Abstract
Complement factor C5a is one of the most powerful pro-inflammatory agents involved in recruitment of leukocytes, activation of phagocytes and other inflammatory responses. C5a triggers inflammatory responses by binding to its G-protein-coupled C5a-receptor (C5aR). Excessive or erroneous activation of the C5aR has been implicated in numerous inflammatory diseases. The C5aR is therefore a key target in the development of specific anti-inflammatory compounds. A very potent natural inhibitor of the C5aR is the 121-residue chemotaxis inhibitory protein of Staphylococcus aureus (CHIPS). Although CHIPS effectively blocks C5aR activation by binding tightly to its extra-cellular N terminus, it is not suitable as a potential anti-inflammatory drug due to its immunogenic properties. As a first step in the development of an improved CHIPS mimic, we designed and synthesized a substantially shorter 50-residue adapted peptide, designated CHOPS. This peptide included all residues important for receptor binding as based on the recent structure of CHIPS in complex with the C5aR N terminus. Using isothermal titration calorimetry we demonstrate that CHOPS has micromolar affinity for a model peptide comprising residues 7–28 of the C5aR N terminus including two O-sulfated tyrosine residues at positions 11 and 14. CD and NMR spectroscopy showed that CHOPS is unstructured free in solution. Upon addition of the doubly sulfated model peptide, however, the NMR and CD spectra reveal the formation of structural elements in CHOPS reminiscent of native CHIPS.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
As part of the host defense system, the human complement cascade initiates inflammatory responses directed against invading infectious microorganisms, injury and other threatening conditions (Lee et al. 2008). Complement factor C5a is the most powerful pro-inflammatory agent generated during complement activation. C5a interacts with the membrane associated G-protein-coupled C5a receptor (C5aR) resulting in chemotaxis of specific white blood cells, activation of phagocytes, release of granule-based enzymes, and the generation of oxidants (Guo and Ward 2005). C5a is a 74-residue glycoprotein comprising a bundle of four anti-parallel α-helices stabilized by three disulfide bonds (PDB ID code: 1KJS). Binding and activation of the C5aR by C5a is considered a two-step process, in which residues in the region between 12 and 46 of C5a bind to a primary binding site located in the extra-cellular N terminus of the C5aR. Subsequently the C-terminal portion of C5a (residues 69–74) binds to the C5aR activation domain located inside the receptor core (Chen et al. 1998; Gerber et al. 2001). Together these two binding sites provide the complex of C5a and the C5aR sub-nanomolar affinity (K d ≈ 0.60 nM; van Epps et al. 1993). Although under normal circumstances C5a-mediated C5aR activation is highly favorable, excessive levels of C5a can be deleterious to the host and have been related to numerous inflammatory and autoimmune diseases (e.g. rheumatoid arthritis, inflammatory bowel disease and reperfusion injury). Specific inhibition of C5aR activation is considered a promising strategy to treat such conditions (Allegretti et al. 2005). So far, the design and development of novel anti-inflammatory agents was primarily focussed on small organic compounds and short C5a peptide analogs, which bind with high affinity to the C5aR activation site (Allegretti et al. 2005; Monk et al. 2007). An alternative approach to inhibit C5aR activation was inspired by the discovery of the Chemotaxis Inhibitory Protein of Staphylococcus aureus abbreviated as CHIPS (Veldkamp et al. 2000). CHIPS is a 121-residue immune evasive protein excreted by Staphylococcus aureus bacteria in order to prevent host inflammatory responses triggered by formylated peptides and C5a. CHIPS binds to the formylated peptide receptor (FPR) and the C5aR with high affinity (K d = 35.4 ± 7.7 nM and K d = 1.1 ± 0.2 nM, respectively; Postma et al. 2004). Mutational studies revealed that the C5aR blocking activity of CHIPS is entirely conserved in a protein fragment lacking the first 30 residues. This truncated protein was designated CHIPS31–121 and its structure was solved by Haas et al. (2005). CHIPS31–121 has an entirely different folding topology compared to C5a and is composed of a single α-helix packed onto a four-stranded anti-parallel β-sheet. This particular topology is present in several other S. aureus proteins with immune modulating properties (Haas et al. 2005).
