Abstract
MADS-box genes are known to play important roles in diverse aspects of growth/devolopment and stress response in several plant species. However, no study has yet examined about MADS-box genes in P. vulgaris. In this study, a total of 79 PvMADS genes were identified and classified as type I and type II according to the phylogenetic analysis. While both type I and type II PvMADS classes were found to contain the MADS domain, the K domain was found to be present only in type II PvMADS proteins, in agreement with the literature. All chromosomes of the common bean were discovered to contain PvMADS genes and 17 paralogous gene pairs were identified. Only two of them were tandemly duplicated gene pairs (PvMADS-19/PvMADS-23 and PvMADS-20/PvMADS-24), and the remaining 15 paralogous gene pairs were segmentally duplicated genes. These duplications were found to play an important role in the expansion of type II PvMADS genes. Moreover, the RNAseq and RT-qPCR analyses showed the importance of PvMADS genes in response to drought stress in P. vulgaris.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In higher plants, transcription factors (TFs) have a range of functions during their life cycle. They are involved in the growth, development, morphogenesis, and stress responses of plants by binding to cis-acting regulatory sequences (Singh et al. 2002). A group of transcription factors known as MADS-box genes are present in eukaryotes like plants, yeasts, insects, amphibians, and mammals. The genes MONOCHROME HOME MAINTENANCE 1 (MCM1) from yeast, AGAMOUS (AG) from Arabidopsis, DEFICIENT (DEF) from Antirrhinum, and serum response factor (SRF) from humans collectively give rise to the name of MADS-box family (Riechmann and Meyerowitz 1997). MADS-box genes can be divided into two primary classes, type I and type II, in plants, mammals, and fungi (Ma et al. 2017). Compared to type I (M type) classes, type II (MIKC type) genes have three extra functional domains: (1) dimerization-inducing keratin-like helix (K) domain, (2) DNA-binding/dimerization domain, and (3) C-terminal domain that participates in the creation of tertiary or quaternary protein complexes during transcriptional activation contribute to functional specificity (Theißen et al. 1996; Ayra et al. 2021). Type I and type II genes are further divided into the subclasses based on structural variations (Henschel et al. 2002).
MADS-box genes play a vital role in diverse essential developmental processes, including flower structure and root development, fruit growth, timing of flowering, and regulation of gametophyte cellular division (Liljegren et al. 2000; Michaels et al. 2003; Zhang et al. 2008; Martel et al. 2011). The majority of the proteins related to flower organ development are regulated by MADS-box genes in plants (Coen and Meyerowitz 1991; Pelaz et al. 2000; Thei and Saedler 2001; Pinyopich et al. 2003; Ditta et al. 2004).
In previous studies, it was shown that MADS-box genes had the ability to respond to diverse abiotic stress conditions (Wei et al. 2014). In a study by Zhang et al. (2012), MADS-box genes were found to be differentially expressed in response to ABA, cold, salt, and drought stresses in maize (Zhang et al. 2012). Another study in rice showed that MADS-box genes were associated with pathogen resistance and drought tolerance (Lee et al. 2008b; Khong et al. 2015). In the study of Chen et al. (2019), transgenic CaMADS-expressing pepper plants have been found to be more resilient to cold, high salt, and mannitol stresses than wild type (Chen et al. 2019). Additionally, MADS-box genes linked to stress tolerance have been found in other species, such as wheat, sheep grass, chinese cabbage, and tomato (Saha et al. 2015; Guo et al. 2016; Jia et al. 2018; Schilling et al. 2020). The findings related to the function of MADS-box genes for biotic and abiotic stress tolerance offer valuable resources for their possible utilization in plant breeding and enhancing crop production. Nonetheless, there is limited information regarding the MADS-box genes in common bean in the literature. However, common bean is the most widely cultivated and produced edible legume globally which is consumed as a fresh vegetable or dried grain (Anlarsal et al. 2000; Idiku et al. 2022). Bioinformatics analysis was used to identify and characterize the MADS-box gene family members in common bean in this study. In addition, the role of PvMADS genes in response to drought stress was evaluated at mRNA level for the first time using RNAseq and RT-qPCR analyses.
Materials and methods
Identification of MADS-box proteins in P. vulgaris
MADS-box sequences of P. vulgaris were acquired from the Phytozome (https://phytozome-next.jgi.doe.gov/) through Pfam (PF00319) code search and the sequences were deposited in Supplementary Table 1 (Goodstein et al. 2012). For characterization of putative proteins, candidate P. vulgaris MADS-box proteins were used as query sequences in NCBI blastp (https://blast.ncbi.nlm.nih.gov/Blast.cgi). Redundant sequences were removed using the decrease redundancy tool (http://web.expasy.org/decrease_ redundancy/), and sequences were checked for MADS box domains using HMMER (http://www.ebi.ac.uk). Physical and chemical properties were calculated using Expasy ParatProm (www.web.expasy.org/protparam). Domains of MADS box proteins were discovered using InterProScan (https://www.ebi.ac.uk/interpro/search/sequence/). The WoLF PSORT: Protein Subcellular Localization Estimator Tool (https://wolfpsort.hgc.jp/) was used to determine where PvMADS proteins were located. Wolfpsort is a tool used to predict the cell nucleus localization of a protein. Based on databases and calculations, Wolfpsort estimates the probability at which site within the cell a protein is located. The ratios obtained as a result of Wolfpsort analysis usually express the probabilities for specific sub-cellular regions of the protein.
Phylogenetic analysis
The peptide sequences of Arabidopsis and common bean MADS-box proteins were aligned in MEGA11 using ClustalW algorithm (Hall 2013; Thompson et al. 2003). The phylogenetic tree was then created using the maximum-likelihood (ML) approach with 1000 bootstrap value using MEGA11 software (Tamura et al. 2021). The phylogenetic tree was ultimately visualized using iTOL (https://itol.embl.de/) online tool (Letunic and Bork 2007).
Chromosomal distribution and duplication analysis of PvMADS genes
The P. vulgaris genomic feature file (GFF3) was downloaded from the JGI Data Portal, and chromosomal locations were plotted using TBtools software (http://cj-chen.github.io/tbtools/). The genomes of specific organisms were acquired from Phytozome v13, and orthologous gene pairs were identified using TBtools MScanX module. In order to comprehend the syntenic connections between MADS-box genes, Dual Synteny Plotter software was utilized to create synteny maps. Collinearity analysis confirmed paralog relationships and then visualized with the Circos tool in TBtools software (Chen et al. 2020). The CLUSTALW program was used to predict the peptide sequences of the transcribed PvMADS genes. The Ka/Ks calculator was utilized to evaluate the selection pressure. Therefore, the veracity of the identified duplication connections was verified utilizing this information (Suyama et al. 2006).
Gene structure, protein motifs, and homology analysis
The exon–intron pattern of PvMADS genes was visualized using the GFF3 file in Tbtools software (Chen et al. 2020). MEME (https://meme-suite.org/meme/) was used for motif extraction of PvMADS proteins (Bailey et al. 2006). Sequence of Motif 1, three-dimensional structure, was obtained using AlfaFold (Jumper et al. 2021).
miRNA and promoter prediction of PvMADS genes
To investigate the cis-regulatory components in the promoter region, the 1.5 kb genomic sequence of each PvMADS gene was extracted from the Phytozome v13 database. Potential cis-regulatory elements in the promoter sequence were evaluated using the PlantCARE online tool (https://bioinformatics.psb.ugent.be/webtools/plantcare/html/). Potential miRNA prediction was obtained using the psRNATarget (http://plantgrn.noble.org/psRNATarget) online tool. PvMADS genes targeted by miRNAs were then visualized using Cytoscape 3.9.1 program (Smoot et al. 2011).
