Bigotti MG, Brunori M (2001) J Mol Biol 309:1177–1187
Pelletier H, Kraut J (1992) Science 258:1748–1755
Pearl NM, Jacobson T, Arisa M, Vitello LB, Erman JE (2007) Biochemistry 46:8263–8272
Antalis TM, Palmer G (1982) J Biol Chem 257:6194–6206
Lappalainen P, Watmough NJ, Greenwood C, Saraste M (1995) Biochemistry 34:5824–5830
Cutruzzola F, Arese M, Ranghino G, van Pouderoyen G, Canters G, Brunori M (2002) J Inorg Biochem 88:353–361
Crowley PB, Ubbink M (2003) Acc Chem Res 36:723–730
Liang ZX, Nocek JM, Huang K, Hayes RT, Kurnikov IV, Beratan DN, Hoffman BM (2002) J Am Chem Soc 124:6849–6859
Crowley PB, Carrondo MA (2004) Proteins 55:603–612
Prudencio M, Ubbink M (2004) J Mol Recognit 17:524–539
Zhuravleva AV, Korzhnev DM, Kupce E, Arseniev AS, Billeter M, Orekhov VY (2004) J Mol Biol 342:1599–1611
Low DW, Gray HB, Duus JØ (1997) J Am Chem Soc 119:1–5
Michel LV, Ye T, Bowman SEJ, Levin BD, Hahn MA, Russell BS, Elliott SJ, Bren KL (2007) Biochemistry 46:11753–11760
Acknowledgments
This work is supported by NIH grant R01-GM63170.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Michel, L.V., Bren, K.L. Submolecular unfolding units of Pseudomonas aeruginosa cytochrome c-551. J Biol Inorg Chem 13, 837–845 (2008). https://doi.org/10.1007/s00775-008-0370-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-008-0370-y