In contrast to C5a, CHIPS binds exclusively to the C5aR N terminus (Postma et al. 2005). This part of the receptor is post-translationally modified by introduction of two sulfate groups on tyrosine residues at positions 11 and 14 (Farzan et al. 2001). Sulfation of these tyrosines appeared to be crucial for tight binding to CHIPS31–121 as was concluded from ITC binding studies using several sulfated and unsulfated mimics of the C5aR N terminus (Bunschoten et al. 2009; Ippel et al. 2009). The highest affinity for CHIPS31–121 (K d = 8.4 ± 1.1 nM) was observed for a peptide composed of residues 7–28 of the C5aR with both tyrosine residues sulfated (designated C5aR7–28S2; Ippel et al. 2009). This peptide binds almost as strong to CHIPS as the native C5aR. This implies that all moieties essential for the interactions between CHIPS and the C5aR are present within this peptide mimic. The free N terminus of the C5aR is virtually unstructured, which is also the case for the short receptor mimics. Upon binding to CHIPS, residues 10–24 of these C5aR mimics adopt a well-defined conformation (PDB ID code: 2K3U; Ippel et al. 2009). In the complex, residues 10–14 and 19–24 of C5aR7–28S2 form two short stretches of β-strand, which are hydrogen bonded in an anti-parallel fashion to strand β4 and residues 104–107 of CHIPS31–121, respectively. These two stretches are interconnected by a single turn comprising residues 15–18. Sulfated tyrosine 11 interacts mainly with residues in the α-helix of CHIPS31–121, while sulfated tyrosine 14 is primarily accommodated by residues in the loop between the α-helix and the first β-strand (residues 52–59; Fig. 1a). The sequence between residues T66 and Y94 of CHIPS31–121 does not contribute to interactions with the receptor, but is essential for its native structure (Ippel et al. 2009).
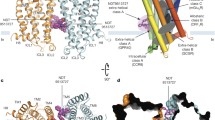
Topology design of the CHOPS construct. a Cartoon representation of one of the NMR structures of CHIPS31–121 (PDB ID code: 1XEE). The two regions interacting with the C5aR are indicated: residues 43–61 (α-helix and β1) and residues 95–111 (β3 and β4). The N and C termini as well as the numbering of the β-strands are indicated. b The amino acid sequence of CHOPS. The D-Pro-Gly linker is indicated in black. c Cartoon representation of CHOPS based on the structure of CHIPS31–121. The D-Pro-Gly linker is shown in stick representation. The numbering of the terminal residues and of the residues flanking the linker is indicated
Despite its strong C5aR inhibitory potency, intact CHIPS itself is not amenable for use as an anti-inflammatory agent. Several immunogenic surface epitopes have been identified by Gustafsson et al. (2009). A recent study of Wright et al. (2007) showed the presence of anti-CHIPS antibodies in numerous serum samples of human donors. Therefore, administration of intact CHIPS protein can potentially lead to adverse immunogenic responses. Here, we describe the design, chemical synthesis, and analysis of a protein construct in which specific segments of CHIPS crucial for interactions with the C5aR have been incorporated while a number of non-interacting segments were omitted. We denote this protein construct with the acronym CHOPS, which stands for ‘CHemotaxis inhibitory cOnstruct Protein of Staphylococcus aureus’. The ultimate goal is to obtain a CHOPS molecule, which is non-immunogenic, but has a high inhibitory potency for the C5aR.
Materials and methods
Materials
Peptide grade DiPEA, DCM, NMP, TFA, piperidine, and HPLC grade solvents were purchased from Biosolve B. V. (Valkenswaard, The Netherlands). Fmoc-protected amino acids were purchased from GL Biochem Ltd. (Shanghai, China). Side-chain protecting groups were chosen as: Boc for lysine, tBu for aspartic acid, glutamic acid, serine, threonine, and tyrosine, Trt for asparagine and glutamine, and Pbf for arginine. Unless stated otherwise, chemicals were obtained from commercial sources and used without further purification.