Investigation of expression level of PvMADS genes in several tissues and under drought
Expression levels of PvMADS genes in several plant organs at different growth stages were retrieved from Phytozome v13. The expression levels in silico were measured using FPKM (fragments per kilobase million). The FPKM values were transformed into log2FC values and a heatmap was then generated utilizing the Heatmap module in TBtools. Expression levels of PvMADS genes under drought stresses were calculated using RNA-seq data from the sequence reading archive (SRA) by Illumina sequencing. SRA data with accession numbers SRR8284481 (drought stress leaf) and SRR8284480 (control leaf) were used as previously described by Aygören et al. (2023). Hierarchical clustering heatmaps were created using TBtools.
Plant materials, environmental factors, and stress management
In this study, “Zülbiye,” a Turkish common bean variety, was included in experimental studies since it has been shown to be sensitive to various abiotic stress factors such as drought and high salinity in previous studies (Yıldız et al 2021; Güler et al. 2012). The plant seeds were gifted by “Traditional Regional Agricultural Research Institute in Eskişehir, Turkey.” Followingly, uniform looking seeds were selected and kept in distilled water for 2 h prior to sowing into the weight adjusted pots. Then, the pots were taken into controlled plant growing chamber which was set to + 25 °C, 70% humidity, and 20000 LUX light exposure (16 h day, 8 h night). From the first day, the shelf positions of the pots in the growing chamber were changed regularly. On the 8th day after planting, plants were separated into two groups, and control group plants were continued to be watered; however, water was withheld for stress group plants. Plant weights were taken regularly every 2 days from the first day of stress application. At the end of the 16th day, as the control group plant samples were continued to be watered, an increase was observed in the growth, height, and weight of the leaves. The stress-exposed plants were observed to be shorter than the control group plants; their leaves were curved and pale, and their weights continuously dropped because of water loss. Followingly, the plants were sampled and were crushed with liquid nitrogen in a mortar and then stored at − 80 °C (Büyük and Aras 2017; Aygören et al. 2023).
RNA isolation, cDNA synthesis, and qPCR analyzes
RNA was extracted using the Machery-Nagel NucleoSpin® RNA Kit. RNAs were then subjected to quality control analysis using NanoDrop Lite Spectrophotometer (NanoDrop Technologies, Wilmington, DE, USA) and agarose gel electrophoresis. For complementary DNA synthesis, the Roche (USA) cDNA synthesis kit was employed. PrimerQuest Tool (https://www.idtdna.com) was used to construct the primers, and primer sequences were presented in Supplementary Table 6. iTaq Universal SYBR Green Supermix (Biorad, USA) was used for RT-qPCR reactions following the conditions previously suggested by Büyük et al. (2022). The Light Cycler® Nano qPCR Instrument was used to perform RT-qPCR reactions (Roche, USA). The ACT (Actin) gene was employed for the standardization of RT-qPCR data through the 2−∆∆CT formula (Livak and Schmittgen 2001). GraphPad Prism 7 software was used for statistical analysis, and Fisher’s least significant difference test at 0.05 significance level was applied with two-way ANOVA method.
Results and discussion
Identification, characterization, and phylogenetic analysis of MADS-box genes in P. vulgaris
Common bean (P. vulgaris L.) is a seasonal vegetable that is high in protein, vitamins, fiber, and antioxidants. For hundreds of years, it has been grown for human food with its green or dried parts all over the world (Vougeleka et al. 2023). Recently, the developing high-quality varieties through novel breeding technologies, enhancing production and quality efficiency, have become important research areas in agricultural biotechnology (Naqvi et al. 2022).
Sequencing technologies have enabled the rapid release of genome sequences, and as such, many gene families have become the focus of attention in recent years. Transcription factor gene families have gained popularity in agricultural biotechnology for the last decades since they interact with cis-acting regions of the genes to control the expression of gene of interest. The structure of MADS-box genes in several plant species has been well studied, and previous studies have indicated the importance of MADS-box transcription factors in growth and abiotic stress response of plants (Parenicova et al. 2003; Zhao et al. 2011). Due to many of these reasons, the MADS-box gene family has the potential to be a valuable resource in promoting the development of genetically modified crops and conventional breeding techniques to improve agricultural yield (Lee et al. 2008b; Khong et al. 2015; Theisse and Becker 2003).
The MADS-box gene family has been systematically characterized in diverse plants up to date, including Arabidopsis thaliana (n = 109) (Parenicova et al. 2003), Populus trichocarpa (n = 105) (Leseberg et al. 2006), Solanum tuberosum (n = 156), Solanum lycopersicum (n = 131) (Zou et al. 2017), Glycine max (n = 106), Malus domestica (n = 146) (Tian et al. 2015), Oryza sativa (n = 75) (Arora et al. 2007), Vitis vinifera (n = 32) (Díaz-Riquelme et al. 2009), Cucumis sativus (n = 43) (Hu and Liu 2012), Prunus mume (n = 80) (Xu et al. 2014), Zea mays L. (n = 75) (Zhang et al. 2012), and Sorghum bicolor (n = 65) (Zhao et al. 2011).
In this study, 79 MADS-box genes have been identified in common bean genome, and they were named from PvMADS-01 to PvMADS-79 according to their chromosomal positions (Supplementary Table 1). Moreover, physico-chemical properties of PvMADS proteins were determined using their peptide sequences. Accordingly, the length and isoelectric points (pI) of PvMADS proteins were found to be ranged from 88 (PvMADS-41) to 381 (PvMADS-03) and from 4.6 (PvMADS-03) to 9.8 (PvMADS-41), respectively. In comparison, 58 PvMADS proteins were determined to be basic (pI > 7) and 21 PvMADS proteins to be acidic (pI < 7) with molecular weights ranging from 10.15 (PvMADS-41) to 42.02 (PvMADS-03) kDa.
In this study, three protein characteristics were also evaluated: (1) aliphatic index, (2) instability index, and (3) hydropathicity (GRAVY). The aliphatic index (1) measures the number and proportion of aliphatic (hydrocarbon-containing) amino acids in a protein’s amino acid sequence. This is used to understand the structural properties and functions of proteins. Aliphatic amino acids can form the hydrophobic internal regions of proteins or be associated with the membrane permeability of proteins (Avrahami et al. 2001). In the current study, the aliphatic index values of PvMADS proteins were found to be between 61.58 (PvMADS-10) and 98.59 (PvMADS-15).
The instability index (2) measures the instability of a protein, and it is used to determine which regions of a protein can change or transform rapidly in a test tube. This is important for identifying the variable regions of proteins and understanding their function. The proteins with instability index values less than 40 are referred to as stable proteins; however, they are known as unstable proteins with an instability index value more than 40 (Guruprasad et al. 1990). In this study, the instability index values of PvMADS proteins were found to vary between 30.12 (PvMADS-67) and 73.88 (PvMADS-17).
The GRAVY score (3) measures the hydrophobicity or hydrophilicity of a protein. Positive GRAVY scores can indicate a hydrophobic protein region, while negative GRAVY scores can indicate a hydrophilic protein region. Moreover, the GRAVY score is used to understand the solubility of proteins and their cellular localization. In this study, the GRAVY scores except for PvMADS-11 protein were found to be negative which means that PvMADS proteins were hydrophilic.
In addition, the subcellular localization prediction of MADS-box proteins, which is helpful for understanding protein function, was also performed, and most of the PvMADS proteins were found to be localized in the nucleus in this study (Supplementary Table 1). These ratios give clues about cellular localization by indicating in which cell sub-region a protein is more likely to be found. Similar to our findings here, MADS-box proteins were also reported to be located in nucleus in Argania spinosa by Louati et al. (2021) and in Setaria italica L. by Zhao et al. (2021) in recent years.
