Abstract
Pellino1 has been shown to regulate proinflammatory genes by activating the nuclear factor kappa B (NF-κB) and Toll-like receptor (TLR) signaling pathways, which are important in the pathological development of lipopolysaccharide (LPS)-induced myocarditis. However, it is still unknown whether silencing Pellino1 (si-Pellino1) has a therapeutic effect on this disease. Here, we showed that silencing Pellino1 can be a potential protective strategy for abnormal myocardial energy metabolism in LPS-induced myocarditis. We used liquid chromatography electrospray–ionization tandem mass spectrometry (LC–MS/MS) to analyze samples from si-Pellino1 neonatal rat cardiac myocytes (NRCMs) treated with LPS or left untreated. After normalization of the data, metabolite interaction analysis of matched KEGG pathway associations following si-Pellino1 treatment was applied, accompanied by interaction analysis of gene and metabolite associations after this treatment. Moreover, we used western blot (WB) and polymerase chain reaction (PCR) analyses to determine the expression of genes involved in regulating cardiac energy and energy metabolism in different groups. LC–MS-based metabolic profiling analysis demonstrated that si-Pellino1 treatment could alleviate or even reverse LPS-induced cellular damage by altering cardiomyocytes energy metabolism accompanied by changes in key genes (Cs, Cpt2, and Acadm) and metabolites (3-oxoocotanoyl-CoA, hydroxypyruvic acid, lauroyl-CoA, and NADPH) in NRCMs. Overall, our study unveiled the promising cardioprotective effect of silencing Pellino1 in LPS-induced myocarditis through fuel and energy metabolic regulation, which can also serve as biomarkers for this disease.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Sepsis is known to diminish oxidative metabolism in the heart and other tissues (Rudiger and Singer 2007). Myocarditis, caused mainly by bacterial or viral infections and autoimmune diseases (Hekimian and Combes 2017), is a multietiological pathological immune process involving myocardial inflammation with various clinical symptoms and outcomes, such as acute heart failure, chronic dilated cardiomyopathy, and even sudden cardiac death (Bracamonte-Baran and Cihakova 2017; Zhang et al. 2013; Sagar et al. 2012).
Notably, the occurrence and development of myocarditis involve various pathophysiological mechanisms of inflammation. The activation of multiple inflammatory regulators and related transcription factors can trigger the excessive inflammatory response, which is essential for the malignant development of cardiomyopathy (Maier et al. 2012; Akira et al. 2006). The inflammatory response and left-ventricular dysfunction induced by lipopolysaccharide (LPS) are mediated, in large part, by the host molecule Toll-like receptor 4 (TLR4) (Nemoto et al. 2002). TLR4 can trigger members of the nuclear factor kappa B (NF-κB) family and other transcription factors involved in the immune response and different cardiovascular diseases, such as ischemia–reperfusion injury and cardiac hypertrophy (Oyama et al. 2004; Timmers et al. 2008; Ha et al. 2005). Previous studies have reported variable effects of LPS on cardiac fuel and energy metabolism, suggesting that targeting inflammatory-metabolic “crosstalk” could improve outcomes in this syndrome.
The Pellino family interacts with Pelle/IRAK and has been identified to mediate the transcriptional regulation of proinflammatory genes in innate immunity through ubiquitination and then activate TLR and/or T-cell receptor (TCR) signaling (Grosshans et al. 1999b; Butler et al. 2007; Moynagh 2014). Pellino1, in particular, has been shown to play a dispensable role in activating the NF-κB family or other signal transduction pathways, which is important in T-cell activation and differentiation by the MyD88/TRIF-dependent TLR pathway (Chang et al. 2009; Kawai and Akira 2007; Choi et al. 2006; Vallabhapurapu and Karin 2009). A previous report has shown that the loss of Pellino1 can attenuate the activation of the TRIF-dependent NF-kB family due to the hyperactivation and nuclear accumulation of c-Rel in response to T-cell receptor-CD28 (TCR-CD28) signaling (Chang et al. 2011; Smith et al. 2011). In this study, we further explored whether silencing Pellino1 has therapeutic effects on LPS-induced myocarditis through multiple known or unknown regulatory mechanisms.
Currently, the immune response is the best-known underlying pathogenic mechanism and therapeutic target of bacterial myocarditis (Zhou and Yu 2017). However, the therapeutic effect is limited, and more strategies are needed to complement or replace this treatment (Kindermann et al. 2012). Since the close relationship between metabolism and signaling programs of immune cells may show a critical physiological regulation in normal or disease conditions by maintaining nutrient uptake and metabolic homeostasis (Andrejeva and Rathmell 2017), we used metabolomics to explore a new possible mechanism in the treatment of bacterial myocarditis.
In our study, we attempted to use LC–MS/MS to analyze the differences in metabolic compositions that could be vital in the occurrence and progression of LPS-induced myocarditis between samples exposed under different conditions. Thus, we could further investigate the therapeutic effect of silencing Pellino1 through metabolic mechanisms on LPS-induced myocarditis.
Materials and methods
Cell culture
NRCMs were prepared from 1-day-old Sprague–Dawley rats (Charles River) as previously described (Paradis et al. 2015). The cells were subsequently treated with 10 µg/mL LPS or PBS for 12 h. Adenovirus silencing Pellino1 (si-Pellino1, 2 × 107 pfu/mL) or adenoviral GFP was purchased from Shanghai GeneChem Co., Ltd. NRCMs were infected with adenovirus expressing GFP (Ad-GFP) or silencing Pellino1 (si-Pellino1) on day 1 after plating and on day 2 were treated with 10 µg/mL LPS or PBS for 12 h, after which mRNA was isolated for RT-PCR. The cells were lysed to obtain protein for western blot analysis.
Western blot analysis
NRCMs were lysed in RIPA buffer (P0013C, Beyotime) supplemented with 1 mM PMSF (ST505, Beyotime). Thirty micrograms of total protein were subjected to electrophoresis on 10% SDS-PAGE gels (Beyotime Biotechnology, China) and transferred to polyvinylidene fluoride (PVDF) (Merck-Millipore, Shanghai, China) membranes. After the samples were blocked with 5% milk powder in TBS-Tween for 1 h, the proteins were probed with primary antibody at 4 °C overnight. After the membranes were washed three times with TBS-Tween, they were incubated for 1 h with the corresponding secondary antibody conjugated to horseradish peroxidase and then subjected to enhanced chemiluminescence (ECL) for detection of protein bands. The primary antibodies used in this study were as follows: anti-CC3 (1:1000 dilution, #9664, Cell Signaling Technology), anti-C3 (1:1000 dilution, #9662, Cell Signaling Technology), anti-Bcl2 (1:1000 dilution, #3498, Cell Signaling Technology), anti-BAX (1:1000 dilution, #5023, Cell Signaling Technology), and anti-Pellino1 (1:200 dilution, sc-271065, Santa Cruz Biotechnology), and β-Actin (1:1000 dilution, #4967, Cell Signaling Technology) was used as a loading control (n = 3).