Compound analysis
Analytical HPLC was performed using an automatic HPLC system (Shimadzu) with an analytical reversed-phase column and a UV detector operating at 214 nm with a flow rate of 0.75 mL/min. A Phenomenex Gemini C18 column (110 Å, 5 μm, 250 × 4.6 mm) was used. TFA buffers were used (buffer A: H2O:MeOH, 80:20 v:v; buffer B: H2O:MeOH, 5:95 v:v, both containing 0.1% TFA). Elution was effected with a linear gradient from 100% A to 100% B over 48 min. Preparative HPLC was performed using an automatic Prep LCMS-QP8000α HPLC system (Shimadzu) with a preparative reversed-phase column and a UV detector operating at 214 nm with a flow rate of 12.5 mL/min. A Reprosil-Pur C18-AQ column (120 Å, 10 μm, 250 × 22 mm) was used. TFA buffers were used (buffer A: H2O:MeOH, 80:20 v:v; buffer B: H2O:MeOH, 5:95 v:v, both containing 0.1% TFA). Elution was effected with a linear gradient from 100% A to 100% B over 100 min. The peptides were characterized using electrospray mass spectrometry (ESI-MS), which was performed on a Thermo Finnigan LCQ DECA XP MAX ion trap mass spectrometer.
Peptide synthesis
CHOPS was synthesized on a Rink Amide PEG resin (0.52 mmol/g) (Matrix Innovation Inc., Montreal, Canada) on a 0.25 mmol scale. The peptide was assembled using an automatic ABI 433A Peptide Synthesizer, equipped with a UV-monitoring system, which was used to monitor the Fmoc removal step, i.e., formation of the dibenzofulvene-piperidine adduct absorbing at 301 nm. ABI FastMoc 0.25 mmol protocols were applied, with the exception of a standard double coupling of 45 min. The synthesis was carried out on 0.48 g resin. The resin was washed with DCM and NMP (five times). Subsequently, 1 mmol of the appropriate amino acid was dissolved in NMP (2 mL), and HBTU/HOBt (1 mmol, 2.78 mL of 0.36 M) in NMP was added. To this mixture, DiPEA (1 mL, 2 M) in NMP was added, and the activated amino acid was then transferred to the reaction vessel. After 45 min, the reaction vessel was drained and the resin was washed with NMP (three times) followed by addition of another batch of pre-activated amino acid, which was allowed to couple for another 45 min. Next, any of the remaining free amino groups were acetylated with an acetic anhydride capping solution (0.5 M Ac2O, 0.125 M DiPEA, and 0.015 M HOBt in NMP) for 15 min. After capping the Fmoc protective group was removed from the N terminus by treatment with 20% piperidine solution in NMP twice (3 and 7.6 min). The last coupling cycle was followed by removal of the Fmoc-group by a 20% piperidine solution, washing the resin with NMP, and acetylation of the N terminus by treatment with the acetic anhydride capping solution for 15 min. Finally, the resin was washed with NMP (five times) and DCM (six times), removed from the reaction vessel, washed with ether, and dried in vacuo over P2O5.
The anchored peptide obtained in this way was de-protected and cleaved from the solid support by treatment with TFA/H2O/TIS (95/2.5/2.5, 25 mL) for 2 h at room temperature. The mixture was filtered and the residue washed thoroughly with TFA (two times 10 mL). The reaction mixture was concentrated in vacuo to a volume of approximately 10 mL and added dropwise to 90 mL MTBE/n-hexane (1/1 v/v) solution. The precipitate was collected by centrifugation (3,000 rpm, 10 min), the supernatant was decanted and the pellet was resuspended in MTBE/n-hexane (1/1 v/v) (100 mL) and centrifuged again. This procedure was repeated twice. Afterwards, the pellets were dissolved in CH3CN/water (1/1 v/v) (ca. 60 mL) and lyophilized to give 595 mg of the crude peptide as a white fluffy solid.
The crude peptide (400 mg) was dissolved in 30 mL buffer A, 10 mL buffer B and purified by preparative HPLC (Reprosil-Pur C18-AQ, TFA buffers) in eight runs. Pure fractions were pooled and lyophilized to give 29 mg of pure CHOPS. The CHOPS sample was characterized by analytical HPLC (Gemini C18, TFA buffers, R t = 36.4 min, Purity > 98%) (Supplementary Figure 1) and by ESI-MS (Supplementary Figure 2; calculated average mass [M + 5H]5+ for C261H422N70O74S: 1,152.34; found: 1,152.30).