To investigate the relationships between MADS-box proteins, a phylogenetic tree was constructed using Arabidopsis and common bean MADS-box peptide sequences (Fig. 1). According to the phylogenetic tree, PvMADS proteins were found to be classified into two main classes as type I (M type (10 Ma, 4 Mβ, 21 Mγ, 4 Mδ)) and type II (MIKC type (2 GLO-like, 6 AGL2-like, 2 EF-like, 4 AGL17-like, 7 STMADS11-llike, 4 AG-like, 1 GGM13-like, 6 SQUA-like, 4 TM3-like, 3 AGL6-like, 1 AGL12-like) in accordance with the previous findings of Parenicova et al. (2003) in Arabidopsis. However, no FLC-like genes, which are known to be control the flowering time, have been identified in P. vulgaris in this study. In most recent studies, Cheng et al. (2023) and Zhang et al. (2023) have also been revealed the absence of FLC-like genes in S. edule, Z. mays, S. bicolor, S. italica, rice, B. distachyon, and Z. latifolia plant species.
Recent studies have indicated that the number of type I MADS-box genes decreased dramatically during evolution of monocotyledons (Arora et al. 2007; Zhao et al. 2021). However, in this study, the number of type I (n = 39) and type II (n = 40) MADS-box genes were very close to each other as contradictory to this statement. In the literature, some studies on rice and Arabidopsis showed that type I genes had a faster birth-and-death evolution rates than type II genes. Gene duplication was suggested to be the main reason of the greater birth rate of type I MADS-box genes, and despite type I genes’ high rate of extinction during evolution, their quantity has been maintained due to duplications (Nam et al. 2004).
PvMADS gene chromosomal locations and gene duplication analyses
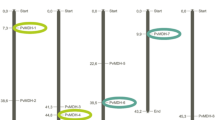
Physical locations of MADS-box genes on P. vulgaris chromosomes were visualized with TBtools software. PvMADS genes showed an unequal distribution across all chromosomes of common bean (Fig. 2). Chromosome 2 was the chromosome with highest number of PvMADS genes (n = 12). The distribution of the remaining PvMADS genes onto chromosomes was as following: 11 PvMADS genes on chromosome 4 and 6; 10 PvMADS genes on chromosome 3; 9 PvMADS genes on chromosome 7; 6 PvMADS genes on chromosome 8; 2 PvMADS genes on chromosome 11; 3 PvMADS genes on chromosome 10; and 2 PvMADS genes on chromosome 5. Meanwhile, two PvMADS genes were found to be present on scaffolds. According to Deng et al. (2023), the exact chromosomal locations of the five PfMADS genes (Paulownia fortunei) could not be ascertained as they were present on scaffolds that had not been stabilized yet (Deng et al. 2023). Similar to the distribution pattern of PvMADS genes in this study, Lakhwani et al. (2022) also reported that the MADS-box genes distributed at different rates across all chromosomes in Musa balbisiana (Lakhwani et al. 2022). According to some studies in eukaryotes, if genes are more widely spread across the genome rather than densely packed on a small number of chromosomes, the organism’s complexity will rise (Toor and MD 2019). Besides, distribution of a gene in different proportions on chromosomes can affect the biological functions, adaptability, and phenotype of the organism. As can be understood, the distribution of gene family members on different chromosomes is important for these gene members to gain different functions (Wang 2020).
Gene duplication are regarded as a crucial element in the evolution of eukaryotes (Taylor and Raes 2004). The discovery of new genes, the development of novel functions, and the extension of gene families are all made possible by the important mechanism of gene duplication. Adaptation of plants to environmental factors, development of new organs, or interactions at the molecular level can appear as evolutionary novelties (Magadum et al. 2013). For example, intensive duplication events have led to expansion of MADS-box genes. Many MADS-box genes have undergone duplication events and differ in their functions (Panchy et al. 2016).
In this study, it was aimed to uncover the molecular mechanism behind the expansion of the PvMADS gene family by detecting paralog genes. Accordingly, 17 paralogous gene pairs were identified, and only two of them were tandemly duplicated gene pairs (PvMADS-19/PvMADS-23 and PvMADS-20/PvMADS-24) (Fig. 2, Supplementary Table 2). The remaining 15 paralogous gene pairs were segmentally duplicated genes (Fig. 2). Ninety-four percent of the detected duplications was observed between type II PvMADS genes. Tandem duplications have been only detected in type II (MIKC) class PvMADS genes, and these were the genes from SQUA-like (PvMADS-19 and PvMADS-23) and from AGL2-like subfamilies (PvMADS-20 and PvMADS-24). This result might be related to late emergence and differentiation of type I genes compared to type II genes during evolutionary time (Zhou et al. 2023). Unsurprisingly, segmentally duplicated gene pairs were classified under same subfamilies in the phylogenetic tree such as PvMADS-05 and PvMADS-79 from AGL2-like, PvMADS-13 and PvMADS-65 from AGL17-like, and PvMADS-50 and PvMADS-68 from STMADS11-like subfamilies. The Ka/Ks ratio can be considered as an indicator of selection pressure during the evolution. Therefore, the Ka/Ks ratio of all homologous genes was calculated to discover the selection pressure on the evolution of MADS-box genes in common bean (Supplementary Table 2). The Ka/Ks ratio was below “1” for all duplications which indicated that the MADS-box genes in the common bean were under strong purifying selection. In a previous study on Camellia sinensis, all of the duplicated MADS-box gene pairs’ Ka/Ks values were also found to be less than 1 similar to our findings here (Hu et al. 2023).
Orthologs are genes that have developed in various species from a common ancestral gene through speciation and typically retain the same function over time. Identification of orthologs is a critical process for better understanding the gene function (Koonin 2005). In this study, orthologous relationships of MADS-box genes between P. vulgaris and other genomes (Arabidopsis thaliana, Glycine max, and Vigna unguiculata) were examined (Fig. 3). As a consequence, there were 38, 61, and 52 orthologous gene pairs between P. vulgaris and those other plants, Arabidopsis, soybean, and kidney bean (Supplementary Table 3). The high number of orthologous relationships between studied genomes might show that the MADS-box gene family spread mostly as a result of duplication events in plant kingdom (Airoldi and Davies 2012).
Synteny analysis of associated genes between P. vulgaris and A. thaliana, G. max, and V. unguiculata. The background displays gray lines indicating the presence of identical linear blocks in the genome of common bean and other plants. The paired syntenic MADS-box genes are highlighted with red lines
Gene structure and conserved protein motifs of PvMADSs
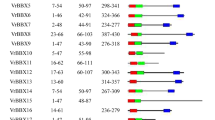
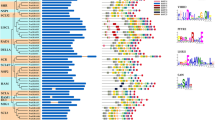
To identify conserved domains in protein structure, the peptide sequences of all PvMADS proteins were retrieved from Phytozome v13. Then, conserved domains were searched in the NCBI CDD database and visualized using TBtools (Fig. 4). The names of detected conserved domains in PvMADS proteins were SRF-TF, MADS superfamily, GOLGA2L5, GOLGA2L5 superfamily, Macoilin, Macoilin superfamily, K domain, K domain super family, CdvA, and CdvA superfamily (Fig. 4).
Human serum response factor (SRF), essential for cell proliferation and differentiation, is a ubiquitous nuclear protein. The SRF core domain plays role for DNA binding and dimerization and belongs to the MADS domain protein family (Pellegrini et al. 1995). In this study, SRF-TF domain was only found in type I PvMADS genes in P. vulgaris. This result was also compatible with results of Alvarez-Buylla et al. (2000) and Parenicova et al. (2003) who previously showed the presence of SRF-like domain proteins in type I MADS-box proteins. The MADS-box domain is commonly known to be associated with the K-box area (Norman et al. 1988). Moreover, Lu et al. (2018) reported that SRF-TF and K domain regions were conserved together between type II MADS-box genes (Lu et al. 2018). Similarly, the findings of our study showed that SRF-TF and K domain regions coexist in type II MADS-box genes.