RNA analysis and real-time quantitative PCR (qRT-PCR)
Total cellular RNA isolation from NRCMs was performed using the RNazol B method and a Qiagen RNeasy kit, according to the manufacturer’s instructions. RNA was reverse transcribed (Applied Biosystems) using random hexamer priming. Real-time qRT-PCR was performed using SYBR Green reagent (Applied Biosystems) and rat-specific primers (Table S1) on the ABI Prism 7500 Sequence Detection system. GAPDH was used as an internal control. The relative gene expression levels were calculated using the 2 − △△Ct method (n = 3).
Flow cytometry
We incubated all collected viable and dead cells with propidium iodide (PI) and Annexin-V (Fcmacs Biotech Co., China) in the provided binding buffer at room temperature for 30 min in the dark, according to the manufacturer’s recommendations. Then, the flow cytometry method was applied to determine the relative cell apoptosis ratio (n = 3).
Metabolite extraction
Cultured NRCMs were quickly washed twice with ice-cold phosphate-buffered saline (PBS) in a cold room to remove medium components and then quickly rinsed with Milli-Q water. After the removal of water, 400 μL of 80% CH3OH/Milli-Q water was added to each culture plate. The cells were placed in liquid N2 for 10 min, thawed, and sonicated for 10 min. This process was repeated three times to completely extract the metabolites from the cells. Combined samples from the extraction process were then centrifuged at 10,000g for 5 min to pellet insoluble debris at 4 °C, with the supernatant transferred to microcentrifuge tubes and dried with a speed vacuum. Then, each dried sample was resuspended, vortexed, centrifuged, and transferred to autosampler vials for LC–MS analysis. Each group had six biological replicates.
LC–MS and data analysis
Separation of metabolites was performed by reversed-phase LC (HP1100, Agilent Technologies, Santa Clara, CA, USA) using a reversed-phase XBridge C18 column (1.7 μm particle size, 1 × 150 mm, Waters, Milford, MA, USA), and 8 μL of each sample was injected on the C18 column for analysis. Mobile phase A consisted of H2O/0.1% formic acid, and mobile phase B consisted of ACN with 0.1% formic acid with a program: 1% B at 0–1 min, 15% B at 3 min, 70% B at 5 min, 85% at 9 min, 100% at 10–12 min, and subsequently return to the initial conditions with 2 min for equilibration. Furthermore, the MS program was conducted on a 6545 Quadrupole-Time of Flight system (all devices from Agilent Technologies, Santa Clare, CA, United States) as follows: both positive- and negative-ion modes with drying gas 300℃ flow 6 L/min, sheath gas 340℃ flow 11 L/min, nebulizer gas 35 psig, capillary voltage 4000 V, and fragmental voltage 135 V. The data collection (MS 100–3200 m/z, MS/MS 30–3200 m/z) was acquired by both centroid and profile stored in autoMSMS scan mode with reference masses at m/z 112.05087 and 922.009798 were set as online accurate mass calibration.
MassHunter Workstation software (version B.07.00; Agilent Technologies) was used to export mzdata format from the acquired MS data (.d). Data analysis was performed by the XCMS Online at https://xcmsonline.scripps.edu/ using three steps: data upload, parameter selection, and result interpretation. A list of the intensities of all the peaks detected was generated using the retention time and the mass-to-ratio data pairs as the parameters for each ion. MS/MS spectra of the selected putative identifications were retrieved and matched with entries in the Metlin, Massbank, the Human Metabolome Database (HMDB). The metabolomics data resulting directly from XCMS Online were used to generate the cloud plot. The binning data were normalized to the total area. PCA, partial least-squares discriminant analysis (PLS-DA), orthogonal partial least-squares discriminant analysis (OPLS-DA), and metabolites and metabolic set enrichment analysis were performed with the web-based software MetaboAnalyst 3.0 (http://www.metaboanalyst.ca).
Statistical analyses
Data was presented as mean ± standard error of the mean (SEM). Using GraphPad Prism 4.0 (GraphPad software Inc., CA, USA), statistical significance among multiple groups was evaluated by one-way analysis of variance (ANOVA) with the Bonferroni post hoc test. A two-tailed P value < 0.05 was considered statistically significant.
Results
Silencing Pellino1 alleviates LPS-induced cardiomyocyte apoptosis and inflammation
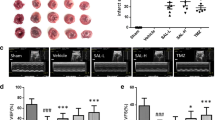
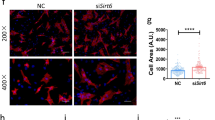
To verify the role of Pellino1 in LPS-induced septic cardiomyopathy, we first examined its expression in cardiomyocytes after LPS treatment. Western blot analysis showed that the expression of Pellino1 increased gradually with LPS treatment time (Fig. 1a, b). To further investigate whether silencing Pellino1 can ameliorate LPS-induced apoptosis, we used adenovirus inhibition of Pellino1 before LPS treatment, significantly reducing the Pellino1 expression levels in NRCMs (Fig. 1c, d). Flow cytometry showed that the LPS-induced increase in apoptosis was partially reversed by silencing Pellino1 (Fig. 1e, f). Besides, given the relationship between septic cardiomyocytes dysfunction and several apoptotic indexes, western blot analysis showed that the protective effect of silencing Pellino1 on septic cardiomyocytes dysfunction was mediated by reduced cleaved caspase-3 and Bax expression but not increased Bcl2 expression (Fig. 1g–i). In addition, LPS-induced inflammatory cytokine release was partially abolished by silencing Pellino1. Collectively, these data suggest that inhibition of Pellino1 attenuates LPS-induced cardiomyocytes dysfunction and promotes survival (Fig. 1j).
Silencing Pellino1 alleviates LPS-induced cardiomyocyte apoptosis and inflammation in vitro. a, b Representative western blot of the expression of Pellino1 in the LPS-induced cardiomyocytes. The bar graph shows that the expression of Pellino1 increased gradually with LPS treatment time. β-actin was detected as the loading control. c, d The efficacy of silencing Pellino1 via an adenovirus was assessed by western blotting. β-actin was detected as the loading control. e, f Apoptosis of cardiomyocytes exposed to LPS with or without the Pellino1-silencing adenovirus. The right bar graph shows the results of statistical analysis of the apoptosis ratios. g–i Representative western blot of the expression of Bax, Bcl2, CC3, and C3 in LPS-induced cardiomyocytes with/without si-Pellino1 treatment. The relative protein levels were normalized to the β-actin levels. The data are shown as the mean ± standard error of the mean of triplicates and are representative of three independent experiments performed. j Representative qRT-PCR of mRNA expression of TNF-α, IL-1β, and IL-6 in the LPS-induced cardiomyocytes with/without adenovirus silencing Pellino1. All mRNA expression was normalized to that of GAPDH. n = 6, results are expressed as the mean ± SD, *P < 0.05, **P < 0.01, ***P < 0.001, compared with the ctrl or GFP group
Silencing Pellino1 alleviates the effects of LPS on cardiac fuel and energy metabolism
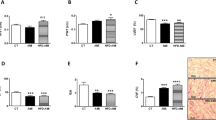
Given the regulatory role of PGC-1 coactivators in maintaining a high-capacity mitochondrial system, we next assessed the effect of silencing Pellino1 on the myocardial expression of LPS-induced cardiac PGC-1α and β genes. RT-PCR showed that LPS-mediated rapid downregulation of the PGC-1α and β mRNA levels was preserved in the Pellino1-silenced cells (Fig. 2a, b). The LPS-induced reduction of the PGC-1 coactivators that mediate metabolic transcription factors [PPARα, ERRα, NRF1 (Nrf1)] was partially reversed by silencing Pellino1 (Fig. 2c–e). However, silencing Pellino1 had no effect on the expression of PPARγ and PPARδ (Fig. 2c). Interestingly, the expression of downstream gene targets of PGC-1 coactivators involved in mitochondrial function [ATP Synthase β (Atp5b), Citrate Synthase (Cit syn)] and fatty acid β-oxidation (FAO) [MCPT-1 (Cpt1b), PDK4, MCAD (Acadm), CD36] was enhanced at the mRNA level following LPS treatment after silencing Pellino1 (Figs. 2f–g, 3a–d). Similar to the effects on the PGC-1 coactivator and their downstream gene targets, mRNA expression of GLUT4 (Slc2a4), the PGC-1 target gene encoding the glucose transporter, was also increased by silencing Pellino1 through adenovirus treatment (Fig. 3e). In addition, other genes involved in the regulation of FAO, such as PLIN2 and the myocardial lipid metabolic gene angiopoietin-like 4 (ANGPTL4), were not differentially expressed in the inflamed NRCMs and the si-Pellino1 group (Fig. 3f, g). Collectively, these data support the view that LPS-mediated inflammation leads to rapid deactivation of the PGC-1 metabolic gene regulatory circuit, and Pellino1 may be a potential target for cardiac fuel and energy metabolism.