The C5a receptor mimic peptides C5aR7–28 and C5aR7–28S2 were synthesized as has been described previously (Bunschoten et al. 2009).
ITC experiments
The binding affinities of the C5aR mimics C5aR7–28 and C5aR7–28S2 with CHOPS were measured using an ITC200 Microcalorimeter (MicroCal) operating at 283 K. The measuring cell was filled with 208 μL of a 0.19 mM solution of CHOPS in a 20 mM sodium phosphate buffer at pH 6.5. The concentration of CHOPS was determined by OD280 measurements. The syringe was loaded with 30 μL of a 2.5 mM solution of one of the C5aR-mimics in the same buffer system. After each incremental addition of the solution in the syringe, the integrated heat change due to binding was measured. The data were analyzed using the Microcal Origin software and fitted by non-linear regression analysis. Three independent experiments were carried out. The experimental errors were estimated by Monte Carlo simulations using the standard deviations of the individual experiments.
NMR spectroscopy
NMR samples of C5aR7–28, C5aR7–28S2 and CHOPS were at concentrations of 0.8–1 mM in 10/90% (v/v) D2O/H2O sodium phosphate buffers (20 mM, pH 6.5). Spectra were recorded at 288 K on a Varian INOVA 500 and a Bruker Avance 750 MHz spectrometer equipped with an HCN triple-resonance pulsed field gradient probe. Sequential 1H NMR assignments of CHOPS were accomplished using a combination of NOESY, TOCSY, and 13C-HSQC spectra.
CD measurements
CD spectra (190–260 nm) were recorded on an Olis RSM1000 spectrophotometer operating at 2 nm spectral resolution (slit size 1.24 mm). Samples of CHIPS31–121 (43 μM) in 20 mM sodium phosphate buffer (pH 6.5), CHOPS (50 μM), C5aR7–28 and C5aR7–28S2 (50 μM) in 10 mM sodium phosphate buffer (pH 7.4) were measured at 298 K using a 0.5 mm cuvette. To gain a satisfactory S/N ratio five scans were summed, with data points averaged by three-point triangular smoothing.
Results and discussion
Design of CHOPS
The Staphylococcal protein CHIPS is one of the most potent inhibitors of C5a-induced inflammatory responses presently known. In contrast to the numerous agents developed to interact directly with the C5aR activation site located inside the receptor core (Proctor et al. 2006; Chen et al. 2010), CHIPS blocks activation by C5a by binding with high affinity to the flexible extra-cellular N-terminal portion of the C5aR (Postma et al. 2005). The interaction surface of CHIPS31–121 with the C5aR comprises ~20% of its solvent accessible surface and is not confined to a limited region of the protein. The interactions between CHIPS and the C5aR involve a substantial number of non-sequential amino acids optimally positioned in the inhibitory protein to provide tight binding. A successful mimic of CHIPS should not only include the amino acid residues (or mimics of these) crucial for C5aR binding, but also the amino acids responsible for the proper spatial arrangement dictated by the CHIPS folding topology. Our first approach to build such a structure is to leave out a limited number of residues which do not interact directly with the C5aR, but with the intention to leave the structural integrity of CHIPS31–121 intact. NMR titration studies revealed that two regions of CHIPS31–121 show relatively large perturbations in the backbone and Cβ chemical shifts (Ippel et al. 2009): the first region includes residues 43–61, which comprises part of the α-helix and the subsequent loop connecting strand β1 (Fig. 1a). The second region is composed of residues 95–111 and comprises strands β3 and β4 of CHIPS31–121. The non-interacting portion of CHIPS31–121 comprises strand β2 and the long loop connecting β2 with β3 (Fig. 1a). This portion could potentially be left out by directly connecting strand β1 with β3 via a tight turn. Inspection of the NMR structural models of CHIPS31–121 reveals that residue L65 at the end of strand β1 and residue K95 at the start of strand β3 are proximal and offer an excellent opportunity to link the two fragments interacting with the C5aR together to form one short, contiguous sequence. Several β-hairpin inducing sequences have been described and reviewed in the literature (Blanco et al. 1998; Stotz and Topp 2004). One of the smallest peptide fragments, which induces a β-turn and facilitates the formation of an anti-parallel β-sheet, is the dipeptide D-Pro-Gly (Haque and Gellman 1997). This fragment was chosen to link the N- and C-terminal segments of CHIPS, which interact with the C5aR, together. These segments were chosen to comprise the complete elements of secondary structure as present in native CHIPS (i.e. the α-helix and β-strands β1, β3, and β4) in order to pursue structural integrity. The resulting construct consisted of the CHIPS amino acid sequences T36-L65 and K95-G112 interconnected by D-Pro-Gly (Fig. 1b). A number of residues suggested to be part of discontinuous immunogenic epitopes by Gustafsson et al. (2009) are not present in this construct (designated CHOPS). A model representation of CHOPS based on the structure of native CHIPS31–121 is presented in Fig. 1c.