CdvA domain is known to organize DNA-containing double-helix filaments and is thought to be involved in cell division. Moreover, CdvA, known to be in archaea, is associated with ESCRT (Moriscot et al. 2011). The ESCRT system plays a crucial role in fostering the growth and maturation of plants. In different studies on Arabidopsis, ESCRT mutants resulted in plant death (Haas et al. 2007; Gao et al. 2014; Nagel et al. 2017). GOLGA2L5, representing the conserved domain found in Golgin subfamily A members, is an unstable region enriched with segmental copies containing the core duplicon. Nuclei represent ancestral copies around which additional replication blocks have been formed and correspond to expansions of gene families, some of which show signatures of positive selection. In this study, PvMADS-06 gene was found to contribute to the expansion of PvMADS gene family by establishing both paralogous and orthologous interactions and was also found to have GOLGA2L5 and GOLGA2L5 superfamily domains.
Introns have been shown to influence gene expression at several levels in higher organisms. The primary aim of introns is to translate diverse proteins by generating various combinations of exons through alternative splicing mechanism, which raises the proteome’s quality. Considering that the splicing efficiency of several tRNAs and mRNAs is differentially altered in coffee and rice plants under abiotic stress conditions (e.g., drought, cold, and heat), it was determined that introns respond to abiotic stresses by differentially regulating gene expression (Dinh et al. 2016).
The exon and intron structures of PvMADS genes in P. vulgaris genome were obtained using the General feature file (GFF3). Accordingly, the number of introns in PvMADS genes was found to be varied between 0 and 11 (Fig. 4). The majority of type I MADS-box genes were found to have only one exon with a short sequence in Arabidopsis, soybean, rice, and apple (Ren et al. 2022; Shah et al. 2022; Zhang et al. 2022b). In this study, type I PvMADS genes were also found to include single exon; however, type II PvMADS genes were found to contain more than one intron. On the other hand, different number of introns might be correlated to different evolution rates of PvMADS genes. Similarly, Lai et al. (2022) determined that SiMADS genes exhibited different exon–intron structures in Setaria italica recently.
Conserved motif structures of PvMADS proteins were also investigated in the current study. Motif 1, which was found to be conserved in all proteins, was searched using AlphaFold in order to check for the presence of MADS-box domain (Fig. 5). As a result of this analysis, the Motif 1 (MADS-box domain) was found to be composed of α-Helices and β-sheets layers and had an antiparallel beta-sheet structure. In a study of Tian et al. (2015), the conserved α-helices and β-sheet structures were also detected in apple MADS-box proteins similar to our findings. Motif composition and distribution pattern showed that PvMADS proteins from the same subfamily mostly contained similar motifs in their protein structure. Motif 8/3/4/6/10/9 were frequently seen in type I group PvMADS proteins, while motif 5/7/2 were mostly observed in type II group PvMADS proteins. Some PvMADS proteins were found to differ from other members of the same subclass in terms of certain motifs. For example, some of the Mγ subfamily PvMADS genes contain motif 9, while others contain motif 3/4/6. The evolution of MADS-box genes in P. vulgaris may account for these differences.
Cis-regulatory element analysis in PvMADS promoters
The cis regulatory elements of the PvMADS genes were investigated, and the following elements were found in different ratios: 88% BOX4, 73% ARE, 55% ABRE, 49% TCT motif, 43% TC-rich repeats, 32% AE-box, 29% TCA-element, 26% MRE, 25% MBS, 20% circadian, 7.5% 3-AF1 binding site, 7.5% ACE (Fig. 6, Supplementary Table 4). These cis-regulatory elements were categorized according to their functions in the cell (Fig. 7).
The WUN motif, LTR, MBS, ARE, and TC rich repeats were cis-regulatory elements belonging to the stress category. Zhao et al. (2020) found that pomegranate MADS-box genes also contained rich repeats of stress-responsive cis elements. Similarly, Tanin et al. (2022) showed that MADS-box genes, which were responsible for multi-abiotic stress tolerance in wheat, were found to contain same cis-regulatory elements in their promoter regions. Additionally, Wang et al. (2018) reported that similar cis-regulatory elements were also detected in bZIP genes which respond to salt and drought stress in V. radiata and V. angularis.
ABRE, CGTCA motif, GACG motif, AuxRR core, TATC box, TGA element, GARE motif, TGA box, and P box were detected cis-regulatory elements in PvMADS genes, and they are known to associate with hormone response in plants. ABRE participates in ABA signaling pathway and is involved in the control of diverse abiotic stressors. According to Fujita et al. (2005), ABRE was linked with drought tolerance in Arabidopsis thaliana (Fujita et al. 2005). Kim et al. (2011) demonstrated that the DREB2A gene, which regulates drought-inducible genes in Arabidopsis, contains the ABRE promoter sequence.
The detected cis regulatory elements of MADS-box genes in the development category were GCN4_motif, MBSI, CAT-box, CCAAT-box, O2-site, HD-Zip 1, MSA-like, RY-element, and circadian. On the other hand, the cis-regulatory elements in the light response category were Box 4, G-Box, GATA-motif, TCT-motif, AE-box, I-box, GT1-motif, 3-AF1 binding site, GA-motif, TCCC-motif, chs-CMA1a, chs-CMA2a, LS7, MRE, AT1-motif ATCT-motif, Gap-box, ACE, Sp1, Box II, AAAC-motif, LAMP-element, ATC-motif, and ACA-motif. Shariatipour and Heidari (2018) explored the genes and their governing mechanisms during the period of drought stress in Arabidopsis and found that 2558 gene entries in root and 3691 gene entries in shoot tissues exhibited considerably distinct expressions as compared to the control state. Certain cis-regulatory elements (ABRE, P-box, TATC-box, CGTCA-motif, GARE-motif, GATA- motifi, TCT-motif, GT1-motif, Box 4, G-Box, I-box, LAMP-elemanı, Sp1, MBS, TC-rich repeats, TCA-element) have been identified as the most important elements in the transcriptional regulatory activity of differentially expressed genes.
In silico identification of miRNAs targeting PvMADS genes
MicroRNAs (miRNAs) are non-coding endogenous small RNAs (20–24 nucleotides long) that suppress the expression of some genes in eukaryotes at post-transcriptional level (Saini et al. 2008; Ha and Kim 2014). Furthermore, miRNAs participate in a wide range of biological mechanisms that affect plant development and biotic/abiotic response (Šečić et al. 2021).
A latest research has indicated that the mechanism of miRNA-mediated regulation can enhance the agronomic characteristics of plants and confer resistance to abiotic stress, thereby promoting sustainable agricultural production. Plants experiencing stress can improve their ability to withstand against harsh environmental conditions like drought by controlling the gene expression of specific miRNAs to ensure survival and adaptation (Sun et al. 2021). Studies on miRNAs provide important information about plant stress resistance and may reveal new insights into the adaptation mechanism in plants (Zhang et al. 2022a). As a result of miRNA analysis, miRNAs targeting PvMADS genes were determined, and various miRNA families have been found (Fig. 8, Supplementary Table 5).
The genes targeted by miRNA-171 were involved in the development of roots and leaves, signaling via photochromes, polarity of lateral organs, formation of meristems, development of vasculature, and response to stress in Arabidopsis thaliana (Lee et al. 2008a; Wang et al. 2010; Zhu et al. 2015). Furthermore, miRNA-171 has been discovered to control the reaction to different non-living environmental factors like salinity, drought, and heat in plants (Sunkar and Zhu 2004; Nguyen et al. 2015; Vakilian 2020).
Visentin et al. (2020) revealed a molecular link between drought and miR156 in tomato. In a previous study, drought recovery has been associated with controlling of stomatal behavior by miR156. Furthermore, miR156 has been found to regulate drought tolerance in alfalfa (Feyissa et al. 2019).
Mohsenifard et al. (2017) showed that the regulation of mir156 and mir396 under drought can induce drought tolerance by triggering GRF and MYB. In another study on rice, Nishad et al. (2022) determined that miRNA530 may play a vital role in salinity tolerance by changing its expression level.