Silencing Pellino1 alleviates the effects of LPS on cardiac mitochondrial function. a–e Representative qRT-PCR of PGC-1α, PGC-1β, PPARα, PPARγ, PPARδ, NRF1, and ERRα mRNA expression following LPS treatment after Pellino1 silencing. f, g qRT-PCR validation of downstream metabolic genes of PGC-1 coactivators, including ATP Synthase β (Atp5b) and Citrate Synthase (Cit syn), involved in mitochondrial function. All mRNA expression was normalized to that of GAPDH. n = 6, results are expressed as the mean ± SD, *P < 0.05, **P < 0.01, ***P < 0.001, compared with the GFP group
Silencing Pellino1 alleviates the effects of LPS on cardiac fuel metabolism. a–d qRT-PCR validation of downstream metabolic genes of PGC-1 coactivators, including MCPT-1 (Cpt1b), PDK4, MCAD (Acadm), and CD36 involved in fatty acid β-oxidation (FAO). e Representative qRT-PCR of the mRNA expression of GLUT4 (Slc2a4), the PGC-1 target gene encoding the glucose transporter, followed by Pellino1-silencing adenovirus treatment. f, g Representative qRT-PCR of the mRNA expression of PLIN2 and angiopoietin-like 4 (ANGPTL4) involved in the regulation of FAO and myocardial lipid metabolism, respectively, following LPS treatment after Pellino1 silencing. All mRNA expression was normalized to GAPDH. n = 6, results are expressed as the mean ± SD, *P < 0.05, **P < 0.01, ***P < 0.001, compared with the GFP group
LC–MS-mediated metabolomics analysis of LPS-treated or Pellino1-silenced NRCMs
To identify metabolite alterations that are induced by LPS or silencing Pellino1 adenovirus, we used LC–MS to analyze and evaluate all samples. First, the total ion flow diagram of the analyzed QC samples was compared with the spectral overlap by a comparison of the QC sample spectrogram. The results showed that the response strength and retention time of each chromatographic peak overlapped, indicating that the variation caused by instrument error was small in the whole experiment (Fig. S1A). Then, metabolic profiling (Chenomx, Edmonton, AB, Canada) was used to identify 169 differentially abundant metabolites between the LPS-treated cells and the untreated control cells (Table 1). After normalization of the data, univariate and multivariate statistical analyses were used to comprehensively evaluate the effects of silencing Pellino1 on the LPS-treated NRCMs (Figs. S1B–E, S2A–D). Our correlation analysis identified good sequencing stability and negative associations among the samples and metabolites (Fig. 4a).
LC–MS-mediated metabolomics analysis of LPS-treated or Pellino1-silenced NRCMs. a Correlation analysis identified good sequencing stability and negative associations among samples. b Different perspectives of two-dimensional principal components (PC) showed the variable reduction outputs analyzed by orthogonal partial least-squares projections to latent structures-discriminant analyses (OPLS-DA) on metabolites from si-Pellino1 and LPS-treated, LPS-treated, and untreated control NRCMs. c, d OPLS-DA score plots of three groups (GFP, GFP + LPS, si-Pellino1 + LPS) according to the differential metabolic profile showed in 2D (c) and 3D (d); each point represents a tested sample. e One-way ANOVA (analysis of variation) of significant levels of all 169 metabolites, P < 0.05
As a multivariate statistical technique applied to determine the relative differences in two or more systems that are large and complex, principal component analysis (PCA) is the most widespread multidimensional data analysis technique, which can systematically reduce the dimensionality of original data required to describe protein dynamics with the preservation of variance. Therefore, to further calculate variable importance in projection (VIP) scores of metabolites, we performed partial least-squares projections for latent structures-discriminant analysis (PLS-DA) of metabolites induced by si-Pellino1 and LPS-treated, LPS-treated, and untreated control NRCMs. Metabolites with VIP scores greater than 1.0 were considered important. To confirm the “goodness” of the model and the predictive quality, we performed orthogonal partial least-squares projections to latent structures-discriminant analyses (OPLS-DA) on data from three groups.
The variable reduction outputs were shown from different perspectives of two-dimensional principal components (PC) contributing a percentage of the variation of a sample (Fig. 4b). As shown in Fig. 4c, d, each point represents a tested sample, and the three groups were clustered separately, suggesting that the metabolic profile was intensely influenced by the treatment with LPS and si-Pellino1. Additionally, the obvious separation among samples from different treatment groups observed in the OPLS-DA score plots according to the differential metabolic profile in each group in both positive and negative ion modes indicated that si-Pellino1 treatment led to significant changes in the metabolites compared with those of the LPS-treated control group. Interestingly, as shown in Table 1, the high − log10(P values) in the OPLS-DA shown in the table could reflect the differences among the three groups of samples. Furthermore, one-way ANOVA of the metabolites showed significant levels of all 169 metabolites (Fig. 4e).
Key metabolic pathways associated with si-Pellino1 treatment
Among the distinguishing metabolites identified by statistical correlations, to further identify which metabolic pathways were the most relevant pathways affected by si-Pellino1 treatment, we generated a metabolome view of the top 50 matched pathways according to P values (Table 2) from enrichment analysis and impact values from topology analysis, as shown in Fig. 5a, b. The topological score (X-axis) and the size of each circle represent the importance of different metabolites and their impact values in the enriched pathways, while the color of each circle based on the P value of the enrichment analysis (Y-axis) represents the statistical significance of the overall metabolic changes in the pathways. The pathway from yellow to red indicates a growing impact (Table 3). From this analysis, the citrate cycle (TCA cycle); fatty acid metabolism; glycolysis or gluconeogenesis; fatty acid elongation in mitochondria; alanine, aspartate, and glutamate metabolism; and glyoxylate and dicarboxylate metabolism were identified as the most perturbed metabolic pathways according to − log(P values) after si-Pellino1 treatment in the LPS-induced NRCMs. Protein interaction analysis found that multiple significantly enriched disturbed metabolic pathways were markedly related to the adipocytokine signaling pathway and PPAR signaling pathway (Fig. 6a, b).