Affinity of CHOPS for the C5aR N terminus
The affinity of the CHOPS fragment for the C5aR was determined using isothermal titration calorimetry (ITC). We synthesized two peptide mimics of the C5aR N terminus: unsulfated peptide C5aR7–28 representing residues 7–28 of the C5aR and peptide C5aR7–28S2, the same sequence with tyrosine residues 11 and 14 O-sulfated. We titrated a solution of CHOPS with these peptides and recorded the subsequent heat exchange upon formation of the complex. Two typical ITC experiments are shown in Supplementary Figure 3. Clearly, titration of the doubly sulfated peptide C5aR7–28S2 to CHOPS resulted in a substantial exothermic effect (Supplementary Figure 3a) while no significant response was detected in the ITC experiment with the unsulfated peptide C5aR7–28 (Supplementary Figure 3b). Gratifyingly, the affinity of CHOPS for C5aR7–28S2 was in the micromolar range (K d = 3.6 ± 0.2 μM; n = 3). The complete thermodynamic analysis of these ITC data plus the comparison with previous ITC studies of CHIPS31–121 and C5a peptide mimics is compiled in Supplementary Table 1.
NMR spectroscopy
Previous NMR studies revealed that the synthetic peptides C5aR7–28 and C5aR7–28S2, which mimic the N-terminal portion of the C5aR, were very flexible in solution and did not have detectable propensity for any preferred secondary structure. Although there is no detailed structure available for the intact C5aR, it is expected that its free extra-cellular N terminus (residues 1–35) is unstructured as well. The protein CHIPS31–121 does adopt a well-defined conformation with flexible regions at the termini and some disorder in the loop region between the α-helix and strand β1 (Haas et al. 2005). As could be inferred from 15N relaxation studies this particular loop region adopts an ordered conformation in the complex with C5aR7–28S2 (Ippel et al. 2009). NMR spectra of the free CHOPS construct appear to be typical for a largely unstructured polypeptide chain (Supplementary Figure 4a). 2D NOE spectra of free CHOPS contain predominantly sequential NOEs, but a few long-range contacts could be identified. These non-sequential cross-peaks are indicative for an anti-parallel orientation of strands β1 and β3, which are bridged by the β-hairpin inducing D-Pro-Gly sequence (Fig. 2).
Titration of CHOPS with the unsulfated receptor mimic C5aR7–28 did not result in any changes in the 1H-spectrum of the latter. In contrast, titration of CHOPS with the sulfated receptor mimic C5aR7–28S2 resulted in increased dispersion of resonance lines, which is characteristic for non-random coil behavior (Supplementary Figure 4b). The complex between CHOPS and C5aR7–28S2 is still flexible and the 1H-spectra show a high degree of overlap. Nevertheless, we could assign some of the NMR signals as indicated in Supplementary Figure 4b. These new signals are at comparable positions as in spectra of the complex between CHIPS31–121 and C5aR7–28S2, and indicative for the formation of native-like structure. Similar features were observed in 1H–13C HSQC spectra upon titration of C5aR7–28S2 to CHOPS (Fig. 3).