Expression levels of PvMADS genes in different tissues of P. vulgaris
To examine the tissue-specific expression profiles of PvMADS, expression data in 11 tissues at different developmental stages were examined (Fig. 9a). According to the findings, expression levels were observed on different tissues with different levels Similarly, Zhao et al. (2021) demonstrated that the SiMADS-box genes had remarkably variable expression profiles in different plant tissues. According to the results of our study, among all PvMADS genes, PvMADS-51 showed the highest expression level in flower buds. Intron-rich PvMADS-07 is the gene with relatively higher expression levels in tissues than the other PvMADS genes. The genes that exhibit high expression levels in most of the plant tissues were generally found to contain larger introns. For example, PvMADS-74 contains the longest intron, which exhibits high expression level in different tissues. It was also suggested by Zhou et al. (2023) that the genes which have more or longer introns are expressed more than others in plant cells.
a Heatmap representing expression profiles of PvMADS in various tissues. b Heatmap representing the expression profiles of PvMADS under drought. The color scale shows log2FC values from high (red color) to low (white color). Twelve RNA samples were extracted in two replicates under either drought or control conditions. Gray rectangles indicate no data results
Investigation of PvMADS gene expression levels against drought stress by RNAseq data and RT-qPCR analysis
Transcription factors and hormones regulate important aspects of growth, development, and environmental stress responses. Plant development and survival are significantly impacted by abiotic stressors such as salinity, heat, cold, and flooding. To overcome these situations, plants have developed adaptation, tolerance, complex sensing, and signaling mechanisms. To create strategies to shield plants from the detrimental impacts of abiotic stresses and to ensure future demand for herbal products, it is necessary to explore stress response mechanisms at molecular level (Nair et al. 2022; Waadt et al. 2022). Drought is a serious environmental constraint for plant productivity. Considering the climate crisis, its effects reach a devastating level (Farooq et al. 2018). Reactive oxygen species (ROS) production is inevitably increased under drought stress in various cellular compartments, specifically in chloroplasts and mitochondria. Nonetheless, when faced with drought, plants elicit an extensive array of reactions, spanning from physiological, biochemical, and molecular levels (Kaur and Asthir 2017). In the light of this information, a heat-map graph was created using drought RNAseq data as described in M&M section (Fig. 9b). Accordingly, expression changes in PvMADS genes were investigated between control and drought stress conditions. As a result of the analysis, PvMADS-10, PvMADS-32, PvMADS-46, PvMADS-58, PvMADS-77 genes were found to be upregulated dramatically in response to drought (Fig. 9b).
According to studies in different plant species, abiotic stress response and hormone regulation mechanisms are regulated by MADS-box genes (Castelán-Muñoz et al. 2019; Khong et al. 2015). To deeply understand the roles of PvMADS genes in drought response, their expression profiles were assessed using RT-qPCR technique, and 5 PvMADS genes (type I PvMADS genes: PvMADS-10, PvMADS-32, PvMADS-46, PvMADS-58, PvMADS-77) were found to be upregulated in both leaf and root tissues in consistent with RNAseq data (Fig. 10).
RT-qPCR analysis of selected candidate PvMADS genes. Relative gene expression levels of PvMADS genes in leaf (shown as black box) and root (shown as gray box) tissues. Errors bars represent the standard errors of three biological and two technical replicates, and asterisks signify the level of statistical significance (*p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001, ns = non-significant)
In a previous study, Zhao et al. (2021) were generated transgenic Arabidopsis and rice lines by overexpressing SiMADS51 gene, and they were then subjected those lines to drought stress. As a result of their study, the overexpression of SiMADS51 has been shown to led to drought sensitivity in Arabidopsis and rice lines. Saha et al. (2015) found that 8 BrMADS genes were induced in B. rapa against drought stress (Abbas et al. 2014). Additionaly, Yin et al. (2017) measured the physiological parameters of WT and transgenic plants in order to better evaluate the damage caused by drought stress, and it was revealed that MADS-box genes increased drought tolerance in tomato. Similar to those previous findings in the literature, we suggest that the PvMADS genes may have important roles in abiotic stress response in P. vulgaris. In addition, the study by Ayra et al. (2021) revealed that MADS-box genes are positive regulators of the bean-rhizobia symbiosis with functions such as root development, rhizobial infection, nodule organogenesis/function, and autoregulation of nodulation.
Conclusions
This study offers insight into the MADS genes in the common bean. A total of 79 PvMADS genes were identified. Exon–intron and motif structure and phylogenetic relationships with A. thaliana were characterized. Considering evolutionary relationships, PvMADS genes were divided into two main classes as type I and type II. The MADS domain was found in type I and type II PvMADS proteins, while the K domain was found to be specific to type II PvMADS proteins. All chromosomes of the common bean were discovered to contain PvMADS genes. The orthologs of A. thaliana, G. max, and V. unguiculata provide an evolutionary insight, indicating that expression profiles in different tissues may be involved in developmental stages. Information from RNA-seq data was confirmed by RT-qPCR, indicating their potential role in stress regulation, revealing their involvement in drought stress tolerance. Overall, the data gave a clearer picture of the PvMADS genes in common bean, which could then be used to illustrate how they affect growth and development and increase the level of resistance to drought conditions. This study might be a reference for the future studies with MADS genes in different plant species.
Data availability
All data generated or analyzed during this study are included in this published article.
References
Abbas AK, Lichtman AH, Pillai S (2014) Cellular and molecular immunology E-book. Elsevier Health Sciences
Airoldi CA, Davies B (2012) Gene duplication and the evolution of plant MADS-box transcription factors. J Genet Genomics 39(4):157–165
Alvarez-Buylla ER, Pelaz S, Liljegren SJ, Gold SE, Burgeff C, Ditta GS, Ribas de Pouplana L, Martínez-Castilla L, Yanofsky MF (2000) "An ancestral MADS-box gene duplication occurred before the divergence of plants and animals. “ Proc Natl Acad Sci 97(10):5328–33