Metabolome view of the top 50 matched metabolic pathways associated with si-Pellino1 treatment. a Impact analysis of metabolites associated with si-Pellino1 treatment relative to the control according to P values (shown in Table 2) from enrichment analysis and impact values from topology analysis. The topological score (X-axis) and the size of each circle represent the importance of different metabolites and their impact values in the enriched pathways, while the color of each circle based on the P value of the enrichment analysis (Y-axis) represents the statistical significance of the overall metabolic changes in the pathways. b Enrichment analysis of metabolites associated with si-Pellino1 treatment relative to the control. The pathway from yellow to red indicates a growing impact
Protein interaction analysis of matched KEGG pathway associations with siPellino1 treatment. a Predicted protein interactions among PGC-1α, PPARα, Slc2a1, CD36, and Cpt1b involved in the adipocytokine signaling pathway. b Predicted protein interactions among PPARγ, PPARα, Angptl4, CD36, Cpt1b, and Acadm involved in the PPAR signaling pathway
To investigate the relationships between metabolites and genes that we previously explored in the main enriched metabolic pathways showing alterations, we generated specific correlation networks of the citrate cycle (TCA cycle), fatty acid metabolism, glyoxylate, and dicarboxylate metabolism and propanoate metabolism, as well as valine leucine and isoleucine degradation and beta-alanine metabolism, as shown in Fig. 7. The results shown above verified that si-Pellino1 treatment could alleviate or even reverse the LPS-induced cellular damage by altering cardiac fuel and energy metabolism accompanied by changes in key genes (Cs, Cpt2, and Acadm) and metabolites (3-oxoocotanoyl-CoA, hydroxypyruvic acid, lauroyl-CoA, and NADPH).
Interaction analysis of gene and metabolite associations with si-Pellino1 treatment. Correlation networks of the citrate cycle (TCA cycle); fatty acid metabolism; glyoxylate and dicarboxylate metabolism; propanoate metabolism; valine, leucine, and isoleucine degradation and beta-alanine metabolism showed the inner relationships between metabolites and genes that we previously explored in the main enriched metabolic pathways showing changes
Discussion
Bacterial myocarditis is a common clinical sepsis syndrome caused mainly by inflammation, which can contribute to adverse cardiovascular outcomes, end-organ dysfunction, and even mortality (Li and Sun 2007; Schauvliege et al. 2007).
LPS-trigged excessive host responses have been established to result in systemic inflammatory conditions, including myocarditis and sepsis, mainly by activating the Toll-like receptor (TLR) family, the major pattern-recognition receptors (PRRs) family of host cells (O’Neill et al. 2009; Akira et al. 2006), which fuels our interest in exploring the signaling function of TLRs after LPS stimulation. Previous studies have shown that TLRs-activated signal transduction pathways which determine the vital role of TLRs in inflammatory disorders depend on the binding between their intracellular domain (called TIR domain) and intracellular adaptors, including myeloid differentiation factor 88 (MyD88), Mal (TIRAP), TRIF (TICAM-1), and TRAM (TICAM-2) (Kawai and Akira 2007; Dunne and O’Neill 2003). Upon LPS stimulation, the phosphorylation cascade of MyD88 initiated by TLRs mediates the recruitment and activation of IL-1R-associated kinases (IRAKs), further triggering the activation of the nuclear factor-κB (NF-κB) family, resulting in the production of pro-inflammatory mediators helping to shape the immune response and eradicate infection (Janssens and Beyaert 2003).
As an intrinsic ubiquitin E3 ligase responsible for receptor-interacting protein 1 (RIP1) ubiquitination in the TLR/IL-1R signaling, Pellino is an evolutionarily conserved scaffold protein that could catalyze the polyubiquitylation of key molecule involved in the tumor necrosis factor receptor-associated factor (TRAF)-dependent TLR signaling pathway by interacting with its highly homologous family, IRAKs, in an MyD88-dependent manner after stimulation in mammals (Grosshans et al. 1999a; Schauvliege et al. 2007).
Pellino1, a mammalian counterpart of Pellino, have been recognized to be highly required for maintaining TLR-mediated NF-κB nuclear translocation and binding activity due to its value in forming Pellino1–IRAK1–IRAK4–TRAF6 signaling complex after the induction of inflammatory response (Moynagh 2009; Jiang et al. 2003), indicating the causal relationship between Pellino1 and TLR-mediated inflammatory response. In addition to TRAF6, a recent study found that the activation of TIR-domain-containing adapter-inducing interferon-β (TRIF)-dependent signaling induced-by Pellino1-mediated ubiquitination of RIP1 also contributed to the upregulation of NF-κB (Sato et al. 2003; Chang et al. 2009). Moreover, Pellino1 deficiency may interfere with the ubiquitination of RIP1 and TRAF6 which are needed for TLR3/4-mediated pro-inflammatory gene induction, thereby increasing the resistance of mice to LPS-induced lethality (Chang et al. 2009; Skaug et al. 2009). Collectively, the importance of Pellino1 in LPS-induced inflammatory signaling makes it reasonable to assume that suppressing Pellino1 do be especially effective in attenuating LPS-induced myocarditis.
As the unifying link between heart and cardiac function, changed cardiac metabolism has been evidenced to be causative in cardiomyocytes dysfunction induced by various physiological and pathological causes, including exercise and pressure overload (Gibb and Hill 2018). During infections, cell-intrinsic metabolism is no longer considered as a physical process that meets the basic demand of energy for heart contraction (Newsholme et al. 1986). In addition to supporting tissue homeostasis, pathogen-activated reprogramming of metabolic pathways also contributes to the production of metabolic intermediates and end products (Williams and O’Neill 2018). The immune-metabolic framework consisted of kinds of regulatory metabolites that play a vital role in regulating cardiac function, and determining the fate of an infection (Chauhan and Saha 2018).
Recent advances in LC–MS/MS-based omics uncovered that changed metabolites under pathological states could lead to the progression of inflammation (Ishihara et al. 2019). It has been shown that myeloperoxidase (MPO) could mediate primarily host defense reactions by regulating peroxidation of polyunsaturated fatty acids which is the characteristic feature of inflammation during acute inflammation (Winterbourn et al. 2000; Kubala et al. 2010). The withdrawal of the synthesis of bioactive metabolites of arachidonic acid (AA) and linoleic acid (LA) metabolism may ameliorate the severity of sepsis by modifying inflammatory-related cellular signaling pathways (Cook 2005; Nauseef 2001). Besides, the l-Tryptophan (Trp)–kynurenine (Kyn) pathway, which is essential for protein synthesis, was reported to improve acute viral myocarditis (Hoshi et al. 2012).