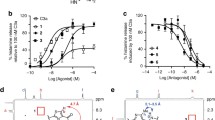
The NOESY spectra of the CHOPS:C5aR7–28S2 complex suffer from severe overlap, but still a limited number of long-range intra-molecular and inter-molecular NOE cross-peaks could be assigned. Inter-molecular NOEs were identified between the aromatic side-chain protons of sulfated tyrosine sY14 of C5aR7–28S2 and the side-chains of T53 (Fig. 4a) and Y108 (Fig. 4c) of CHOPS. H13 of C5aR7–28S2 has an NOE contact with V109 of CHOPS (Fig. 4b). Intra-molecular NOEs were identified between T53 and L49 and between Y97 and V109 (Fig. 4a). Similar peaks were observed in NOESY spectra of the complex between CHIPS31–121 and C5aR7–28S2. The position of these residues in the NMR structure of the CHIPS31–121:C5aR7–28S2 complex is indicated in Fig. 4d, e.
Observed cross-peaks in the NOESY spectrum of the CHOPS:C5aR7–28S2 complex in relation to structural features of the CHIPS31–121:C5aR7–28S2 complex (PDB ID code: 2K3U). a – c Different sections of the NOESY spectrum recorded at 288 K of a 1:1 mixture of CHOPS and C5aR7–28S2. Identified long-range intra-molecular and inter-molecular NOE cross-peaks are indicated (normal fonts for residues belonging to CHOPS and italic fonts for residues belonging to C5aR7–28S2). d – e Cartoon representation of the structure of the CHIPS31–121:C5aR7–28S2 complex. The side-chains of the residues identified in a – c are shown in stick representation. NOE cross-peaks observed in a – c are indicated by arrows in the structure
CD spectroscopy
The presence of residual structure in the C5aR mimics and CHOPS was also monitored by CD spectroscopy. The CD spectra of the C5a-receptor mimics C5aR7–28 and C5aR7–28S2, and CHOPS show no structural features apart from a shallow minimum at 200 nm (Fig. 5). The CD spectrum of free CHIPS31–121 comprises a large positive signal around 190 nm and a minimum around 205 nm (Fig. 5). This spectrum does not change significantly upon binding of C5aR7–28S2 (data not shown). Titration of CHOPS with a stoichiometric amount of unsulfated receptor mimic C5aR7–28 resulted in an increase of the CD signal, although the shape of the spectrum did not change (Fig. 5a). Stoichiometric titration of CHOPS with sulfated receptor mimic C5aR7–28S2, on the other hand yielded a clear change of the CD spectrum: an increase in the signal around 190 nm and a shift of the minimum from 200 to 205 nm (Fig. 5b). These changes shift the appearance of the CD spectrum towards that of CHIPS31–121 although at lower intensities.
In this study we aimed to create a shorter version of the immune evasive protein CHIPS based on the NMR structure of the complex between CHIPS31–121 and C5aR7–28S2. The construct we designed (CHOPS) comprises all portions of CHIPS31–121 important in the interaction with the C5aR. Portions outside the binding region including strand β2 and the connecting loop between β2 and β3 were discarded. This was accomplished by coupling strand β1 and β3 together via a D-Pro-Gly linker segment. The resulting 50-residue long peptide appeared to be largely unfolded apart from some residual structure around the β-hairpin inducing D-Pro-Gly linker sequence. ITC studies revealed that CHOPS binds to the doubly sulfated C5a-receptor mimic C5aR7–28S2 with micromolar affinity (K d = 3.6 ± 0.2 μM). Although the affinity of C5aR7–28S2 to CHOPS is three orders of magnitude lower compared to binding to CHIPS31–121 (K d = 8.4 ± 1.2 nM; Ippel et al. 2009), this is a very promising result for a first lead compound. No detectable affinity of CHOPS was observed in the ITC measurements using the unsulfated mimic C5aR7–28. This is consistent with previous measurements of CHIPS31–121 and C5aR7–28, which revealed that the absence of the two sulfate moieties results in an almost 400-fold decrease in affinity.