Anlarsal AE, Yücel C, Özveren D (2000) Çukurova koşullarında bazı fasulye (Phaseolus vulgaris L.) çeşitlerinde tane verimi ve verimle ilgili özellikler ile bu özellikler arası ilişkilerin saptanması. Turk J Agric For 24(1):19–29
Arora R, Agarwal P, Ray S, Singh AK, Singh VP, Tyagi AK, Kapoor S (2007) MADS-box gene family in rice: genome-wide identification, organization and expression profiling during reproductive development and stress. BMC Genomics 8(1):1–21
Avrahami D, Oren Z, Shai Y (2001) ’Effect of multiple aliphatic amino acids substitutions on the structure, function, and mode of action of diastereomeric membrane active peptides. Biochemistry 40(42):12591–12603
Aygören AS, Güneş E, Muslu S, Kasapoğlu AG, Yiğider E, Aydın M, Büyük İ, İlhan E (2023) Genome-wide analysis and characterization of SABATH gene family in Phaseolus vulgaris genotypes subject to melatonin under drought and salinity stresses. Plant Mol Biol Report 41(2):242–259
Ayra L, Reyero-Saavedra MDR, Isidra-Arellano MC, Lozano L, Ramírez M, Leija A, Hernández G (2021) Control of the rhizobia nitrogen-fixing symbiosis by common bean MADS-domain/AGL transcription factors. Front Plant Sci 12:679463
Bailey TL, Williams N, Misleh C, Li WW (2006) MEME: discovering and analyzing DNA and protein sequence motifs. Nucleic Acids Res 34(suppl_2):W369–W373
Büyük İ, Aras S (2017) Genome-wide in silico identification, characterization and transcriptional analysis of the family of growth-regulating factors in common bean (Phaseolus vulgaris L.) subjected to polyethylene glycol-induced drought stress. Arch Biol Sci 69(1):5–14
Büyük İ, Okay A, Aras S (2022) Identification and characterization of SRS genes in Phaseolus vulgaris genome and their responses under salt stress. Biochem Genet 60:482–503
Castelán-Muñoz N, Herrera J, Cajero-Sánchez W, Arrizubieta M, Trejo C, García-Ponce B, MdlP S, Álvarez-Buylla ER, Garay-Arroyo A (2019) MADS-box genes are key components of genetic regulatory networks involved in abiotic stress and plastic developmental responses in plants. Front Plant Sci 10:853
Chen C, Chen H, Zhang Y, Thomas HR, Frank MH, He Y, Xia R (2020) TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol Plant 13(8):1194–1202
Chen R, Ma J, Luo D, Hou X, Ma F, Zhang Y, Meng Y, Zhang H, Guo W (2019) CaMADS, a MADS-box transcription factor from pepper, plays an important role in the response to cold, salt, and osmotic stress. Plant Sci 280:164–174
Cheng S, Jia M, Su L, Liu X, Chu Q, He Z, Zhou X, Lu W, Jiang C (2023) Genome-wide identification of the MADS-box gene family during male and female flower development in chayote (Sechium edule). Int J Mol Sci 24(7):6114
Coen ES, Meyerowitz EM (1991) The war of the whorls: genetic interactions controlling flower development. Nature 353(6339):31–37
Deng M, Dong Y, Xu S, Huang S, Zhai X, Fan G (2023) Genome-wide identification and expression of the Paulownia fortunei MADS-box gene family in response to phytoplasma infection. Genes 14(3):696
Dinh SN, Sai TZT, Nawaz G, Lee K, Kang H (2016) Abiotic stresses affect differently the intron splicing and expression of chloroplast genes in coffee plants (Coffea arabica) and rice (Oryza sativa). J Plant Physiol 201:85–94
Díaz-Riquelme J, Lijavetzky D, Martínez-Zapater JM, Carmona MJ (2009) Genome-wide analysis of MIKCC-type MADS box genes in grapevine. Plant Physiol 149(1):354–369
Ditta G, Pinyopich A, Robles P, Pelaz S, Yanofsky MF (2004) The SEP4 gene of Arabidopsis thaliana functions in floral organ and meristem identity. Curr Biol 14(21):1935–1940
Farooq M, Ullah A, Lee DJ, Alghamdi SS, Siddique KH (2018) Desi chickpea genotypes tolerate drought stress better than kabuli types by modulating germination metabolism, trehalose accumulation, and carbon assimilation. Plant Physiol Biochem 126:47–54
Feyissa BA, Arshad M, Gruber MY, Kohalmi SE, Hannoufa A (2019) The interplay between miR156/SPL13 and DFR/WD40–1 regulate drought tolerance in alfalfa. BMC Plant Biol 19(1):1–19
Fujita Y, Fujita M, Satoh R, Maruyama K, Parvez MM, Seki M, Hiratsu K, Ohme-Takagi M, Shinozaki K, Yamaguchi-Shinozaki K (2005) AREB1 is a transcription activator of novel ABRE-dependent ABA signaling that enhances drought stress tolerance in Arabidopsis. Plant Cell 17(12):3470–3488
Gao C, Luo M, Zhao Q, Yang R, Cui Y, Zeng Y, Xia J, Jiang L (2014) A unique plant ESCRT component, FREE1, regulates multivesicular body protein sorting and plant growth. Curr Biol 24(21):2556–2563
Guruprasad K, Reddy BB, Pandit MW (1990) Correlation between stability of a protein and its dipeptide composition: a novel approach for predicting in vivo stability of a protein from its primary sequence. Protein Eng Des Sel 4(2):155–161
Güler NS, Sağlam A, Demiralay M, KADIOĞLU A, (2012) Apoplastic and symplastic solute concentrations contribute to osmotic adjustment in bean genotypes during drought stress. Turk J Biol 36(2):151–160
Goodstein DM, Shu S, Howson R, Neupane R, Hayes RD, Fazo J, Mitros T, Dirks W, Hellsten U, Putnam N (2012) Phytozome: a comparative platform for green plant genomics. Nucleic Acids Res 40(D1):D1178–D1186
Guo X, Chen G, Cui B, Gao Q, Guo JE, Li A, Zhang L, Hu Z (2016) Solanum lycopersicum agamous-like MADS-box protein AGL15-like gene, SlMBP11, confers salt stress tolerance. Mol Breeding 36:1–15
Ha M, Kim N (2014) Regulation of microRNA biogenesis. Nat Rev Mol Cell Biol 15:509–24
Haas TJ, Sliwinski MK, Martínez DE, Preuss M, Ebine K, Ueda T, Nielsen E, Odorizzi G, Otegui MS (2007) The Arabidopsis AAA ATPase SKD1 is involved in multivesicular endosome function and interacts with its positive regulator LYST-INTERACTING PROTEIN5. Plant Cell 19(4):1295–1312
Hall BG (2013) Building phylogenetic trees from molecular data with MEGA. Mol Biol Evol 30(5):1229–1235
Henschel K, Kofuji R, Hasebe M, Saedler H, Münster T, Theißen G (2002) Two ancient classes of MIKC-type MADS-box genes are present in the moss Physcomitrella patens. Mol Biol Evol 19(6):801–814
Hu L, Liu S (2012) Genome-wide analysis of the MADS-box gene family in cucumber. Genome 55(3):245–256
Hu J, Chen Q, Idrees A, Bi W, Lai Z, Sun Y (2023) Structural and functional analysis of the MADS-box genes reveals their functions in cold stress responses and flower development in tea plant (Camellia sinensis). Plants 12(16):2929
Idiku L, Odabaşioğlu E, Uskutoğlu D (2022) İkinci Ürün Olarak Yetiştirilen Fasulye Çeşitlerinin Bitki Aksamının Besin Değerleri. Toprak Bilimi Bitki Besleme Dergisi 10(1):1–9
Jia J, Zhao P, Cheng L, Yuan G, Yang W, Liu S, Chen S, Qi D, Liu G, Li X (2018) MADS-box family genes in sheepgrass and their involvement in abiotic stress responses. BMC Plant Biol 18:1–11
Jumper J, Evans R, Pritzel A, Green T, Figurnov M, Ronneberger O, ... Hassabis D (2021) Highly accurate protein structure prediction with AlphaFold. Nature 596(7873):583–589
Kaur G, Asthir B (2017) Molecular responses to drought stress in plants. Biol Plant 61(2):201–209
Khong GN, Pati PK, Richaud F, Parizot B, Bidzinski P, Mai CD, Bès M, Bourrié I, Meynard D, Beeckman T (2015) OsMADS26 negatively regulates resistance to pathogens and drought tolerance in rice. Plant Physiol 169(4):2935–2949