Metabolic remodeling after LPS stimulation may modify the downstream genes and proteins by triggering the mammalian target of rapamycin (mTOR), AMP-activated protein kinase (AMPK), and nuclear factor-κB (NF-κB) signaling pathways involved in many pathological conditions, such as atherosclerosis, indicating that it is necessary to emphasize the importance of metabolites in LPS-induced pro-inflammatory response (Byles et al. 2013; Sag et al. 2008; Wu et al. 2020). As the main source of energy for cardiac mechanical work, fatty acid (FA) can be transported by fatty acid translocase (FAT/CD36) and other transport proteins from the cytosol to the mitochondria and then form acyl-CoA to enter β-oxidation (van der Vusse et al. 2000; Berndt et al. 2019; Girones et al. 2014). The peroxisome proliferator-activated receptor subfamily is vital in the epigenetic regulation of FA utilization. It has been reported that the main three members of the PPAR nuclear receptor subfamily, PPARα, PPARδ, and PPARγ, are all involved in synergistic modulation of cardiomyocytes energy metabolism (Lopaschuk et al. 2010). Furthermore, in in vitro models of mice, the activation of PPARα has been identified to increase the expression of downstream genes, including CD36, MCPT1, and MCAD, which play different important roles in cardiac FA utilization (Lopaschuk et al. 2010; Schoonjans et al. 1996; Mistry and Cresci 2010; Huss and Kelly 2004). The induced transcriptional activities of both PPARs and ERRs could be coactivated by binding to PGC-1α (Huss et al. 2004; Schreiber et al. 2004). Recently, the PGC-1α/PPARα and PGC-1α/ERRα pathways are involved in regulating cardiac mitochondrial fatty acid oxidation genes (Lehman et al. 2000).
PGC-1α, an important regulator of mitochondrial activity and fatty acid oxidative metabolism, is enriched in the heart, which has high oxidative metabolic rates (Lehman et al. 2000; Lin et al. 2002; Knutti and Kralli 2001). Compared to those of PGC-1β, which is also confirmed to be necessary for aerobic mitochondrial respiration, post-translational modifications of PGC-1α bound with transcriptional partners could better control complex cardiac cellular energy metabolism by activating downstream endogenous gene promoters, including MCAD (St-Pierre et al. 2003; Rowe et al. 2010; Huss et al. 2002). Interestingly, in our study, silencing Pellino1 reversed the LPS-induced decrease in the expression of transcription factors and regulators in NRCMs, showing that Pellino1 plays a role in regulating fatty acid metabolism.
After analyzing the metabolome data, we found 169 unique metabolites that differed significantly in si-Pellino1 and LPS-treated, LPS-treated, and untreated control NRCMs. Then, the TCA cycle, fatty acid metabolism, and glycolysis or gluconeogenesis, along with fatty acid elongation in mitochondria and alanine, aspartate and glutamate metabolism, as well as glyoxylate and dicarboxylate metabolism, were identified as the most perturbed metabolic pathways according to P values from enrichment analysis and impact values from topology analysis.
Moreover, lipopolysaccharide has been reported to regulate the myocardial expression of the FAO enzyme and oxidation rates by deactivating the kinetic response of the PGC-1 axis, which is associated with systemic inflammation and sepsis-induced by LPS stimulation (Beutler and Rietschel 2003; Rossi et al. 2007; Puigserver et al. 2001; Feingold et al. 2004). Our results found that LPS altered cardiac fuel and energy metabolism accompanied by changes in key genes (Cs, Cpt2, and Acadm) and metabolites (3-oxoocotanol-CoA, hydroxypyruvic acid, lauroyl-CoA, and NADPH) in NRCMs. Furthermore, for the first time, we confirmed that Pellino1, which has been identified to activate NF-κB and TLR signaling pathways by mediating proinflammatory genes in innate immunity (Chang et al. 2009; Kawai and Akira 2007; Choi et al. 2006; Vallabhapurapu and Karin 2009), could partly reverse or restore the disturbed cardiomyocytes energy metabolism through mechanisms involving glycolysis, lipid metabolism, citrate cycle, and some other types of metabolism after LPS stimulation.
Glucose utilization is also highly linked to various physiological processes, including cardiomyocytes energy metabolism, which can affect mitochondrial function by coregulation with FA utilization (Randle 1998). GLUT1 and GLUT4, the common and major glucose transporters of glycogen-derived glucose in the heart, play a basic and important role in basal myocardial glucose uptake, suggesting that the decreased expression of GLUT4 may be harmful to cardiac glucose metabolism (Henning et al. 1996; Aerni-Flessner et al. 2012). It has been reported that pathological cardiac hypertrophy could accelerate glycolysis (Allard et al. 1994; Cheng et al. 2017).
In our study, si-Pellino1 alleviated the LPS-induced decrease in GLUT4 expression in NRCMs. The observed changes in two pivotal intermediates, succinic acid, and malic acid, were partly reversed after si-Pellino1 treatment, suggesting that silencing Pellino1 in LPS-induced myocarditis improved LPS-induced glucose metabolic dysfunction.
The metabolic profiles analyzed in our study could partly explain the therapeutic value of silencing Pellino1 in LPS-induced myocarditis. However, this report has some limitations that need to be considered. In our study, the samples were extracted from NRCMs cultivated in vitro. As a result, given the different metabolic environments, we lack experimental data from in vivo studies. Additionally, targeted metabolomics is an optimal method to identify the differences in metabolites. Consequently, these issues remain to be examined in the future.
Thus, the LC–MS/MS metabonomic method shed new light on the mechanism of the pathological development of bacterial myocarditis and was also used to assess the efficacy of si-Pellino1 treatment. Importantly, the altered metabolites associated with the TCA cycle, fatty acid metabolism, and glycolytic metabolism may be used as potential specific biomarkers corresponding to LPS-induced myocarditis. Moreover, combined with restoration of the abnormal metabolism and disturbed expression of transcription factors involved in cardiac fuel and energy metabolism, silencing Pellino1 showed cardioprotective value, revealing that silencing Pellino1 could be a promising therapeutic strategy against LPS-induced myocarditis or other pathologies in future clinical trials. In addition, considering the pro-inflammatory role of Pellino1 reported in the sepsis-induced lung injury model and COPD-induced inflammatory airway responses, silencing Pellino1 can also ameliorate acute or chronic lung pneumonia (Liu et al. 2021; Marsh et al. 2020; Hughes et al. 2019). Other than the heart and lung, the regulatory effects of Pellino1 on TLR/IL-1R signaling make it reasonable to assume that silencing Pellino1 may be targeted therapeutically in different inflammatory disorders associated with TLR/IL-1R signaling, which remains to be further explored in the future.