NMR spectroscopy confirmed the results obtained by ITC. Titration of CHOPS with unsulfated receptor mimic C5aR7–28 resulted in the sum of its constituent 1H-spectra, while titration of doubly sulfated peptide C5aR7–28S2 yielded a completely different 1H-spectrum with signals shifted from their random coil position. The latter is indicative for the formation of defined structural elements. The NMR titration experiments using C5aR7–28S2 showed binding in a fast-exchange regime, compatible with the observed micromolar affinity by ITC (Cavanagh et al. 2007). Several characteristic features observed in spectra of the CHIPS31–121:C5aR7–28S2 complex were also present in spectra of CHOPS:C5aR7–28S2. Specific resonances of residues L49, T53, T96, Y97, Y108, V109 and N111 have chemical shifts comparable with their counterparts in the CHIPS31–121:C5aR7–28S2 complex. We also observed NOE cross-peaks between residues sY14, T53, L49, and Y108 and between residues H13, V109, and Y97 in NOESY spectra of CHOPS:C5aR7–28S2. These peaks are reminiscent of NOE contacts observed in spectra of the complex between CHIPS31–121 and C5aR7–28S2 and reveal that CHOPS, in the presence of sulfated peptide C5aR7–28S2, adopts similar structures as native CHIPS31–121.
The structural characteristics of CHOPS, either free in solution or in complex with the C5aR mimics, were also studied by CD spectroscopy. The spectra of the separate peptides (CHOPS, C5aR7–28 or C5aR7–28S2) did not show any significant absorption apart from a shallow minimum around 200 nm. Titration of CHOPS with receptor mimic C5aR7–28 increased the amount of absorption, but not the position of its minimum. Titration of CHOPS with the doubly sulfated receptor mimic C5aR7–28S2 caused an increase in absorption around 190 nm as well as a shift of the absorption minimum from 200 to 205 nm. Although the maximum and minimum intensities are smaller, the overall shape of the CD spectrum of the CHOPS:C5aR7–28S2 complex resembles that of native CHIPS31–121.
Conclusions
We have designed and synthesized a significantly reduced-size mimic of the protein CHIPS, which we coined CHOPS. This construct has a binding affinity of 3.6 μM with respect to the sulfated receptor mimic C5aR7–28S2, but no affinity for the unsulfated receptor mimic C5aR7–28. We conclude, based on NMR and CD studies, that CHOPS adopts structures comparable with native CHIPS31–121 upon binding of C5aR7–28S2. The CHOPS:C5aR7–28S2 complex is, however, still flexible. We expect that improved affinity can be achieved by introduction of more rigid moieties, which will force this mimic more into the structure of native CHIPS.
References
Allegretti M, Moriconi A, Beccari AR, Di Bitondo R, Bizzarri C, Bertini R, Colotta F (2005) Targeting C5a: recent advances in drug discovery. Curr Med Chem 12:217–236
Blanco F, Ramirez-Alvarado M, Serrano L (1998) Formation and stability of beta-hairpin structures in polypeptides. Curr Opin Struct Biol 8:107–111
Bunschoten A, Kruijtzer JAW, Ippel JH, de Haas CJC, van Strijp JAG, Kemmink J, Liskamp RMJ (2009) A general sequence independent solid phase method for the site specific synthesis of multiple sulfated-tyrosine containing peptides. Chem Commun 2999–3001
Cavanagh J, Fairbrother WJ, Palmer AG, Rance M, Skelton NJ (2007) Protein NMR spectroscopy: principles and practice, 2nd edn. Elsevier Academic Press, San Diego
Chen ZG, Zhang XL, Gonnella NC, Pellas TC, Boyar WC, Ni F (1998) Residues 21–30 within the extracellular N-terminal region of the C5a receptor represent a binding do main for the C5a anaphylatoxin. J Biol Chem 273:10411–10419
Chen JJ, Cole DC, Ciszewski G, Crouse K, Ellingboe JW, Nowak P, Tawa GJ, Berstein G, Li W (2010) Identification of a new class of small molecule C5a receptor antagonists. Bioorg Med Chem Lett 20:662–664