Kim JS, Mizoi J, Yoshida T, Fujita Y, Nakajima J, Ohori T, Todaka D, Nakashima K, Hirayama T, Shinozaki K (2011) An ABRE promoter sequence is involved in osmotic stress-responsive expression of the DREB2A gene, which encodes a transcription factor regulating drought-inducible genes in Arabidopsis. Plant Cell Physiol 52(12):2136–2146
Koonin EV (2005) Orthologs, paralogs, and evolutionary genomics. Annu Rev Genet 39:309–338
Lai D, Yan J, He A, Xue G, Yang H, Feng L, Wei X, Li L, Xiang D, Ruan J (2022) Genome-wide identification, phylogenetic and expression pattern analysis of MADS-box family genes in foxtail millet (Setaria italica). Sci Rep 12(1):1–17
Lakhwani D, Dhar YV, Singh S, Pandey A, Trivedi PK, Asif MH (2022) Genome wide identification of MADS box gene family in Musa balbisiana and their divergence during evolution. Gene 836:146666
Lee MH, Kim B, Song SK, Heo JO, Yu NI, Lee S, Kim M, Kim DG, Sohn SO, Lim CE (2008a) Large-scale analysis of the GRAS gene family in Arabidopsis thaliana. Plant Mol Biol 67(6):659–670
Lee S, Woo YM, Ryu SI, Shin YD, Kim WT, Park KY, Lee IJ, An G (2008b) Further characterization of a rice AGL12 group MADS-box gene, OsMADS26. Plant Physiol 147(1):156–168
Leseberg CH, Li A, Kang H, Duvall M, Mao L (2006) Genome-wide analysis of the MADS-box gene family in Populus trichocarpa. Gene 378:84–94
Letunic I, Bork P (2007) Interactive Tree Of Life (iTOL): an online tool for phylogenetic tree display and annotation. Bioinformatics 23(1):127–128
Liljegren SJ, Ditta GS, Eshed Y, Savidge B, Bowman JL, Yanofsky MF (2000) SHATTERPROOF MADS-box genes control seed dispersal in Arabidopsis. Nature 404(6779):766–770
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2− ΔΔCT method. Methods 25(4):402–408
Louati M, Salazar-Sarasua B, Roque E, Beltrán JP, Salhi Hannachi A, Gómez-Mena C (2021) Isolation and functional analysis of a PISTILLATA-like MADS-Box gene from argan tree (Argania spinosa). Plants 10(8):1665
Lu W, Chen J, Ren X, Yuan J, Han X, Mao L, Ying T, Luo Z (2018) One novel strawberry MADS-box transcription factor FaMADS1a acts as a negative regulator in fruit ripening. Sci Hortic 227:124–131
Ma J, Yang Y, Luo W, Yang C, Ding P, Liu Y, Lan X (2017) Genome-wide identification and analysis of the MADS-box gene family in bread wheat (Triticum aestivum L.). PLoS One 12(7):e0181443
Magadum S, Banerjee U, Murugan P, Gangapur D, Ravikesavan R (2013) Gene duplication as a major force in evolution. J Genet 92(1):155–161
Martel C, Vrebalov J, Tafelmeyer P, Giovannoni JJ (2011) The tomato MADS-box transcription factor RIPENING INHIBITOR interacts with promoters involved in numerous ripening processes in a COLORLESS NONRIPENING-dependent manner. Plant Physiol 157(3):1568–1579
Michaels SD, Ditta G, Gustafson-Brown C, Pelaz S, Yanofsky M, Amasino RM (2003) AGL24 acts as a promoter of flowering in Arabidopsis and is positively regulated by vernalization. Plant J 33(5):867–874
Mohsenifard E, Ghabooli M, Mehri N, Bakhshi B (2017) Regulation of miR159 and miR396 mediated by Piriformospora indica confer drought tolerance in rice. Journal of Plant Molecular Breeding 5(1):10–18
Moriscot C, Gribaldo S, Jault JM, Krupovic M, Arnaud J, Jamin M, Schoehn G, Forterre P, Weissenhorn W, Renesto P (2011) Crenarchaeal CdvA forms double-helical filaments containing DNA and interacts with ESCRT-III-like CdvB. PLoS One 6(7):e21921
Nagel MK, Kalinowska K, Vogel K, Reynolds GD, Wu Z, Anzenberger F, Ichikawa M, Tsutsumi C, Sato MH, Kuster B (2017) “Arabidopsis SH3P2 is an ubiquitin-binding protein that functions together with ESCRT-I and the deubiquitylating enzyme AMSH3.” Proc Natl AcadSci 114(34):E7197-E7204
Nair AU, Bhukya DPN, Sunkar R, Chavali S, Allu AD (2022) Molecular basis of priming-induced acquired tolerance to multiple abiotic stresses in plants. J Exp Bot 73(11):3355–3371
Nam J, Kim J, Lee S, An G, Ma H, Nei M (2004) “Type I MADS-box genes have experienced faster birth-and-death evolution than type II MADS-box genes in angiosperms.” Proc Natl Acad Sci 101(7):1910-1915
Naqvi RZ, Siddiqui HA, Mahmood MA, Najeebullah S, Ehsan A, Azhar M, Asif M (2022) Smart breeding approaches in post-genomics era for developing climate-resilient food crops. Front Plant Sci 13:972164. https://doi.org/10.3389/fpls.2022.972164
Nguyen GN, Rothstein SJ, Spangenberg G, Kant S (2015) Role of microRNAs involved in plant response to nitrogen and phosphorous limiting conditions. Front Plant Sci 6:629
Nishad J, Panda AK, Chowrasia S, Nirmala C, Mondal TK (2022) Allantoin mediated regulation of miRNAs for short term salinity stress tolerance in Oryza sativa L. cv. IR-29. J Plant Biochem Biotechnol 31(4): 953–960
Norman C, Runswick M, Pollock R, Treisman R (1988) Isolation and properties of cDNA clones encoding SRF, a transcription factor that binds to the c-fos serum response element. Cell 55(6):989–1003. https://doi.org/10.1016/0092-8674(88)90244-9
Panchy N, Lehti-Shiu M, Shiu SH (2016) Evolution of gene duplication in plants. Plant Physiol 171(4):2294–2316. https://doi.org/10.1104/pp.16.00523
Parenicova L, de Folter S, Kieffer M, Horner DS, Favalli C, Busscher J, Cook HE, Ingram RM, Kater MM, Davies B (2003) Molecular and phylogenetic analyses of the complete MADS-box transcription factor family in Arabidopsis: new openings to the MADS world. Plant Cell 15(7):1538–1551
Pelaz S, Ditta GS, Baumann E, Wisman E, Yanofsky MF (2000) B and C floral organ identity functions require SEPALLATA MADS-box genes. Nature 405(6783):200–203
Pellegrini L, Tan S, Richmond TJ (1995) Structure of serum response factor core bound to DNA. Nature 376(6540):490–498
Pinyopich A, Ditta GS, Savidge B, Liljegren SJ, Baumann E, Wisman E, Yanofsky MF (2003) Assessing the redundancy of MADS-box genes during carpel and ovule development. Nature 424(6944):85–88
Ren Q, Jiang H, Xiang W, Nie Y, Xue S, Zhi H, Li K, Gai J (2022) A MADS-box gene is involved in soybean resistance to multiple Soybean mosaic virus strains. The Crop Journal 10(3):802–808
Riechmann JL, Meyerowitz EM (1997) MADS domain proteins in plant development. Biol Chem 378(10):1079–1102
Saha G, Park JI, Jung HJ, Ahmed NU, Kayum M, Chung MY, Hur Y, Cho YG, Watanabe M, Nou IS (2015) Genome-wide identification and characterization of MADS-box family genes related to organ development and stress resistance in Brassica rapa. BMC Genomics 16(1):1–21. https://doi.org/10.1186/s12864-015-1349-z
Saini S, Dongen H, Enrigh S (2008) miRBase: tools formicroRNA genomics. Nucleic Acids Res 36:D154–D158
Schilling S, Kennedy A, Pan S, Jermiin LS, Melzer R (2020) Genome-wide analysis of MIKC-type MADS-box genes in wheat: pervasive duplications, functional conservation and putative neofunctionalization. New Phytol 225(1):511–529
Šečić E, Kogel KH, Ladera-Carmona MJ (2021) Biotic stress-associated microRNA families in plants. J Plant Physiol 263:153451
Shah L, Sohail A, Ahmad R, Cheng S, Cao L, Wu W (2022) The roles of MADS-box genes from root growth to maturity in Arabidopsis and rice. Agronomy 12(3):582