References
Aerni-Flessner L, Abi-Jaoude M, Koenig A, Payne M, Hruz PW (2012) GLUT4, GLUT1, and GLUT8 are the dominant GLUT transcripts expressed in the murine left ventricle. Cardiovasc Diabetol 11:63. https://doi.org/10.1186/1475-2840-11-63
Akira S, Uematsu S, Takeuchi O (2006) Pathogen recognition and innate immunity. Cell 124(4):783–801. https://doi.org/10.1016/j.cell.2006.02.015
Allard MF, Schonekess BO, Henning SL, English DR, Lopaschuk GD (1994) Contribution of oxidative metabolism and glycolysis to ATP production in hypertrophied hearts. Am J Physiol 267(2 Pt 2):H742-750. https://doi.org/10.1152/ajpheart.1994.267.2.H742
Andrejeva G, Rathmell JC (2017) Similarities and distinctions of cancer and immune metabolism in inflammation and tumors. Cell Metab 26(1):49–70. https://doi.org/10.1016/j.cmet.2017.06.004
Berndt N, Eckstein J, Heucke N, Gajowski R, Stockmann M, Meierhofer D, Holzhutter HG (2019) Characterization of lipid and lipid droplet metabolism in human HCC. Cells. https://doi.org/10.3390/cells8050512
Beutler B, Rietschel ET (2003) Innate immune sensing and its roots: the story of endotoxin. Nat Rev Immunol 3(2):169–176. https://doi.org/10.1038/nri1004
Bracamonte-Baran W, Cihakova D (2017) Cardiac autoimmunity: myocarditis. Adv Exp Med Biol 1003:187–221. https://doi.org/10.1007/978-3-319-57613-8_10
Butler MP, Hanly JA, Moynagh PN (2007) Kinase-active interleukin-1 receptor-associated kinases promote polyubiquitination and degradation of the Pellino family: direct evidence for PELLINO proteins being ubiquitin-protein isopeptide ligases. J Biol Chem 282(41):29729–29737. https://doi.org/10.1074/jbc.M704558200
Chang M, Jin W, Sun SC (2009) Peli1 facilitates TRIF-dependent Toll-like receptor signaling and proinflammatory cytokine production. Nat Immunol 10(10):1089–1095. https://doi.org/10.1038/ni.1777
Chang M, Jin W, Chang JH, Xiao Y, Brittain GC, Yu J, Zhou X, Wang YH, Cheng X, Li P, Rabinovich BA, Hwu P, Sun SC (2011) The ubiquitin ligase Peli1 negatively regulates T cell activation and prevents autoimmunity. Nat Immunol 12(10):1002–1009. https://doi.org/10.1038/ni.2090
Chauhan P, Saha B (2018) Metabolic regulation of infection and inflammation. Cytokine 112:1–11. https://doi.org/10.1016/j.cyto.2018.11.016
Cheng S, Shah SH, Corwin EJ, Fiehn O, Fitzgerald RL, Gerszten RE, Illig T, Rhee EP, Srinivas PR, Wang TJ, Jain M, American Heart Association Council on Functional Genomics and Translational Biology, Council on Cardiovascular and Stroke Nursing, Council on Clinical Cardiology, Stroke Council (2017) Potential impact and study considerations of metabolomics in cardiovascular health and disease: a scientific statement from the American heart association. Circ Cardiovasc Genet. https://doi.org/10.1161/HCG.0000000000000032
Choi KC, Lee YS, Lim S, Choi HK, Lee CH, Lee EK, Hong S, Kim IH, Kim SJ, Park SH (2006) Smad6 negatively regulates interleukin 1-receptor-Toll-like receptor signaling through direct interaction with the adaptor Pellino-1. Nat Immunol 7(10):1057–1065. https://doi.org/10.1038/ni1383
Cook JA (2005) Eicosanoids. Crit Care Med 33(12 Suppl):S488-491. https://doi.org/10.1097/01.ccm.0000196028.19746.42
Dunne A (2003) O’Neill LA (2003) The interleukin-1 receptor/Toll-like receptor superfamily: signal transduction during inflammation and host defense. Sci STKE 171:re3. https://doi.org/10.1126/stke.2003.171.re3
Feingold K, Kim MS, Shigenaga J, Moser A, Grunfeld C (2004) Altered expression of nuclear hormone receptors and coactivators in mouse heart during the acute-phase response. Am J Physiol Endocrinol Metab 286(2):E201-207. https://doi.org/10.1152/ajpendo.00205.2003
Gibb AA, Hill BG (2018) Metabolic coordination of physiological and pathological cardiac remodeling. Circ Res 123(1):107–128. https://doi.org/10.1161/circresaha.118.312017
Girones N, Carbajosa S, Guerrero NA, Poveda C, Chillon-Marinas C, Fresno M (2014) Global metabolomic profiling of acute myocarditis caused by Trypanosoma cruzi infection. PLoS Negl Trop Dis 8(11):e3337. https://doi.org/10.1371/journal.pntd.0003337
Grosshans J, Schnorrer F, Nüsslein-Volhard C (1999a) Oligomerisation of tube and pelle leads to nuclear localisation of dorsal. Mech Dev 81(1–2):127–138. https://doi.org/10.1016/s0925-4773(98)00236-6
Grosshans J, Schnorrer F, Nusslein-Volhard C (1999b) Oligomerisation of tube and pelle leads to nuclear localisation of dorsal. Mech Dev 81(1–2):127–138. https://doi.org/10.1016/s0925-4773(98)00236-6
Ha T, Li Y, Hua F, Ma J, Gao X, Kelley J, Zhao A, Haddad GE, Williams DL, Browder IW, Kao RL, Li C (2005) Reduced cardiac hypertrophy in toll-like receptor 4-deficient mice following pressure overload. Cardiovasc Res 68(2):224–234. https://doi.org/10.1016/j.cardiores.2005.05.025
Hekimian G, Combes A (2017) Myocarditis. Rev Med Intern 38(8):531–538. https://doi.org/10.1016/j.revmed.2016.12.022
Henning SL, Wambolt RB, Schonekess BO, Lopaschuk GD, Allard MF (1996) Contribution of glycogen to aerobic myocardial glucose utilization. Circulation 93(8):1549–1555. https://doi.org/10.1161/01.cir.93.8.1549
Hoshi M, Matsumoto K, Ito H, Ohtaki H, Arioka Y, Osawa Y, Yamamoto Y, Matsunami H, Hara A, Seishima M, Saito K (2012) L-tryptophan-kynurenine pathway metabolites regulate type I IFNs of acute viral myocarditis in mice. J Immunol 188(8):3980–3987. https://doi.org/10.4049/jimmunol.1100997
Hughes BM, Burton CS, Reese A, Jabeen MF, Wright C, Willis J, Khoshaein N, Marsh EK, Peachell P, Sun SC, Dockrell DH, Marriott HM, Sabroe I, Condliffe AM, Prince LR (2019) Pellino-1 regulates immune responses to haemophilus influenzae in models of inflammatory lung disease. Front Immunol 10:1721. https://doi.org/10.3389/fimmu.2019.01721
Huss JM, Kelly DP (2004) Nuclear receptor signaling and cardiac energetics. Circ Res 95(6):568–578. https://doi.org/10.1161/01.RES.0000141774.29937.e3
Huss JM, Kopp RP, Kelly DP (2002) Peroxisome proliferator-activated receptor coactivator-1alpha (PGC-1alpha) coactivates the cardiac-enriched nuclear receptors estrogen-related receptor-alpha and –gamma. Identification of novel leucine-rich interaction motif within PGC-1alpha. J Biol Chem 277(43):40265–40274. https://doi.org/10.1074/jbc.M206324200