Farzan M, Schnitzler CE, Vasilieva N, Leung D, Kuhn J, Gerard C, Gerard NP, Choe H (2001) Sulfated tyrosines contribute to the formation of the C5a docking site of the human C5a anaphylatoxin receptor. J Exp Med 193:1059–1065
Gerber BO, Meng EC, Dotsch V, Baranski TJ, Bourne HR (2001) An activation switch in the ligand binding pocket of the C5a receptor. J Biol Chem 276:3394–3400
Guo RF, Ward PA (2005) Role of C5A in inflammatory responses. Annu Rev Immunol 23:821–852
Gustafsson E, Haas PJ, Walse B, Hijnen M, Furebring C, Ohlin M, van Strijp JAG, van Kessel KPM (2009) Identification of conformational epitopes for human IgG on Chemotaxis inhibitory protein of Staphylococcus aureus. BMC Immunol 10:13
Haas PJ, de Haas CJC, Poppelier MJJC, van Kessel KPM, van Strijp JAG, Dijkstra K, Scheek RM, Fan H, Kruijtzer JAW, Liskamp RMJ, Kemmink J (2005) The structure of the C5a receptor-blocking domain of chemotaxis inhibitory protein of Staphylococcus aureus is related to a group of immune evasive molecules. J Mol Biol 353:859–872
Haque TS, Gellman SH (1997) Insights on beta-hairpin stability in aqueous solution from peptides with enforced type I′ and type II′ beta-turns. J Am Chem Soc 119:2303–2304
Ippel JH, De Haas CJC, Bunschoten A, van Strijp JAG, Kruijtzer JAW, Liskamp RMJ, Kemmink J (2009) Structure of the tyrosine-sulfated C5a-receptor N-terminus in complex with chemotaxis inhibitory protein of Staphylococcus aureus. J Biol Chem 284:12363–12372
Lee H, Whitfeld PL, Mackay CR (2008) Receptors for complement C5a. The importance of C5aR and the enigmatic role of C5L2. Immunol. Cell Biol 86:153–160
Monk PN, Scola A, Madala PK, Fairlie DP (2007) Function, structure and therapeutic potential of complement C5a receptors. Br J Pharmacol 152:429–448
Postma B, Poppelier MJ, van Galen JC, Prossnitz ER, van Strijp JAG, de Haas CJC, van Kessel KPM (2004) Chemotaxis inhibitory protein of Staphylococcus aureus binds specifically to the C5a and formylated peptide receptor. J Immunol 172:6994–7001
Postma B, Kleibeuker W, Poppelier MJJG, Boonstra M, Van Kessel KPM, Van Strijp JAG, de Haas CJC (2005) Residues 10–18 within the C5a receptor N-terminus compose a binding domain for chemotaxis inhibitory protein of Staphylococcus aureus. J Biol Chem 280:2020–2027
Proctor LM, Woodruff TM, Taylor SM (2006) Recent developments in C5/C5a inhibitors. Expert Opin Ther Pat 16:445–458
Stotz CE, Topp EM (2004) Applications of model beta-hairpin peptides. J Pharm Sci 93:2881–2894
van Epps DE, Simpson SJ, Johnson R (1993) Relationship of C5a receptor modulation to the functional responsiveness of human polymorphonuclear leukocytes to C5a. J Immunol 150:246–252
Veldkamp KE, Heezius HCJM, Verhoef J, van Strijp JAG, van Kessel KPM (2000) Modulation of neutrophil chemokine receptors by Staphylococcus aureus supernate. Infect Immun 68:5908–5913
Wright AJ, Higginbottom A, Philippe D, Upadhyay A, Bagby S, Read RC, Monk PN, Partridge LJ (2007) Characterisation of receptor binding by the chemotaxis inhibitory protein of Staphylococcus aureus and the effects of the host immune response. Mol Immunol 44:2507–2517
Acknowledgments
This work was supported in part by the Dutch Technology Foundation STW, Applied Science Division of NWO, and the Technology Program of the Ministry of Economic Affairs (UKG.06609).
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Bunschoten, A., Ippel, J.H., Kruijtzer, J.A.W. et al. A peptide mimic of the chemotaxis inhibitory protein of Staphylococcus aureus: towards the development of novel anti-inflammatory compounds. Amino Acids 40, 731–740 (2011). https://doi.org/10.1007/s00726-010-0711-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00726-010-0711-3