Huss JM, Torra IP, Staels B, Giguere V, Kelly DP (2004) Estrogen-related receptor alpha directs peroxisome proliferator-activated receptor alpha signaling in the transcriptional control of energy metabolism in cardiac and skeletal muscle. Mol Cell Biol 24(20):9079–9091. https://doi.org/10.1128/mcb.24.20.9079-9091.2004
Ishihara T, Yoshida M, Arita M (2019) Omega-3 fatty acid-derived mediators that control inflammation and tissue homeostasis. Int Immunol 31(9):559–567. https://doi.org/10.1093/intimm/dxz001
Janssens S, Beyaert R (2003) Functional diversity and regulation of different interleukin-1 receptor-associated kinase (IRAK) family members. Mol Cell 11(2):293–302. https://doi.org/10.1016/s1097-2765(03)00053-4
Jiang Z, Johnson HJ, Nie H, Qin J, Bird TA, Li X (2003) Pellino 1 is required for interleukin-1 (IL-1)-mediated signaling through its interaction with the IL-1 receptor-associated kinase 4 (IRAK4)-IRAK-tumor necrosis factor receptor-associated factor 6 (TRAF6) complex. J Biol Chem 278(13):10952–10956. https://doi.org/10.1074/jbc.M212112200
Kawai T, Akira S (2007) TLR signaling. Semin Immunol 19(1):24–32. https://doi.org/10.1016/j.smim.2006.12.004
Kindermann I, Barth C, Mahfoud F, Ukena C, Lenski M, Yilmaz A, Klingel K, Kandolf R, Sechtem U, Cooper LT, Bohm M (2012) Update on myocarditis. J Am Coll Cardiol 59(9):779–792. https://doi.org/10.1016/j.jacc.2011.09.074
Knutti D, Kralli A (2001) PGC-1, a versatile coactivator. Trends Endocrinol Metab 12(8):360–365
Kubala L, Schmelzer KR, Klinke A, Kolarova H, Baldus S, Hammock BD, Eiserich JP (2010) Modulation of arachidonic and linoleic acid metabolites in myeloperoxidase-deficient mice during acute inflammation. Free Radical Biol Med 48(10):1311–1320. https://doi.org/10.1016/j.freeradbiomed.2010.02.010
Lakio L, Lehto M, Tuomainen AM, Jauhiainen M, Malle E, Asikainen S, Pussinen PJ (2006) Pro-atherogenic properties of lipopolysaccharide from the periodontal pathogen Actinobacillus actinomycetemcomitans. J Endotoxin Res 12(1):57–64. https://doi.org/10.1179/096805106x89099
Lehman JJ, Barger PM, Kovacs A, Saffitz JE, Medeiros DM, Kelly DP (2000) Peroxisome proliferator-activated receptor gamma coactivator-1 promotes cardiac mitochondrial biogenesis. J Clin Investig 106(7):847–856. https://doi.org/10.1172/jci10268
Li H, Sun B (2007) Toll-like receptor 4 in atherosclerosis. J Cell Mol Med 11(1):88–95. https://doi.org/10.1111/j.1582-4934.2007.00011.x
Lin J, Wu H, Tarr PT, Zhang CY, Wu Z, Boss O, Michael LF, Puigserver P, Isotani E, Olson EN, Lowell BB, Bassel-Duby R, Spiegelman BM (2002) Transcriptional co-activator PGC-1 alpha drives the formation of slow-twitch muscle fibres. Nature 418(6899):797–801. https://doi.org/10.1038/nature00904
Liu X, Lin Z, Xu Y (2021) Pellino1 promoted inflammation in lung injury model of sepsis by TRAF6/ NF-κB signal pathway. J Inflamm 18(1):11. https://doi.org/10.1186/s12950-021-00276-6
Lopaschuk GD, Ussher JR, Folmes CD, Jaswal JS, Stanley WC (2010) Myocardial fatty acid metabolism in health and disease. Physiol Rev 90(1):207–258. https://doi.org/10.1152/physrev.00015.2009
Lu YC, Yeh WC, Ohashi PS (2008) LPS/TLR4 signal transduction pathway. Cytokine 42(2):145–151. https://doi.org/10.1016/j.cyto.2008.01.006
Maier HJ, Schips TG, Wietelmann A, Kruger M, Brunner C, Sauter M, Klingel K, Bottger T, Braun T, Wirth T (2012) Cardiomyocyte-specific IkappaB kinase (IKK)/NF-kappaB activation induces reversible inflammatory cardiomyopathy and heart failure. Proc Natl Acad Sci USA 109(29):11794–11799. https://doi.org/10.1073/pnas.1116584109
Marsh EK, Prestwich EC, Williams L, Hart AR, Muir CF, Parker LC, Jonker MR, Heijink IH, Timens W, Fife M, Hussell T, Hershenson MB, Bentley JK, Sun SC, Barksby BS, Borthwick LA, Stewart JP, Sabroe I, Dockrell DH, Marriott HM (2020) Pellino-1 regulates the responses of the airway to viral infection. Front Cell Infect Microbiol 10:456. https://doi.org/10.3389/fcimb.2020.00456
Mistry NF, Cresci S (2010) PPAR transcriptional activator complex polymorphisms and the promise of individualized therapy for heart failure. Heart Fail Rev 15(3):197–207. https://doi.org/10.1007/s10741-008-9114-x
Moynagh PN (2009) The Pellino family: IRAK E3 ligases with emerging roles in innate immune signalling. Trends Immunol 30(1):33–42. https://doi.org/10.1016/j.it.2008.10.001
Moynagh PN (2014) The roles of Pellino E3 ubiquitin ligases in immunity. Nat Rev Immunol 14(2):122–131. https://doi.org/10.1038/nri3599
Nauseef WM (2001) Contributions of myeloperoxidase to proinflammatory events: more than an antimicrobial system. Int J Hematol 74(2):125–133. https://doi.org/10.1007/bf02981994
Nemoto S, Vallejo JG, Knuefermann P, Misra A, Defreitas G, Carabello BA, Mann DL (2002) Escherichia coli LPS-induced LV dysfunction: role of toll-like receptor-4 in the adult heart. Am J Physiol Heart Circ Physiol 282(6):H2316-2323. https://doi.org/10.1152/ajpheart.00763.2001
Newsholme P, Curi R, Gordon S, Newsholme EA (1986) Metabolism of glucose, glutamine, long-chain fatty acids and ketone bodies by murine macrophages. Biochem J 239(1):121–125. https://doi.org/10.1042/bj2390121
O’Neill LA, Bryant CE, Doyle SL (2009) Therapeutic targeting of Toll-like receptors for infectious and inflammatory diseases and cancer. Pharmacol Rev 61(2):177–197. https://doi.org/10.1124/pr.109.001073
Oyama J, Blais C Jr, Liu X, Pu M, Kobzik L, Kelly RA, Bourcier T (2004) Reduced myocardial ischemia-reperfusion injury in toll-like receptor 4-deficient mice. Circulation 109(6):784–789. https://doi.org/10.1161/01.CIR.0000112575.66565.84
Paradis AN, Gay MS, Wilson CG, Zhang L (2015) Newborn hypoxia/anoxia inhibits cardiomyocyte proliferation and decreases cardiomyocyte endowment in the developing heart: role of endothelin-1. PLoS ONE 10(2):e0116600. https://doi.org/10.1371/journal.pone.0116600
Puigserver P, Rhee J, Lin J, Wu Z, Yoon JC, Zhang CY, Krauss S, Mootha VK, Lowell BB, Spiegelman BM (2001) Cytokine stimulation of energy expenditure through p38 MAP kinase activation of PPARgamma coactivator-1. Mol Cell 8(5):971–982. https://doi.org/10.1016/s1097-2765(01)00390-2
Randle PJ (1998) Regulatory interactions between lipids and carbohydrates: the glucose fatty acid cycle after 35 years. Diabetes Metab Rev 14(4):263–283
Rossi MA, Celes MR, Prado CM, Saggioro FP (2007) Myocardial structural changes in long-term human severe sepsis/septic shock may be responsible for cardiac dysfunction. Shock 27(1):10–18. https://doi.org/10.1097/01.shk.0000235141.05528.47
Rowe GC, Jiang A, Arany Z (2010) PGC-1 coactivators in cardiac development and disease. Circ Res 107(7):825–838. https://doi.org/10.1161/circresaha.110.223818
Rudiger A, Singer M (2007) Mechanisms of sepsis-induced cardiac dysfunction. Crit Care Med 35(6):1599–1608. https://doi.org/10.1097/01.ccm.0000266683.64081.02
Sagar S, Liu PP, Cooper LT Jr (2012) Myocarditis. Lancet 379(9817):738–747. https://doi.org/10.1016/s0140-6736(11)60648-x
Sato S, Sugiyama M, Yamamoto M, Watanabe Y, Kawai T, Takeda K, Akira S (2003) Toll/IL-1 receptor domain-containing adaptor inducing IFN-beta (TRIF) associates with TNF receptor-associated factor 6 and TANK-binding kinase 1, and activates two distinct transcription factors, NF-kappa B and IFN-regulatory factor-3, in the Toll-like receptor signaling. J Immunol 171(8):4304–4310. https://doi.org/10.4049/jimmunol.171.8.4304
Schauvliege R, Janssens S, Beyaert R (2007) Pellino proteins: novel players in TLR and IL-1R signalling. J Cell Mol Med 11(3):453–461. https://doi.org/10.1111/j.1582-4934.2007.00040.x
Schoonjans K, Staels B, Auwerx J (1996) Role of the peroxisome proliferator-activated receptor (PPAR) in mediating the effects of fibrates and fatty acids on gene expression. J Lipid Res 37(5):907–925
Schreiber SN, Emter R, Hock MB, Knutti D, Cardenas J, Podvinec M, Oakeley EJ, Kralli A (2004) The estrogen-related receptor alpha (ERRalpha) functions in PPARgamma coactivator 1alpha (PGC-1alpha)-induced mitochondrial biogenesis. Proc Natl Acad Sci USA 101(17):6472–6477. https://doi.org/10.1073/pnas.0308686101
Skaug B, Jiang X, Chen ZJ (2009) The role of ubiquitin in NF-kappaB regulatory pathways. Annu Rev Biochem 78:769–796. https://doi.org/10.1146/annurev.biochem.78.070907.102750
Smith H, Liu XY, Dai L, Goh ET, Chan AT, Xi J, Seh CC, Qureshi IA, Lescar J, Ruedl C, Gourlay R, Morton S, Hough J, McIver EG, Cohen P, Cheung PC (2011) The role of TBK1 and IKKepsilon in the expression and activation of Pellino 1. Biochem J 434(3):537–548. https://doi.org/10.1042/bj20101421
St-Pierre J, Lin J, Krauss S, Tarr PT, Yang R, Newgard CB, Spiegelman BM (2003) Bioenergetic analysis of peroxisome proliferator-activated receptor gamma coactivators 1alpha and 1beta (PGC-1alpha and PGC-1beta) in muscle cells. J Biol Chem 278(29):26597–26603. https://doi.org/10.1074/jbc.M301850200
Timmers L, Sluijter JP, van Keulen JK, Hoefer IE, Nederhoff MG, Goumans MJ, Doevendans PA, van Echteld CJ, Joles JA, Quax PH, Piek JJ, Pasterkamp G, de Kleijn DP (2008) Toll-like receptor 4 mediates maladaptive left ventricular remodeling and impairs cardiac function after myocardial infarction. Circ Res 102(2):257–264. https://doi.org/10.1161/CIRCRESAHA.107.158220
Vallabhapurapu S, Karin M (2009) Regulation and function of NF-kappaB transcription factors in the immune system. Annu Rev Immunol 27:693–733. https://doi.org/10.1146/annurev.immunol.021908.132641
van der Vusse GJ, van Bilsen M, Glatz JF (2000) Cardiac fatty acid uptake and transport in health and disease. Cardiovasc Res 45(2):279–293. https://doi.org/10.1016/s0008-6363(99)00263-1
Williams NC, O’Neill LAJ (2018) A role for the Krebs cycle intermediate citrate in metabolic reprogramming in innate immunity and inflammation. Front Immunol 9:141. https://doi.org/10.3389/fimmu.2018.00141
Winterbourn CC, Vissers MC, Kettle AJ (2000) Myeloperoxidase. Curr Opin Hematol 7(1):53–58. https://doi.org/10.1097/00062752-200001000-00010
Zhang Y, Zhuang R, Geng C, Cai X, Lei W, Tian N, Gao F (2013) Insulin promotes T cell recovery in a murine model of autoimmune myocarditis. Clin Exp Immunol 171(1):46–53. https://doi.org/10.1111/j.1365-2249.2012.04662.x
Zhou B, Yu JW (2017) A novel identified circular RNA, circRNA_010567, promotes myocardial fibrosis via suppressing miR-141 by targeting TGF-beta1. Biochem Biophys Res Commun 487(4):769–775. https://doi.org/10.1016/j.bbrc.2017.04.044
Funding
This work was supported by grants from the Natural Science Foundation of Jiangsu Province for Youth (Grant No. BK20151020).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
All authors declare that no competing financial interests exist.
Research involving human participants and/or animals
Human participants were not involved in this study. All rat procedures were approved by the Experimental Animal Care and Use Committee of Nanjing Medical University and conducted in accordance with the Guide for the Care and Use of Laboratory Animals.
Informed consent
All authors gave informed consent to the submission of this manuscript.
Additional information
Handling editor: H. Jakubowski .
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
726_2021_2978_MOESM2_ESM.tif
Supplementary file2 All samples induced by LPS or Pellino1-silencing adenovirus were analyzed and evaluated by LC–MS. 4A: Total ion chromatography (TIC) of the analyzed QC samples was compared with the spectral overlap by means of QC sample spectrogram comparison. x-axis: intensity; y-axis: intension time/minutes. 4B-E: Normalization of the sample. The samples were normalized by median sample (TIF 17783 KB)
726_2021_2978_MOESM3_ESM.tif
Supplementary file3 Metabonimics data were normalized by date scaling. A-D: Normalization of the data. The ion intensity of each metabolite was normalized with mean of metabolite (TIF 15649 KB)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yang, C., Zhao, K., Chen, X. et al. Pellino1 deficiency reprograms cardiomyocytes energy metabolism in lipopolysaccharide-induced myocardial dysfunction. Amino Acids 53, 713–737 (2021). https://doi.org/10.1007/s00726-021-02978-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00726-021-02978-w