Shariatipour N, Heidari B (2018) Investigation of drought and salinity tolerance related genes and their regulatory mechanisms in Arabidopsis (Arabidopsis thaliana). Open Bioinforma J 11(1):12
Singh KB, Foley RC, Oñate-Sánchez L (2002) Transcription factors in plant defense and stress responses. Curr Opin Plant Biol 5:430–436
Smoot ME, Ono K, Ruscheinski J, Wang PL, Ideker T (2011) Cytoscape 2.8: new features for data integration and network visualization. Bioinformatics 27(3):431–432
Sun Y, Xiong X, Wang Q, Zhu L, Wang L, He Y, Zeng H (2021) Integrated analysis of small RNA, transcriptome, and degradome sequencing reveals the MiR156, MiR5488 and MiR399 are involved in the regulation of male sterility in PTGMS rice. Int J Mol Sci 22(5):2260
Sunkar R, Zhu JK (2004) Novel and stress-regulated microRNAs and other small RNAs from Arabidopsis. Plant Cell 16(8):2001–2019
Suyama M, Torrents D, Bork P (2006) PAL2NAL: robust conversion of protein sequence alignments into the corresponding codon alignments. Nucleic Acids Res 34(suppl_2):W609–W612
Tamura K, Stecher G, Kumar S (2021) MEGA11: molecular evolutionary genetics analysis version 11. Mol Biol Evol 38:3022–3027
Tanin MJ, Saini DK, Sandhu KS, Pal N, Gudi S, Chaudhary J, Sharma A (2022) Consensus genomic regions associated with multiple abiotic stress tolerance in wheat and implications for wheat breeding. Sci Rep 12(1):13680
Taylor JS, Raes J (2004) Duplication and divergence: the evolution. Annu Rev Genet 38:615–643
Thei G, Saedler H (2001) Plant biology: floral quartets. Nature 409(6819):469–472
Theisse G, Becker A (2003) The major clades of MADS-box genes and their role in the development and evolution of fl flowering plants. Mol Phylogenet Evol 29:464–489
Theißen G, Kim JT, Saedler H (1996) Classification and phylogeny of the MADS-box multigene family suggest defined roles of MADS-box gene subfamilies in the morphological evolution of eukaryotes. J Mol Evol 43(5):484–516
Thompson JD, Gibson TJ, Higgins DG (2003) Multiple sequence alignment using ClustalW and ClustalX. Curr Prot Bioinformat (1):2–3
Tian Y, Dong Q, Ji Z, Chi F, Cong P, Zhou Z (2015) Genome-wide identification and analysis of the MADS-box gene family in apple. Gene 555(2):277–290
Toor AA, MD, AAT (2019) “On the order of gene distribution on chromosomes across the animal kingdom.” arXiv preprint arXiv:1912.07154
Vakilian KA (2020) Determination of nitrogen deficiency-related microRNAs in plants using fluorescence quenching of graphene oxide nanosheets. Mol Cell Probes 52:101576
Visentin I, Pagliarani C, Deva E, Caracci A, Turečková V, Novák O, Lovisolo C, Schubert A, Cardinale F (2020) A novel strigolactone-miR156 module controls stomatal behaviour during drought recovery. Plant, Cell Environ 43(7):1613–1624
Vougeleka V, Savvas D, Ntatsi G, Ellinas G, Zacharis A, Iannetta PP, Mylona P, Saitanis CJ (2023) Impact of the rootstock genotype on the performance of grafted common bean (Phaseolus vulgaris L.) cultivars. Sci Hortic 311:111821
Waadt R, Seller CA, Hsu PK, Takahashi Y, Munemasa S, Schroeder JI (2022) Plant hormone regulation of abiotic stress responses. Nat Rev Mol Cell Biol 23(10):680–694
Wang RRC (2020) Chromosomal distribution of genes conferring tolerance to abiotic stresses versus that of genes controlling resistance to biotic stresses in plants. Int J Mol Sci 21(5):1820
Wang L, Mai YX, Zhang YC, Luo Q, Yang HQ (2010) MicroRNA171c-targeted SCL6-II, SCL6-III, and SCL6-IV genes regulate shoot branching in Arabidopsis. Mol Plant 3(5):794–806
Wang L, Zhu J, Li X, Wang S, Wu J (2018) Salt and drought stress and ABA responses related to bZIP genes from V. radiata and V. angularis. Gene 651:152–160
Wei B, Zhang RZ, Guo JJ, Liu DM, Li AL, Fan RC, Mao L, Zhang XQ (2014) Genome-wide analysis of the MADS-box gene family in Brachypodium distachyon. PLoS One 9(1):e84781
Xu Z, Zhang Q, Sun L, Du D, Cheng T, Pan H, Yang W, Wang J (2014) Genome-wide identification, characterisation and expression analysis of the MADS-box gene family in Prunus mume. Mol Genet Genomics 289:903–920
Yıldız S, Okay A, Büyük İ (2021) Defining the roles of PvMDH genes in response to salt stress and detailed characterization of the gene family. J Plant BiochemBiotechnol 31:380–393
Yin W, Hu Z, Hu J, Zhu Z, Yu X, Cui B, Chen G (2017) Tomato (Solanum lycopersicum) MADS-box transcription factor SlMBP8 regulates drought, salt tolerance and stress-related genes. Plant Growth Regul 83:55–68
Zhang L, Xu Y, Ma R (2008) Molecular cloning, identification, and chromosomal localization of two MADS box genes in peach (Prunus persica). J Genet Genomics 35(6):365–372
Zhang Z, Li H, Zhang D, Liu Y, Fu J, Shi Y, Song Y, Wang T, Li Y (2012) Characterization and expression analysis of six MADS-box genes in maize (Zea mays L.). J Plant Physiol 169(8):797–806
Zhang F, Yang J, Zhang N, Wu J, Si H (2022a) Roles of microRNAs in abiotic stress response and characteristics regulation of plant. Front Plant Sci 13:919243
Zhang S, Yao J, Wang L, Wu N, van Nocker S, Li Z, Gao M, Wang X (2022b) Role of grapevine SEPALLATA‐related MADS‐box gene VvMADS39 in flower and ovule development. The Plant Journal 111(6):1565–1579
Zhang Z, Xiao M, Song S, Jiang Y, Zhu X, Shi L, Zheng X, Jiang J, Miao M (2023) Genome-wide identification, classification and expression analyses of MADS-box genes reveal their role in stem gall formation and expansion of Zizania latifolia. Agronomy 13(7):1758
Zhao W, Zhang LL, Xu ZS, Fu L, Pang HX, Ma YZ, Min DH (2021) Genome-wide analysis of MADS-Box genes in foxtail millet (Setaria italica L.) and functional assessment of the role of SiMADS51 in the drought stress response. Front Plant Sci 12:659474
Zhao Y, Li X, Chen W, Peng X, Cheng X, Zhu S, Cheng B (2011) “Whole-genome survey and characterization of MADS-box gene family in maize and sorghum.” Plant Cell Tiss Organ Cult (PCTOC) 105:159–173
Zhao Y, Zhao H, Wang Y, Zhang X, Zhao X, Yuan Z (2020) Genome-wide identification and expression analysis of MIKC-type MADS-box gene family in Punica granatum L. Agronomy 10(8):1197
Zhou P, Qu Y, Wang Z, Huang B, Wen Q, Xin Y, Ni Z, Xu L (2023) Gene structural specificity and expression of MADS-box gene family in Camellia chekiangoleosa. Int J Mol Sci 24(4):3434
Zhu X, Leng X, Sun X, Mu Q, Wang B, Li X, Wang C, Fang J (2015) “Discovery of conservation and diversification of miR171 genes by phylogenetic analysis based on global genomes.” Plant Genome 8(2). https://doi.org/10.3835/plantgenome2014.10.0076
Zou H, Jakovlić I, Chen R, Zhang D, Zhang J, Li WX, Wang GT (2017) The complete mitochondrial genome of parasitic nematode Camallanus cotti: extreme discontinuity in the rate of mitogenomic architecture evolution within the Chromadorea class. BMC Genomics 18(1):1–17
Funding
Open access funding provided by the Scientific and Technological Research Council of Türkiye (TÜBİTAK). This work was supported by the Ankara University Research Foundation (Project no.: FHD-2022–2339).
Author information
Authors and Affiliations
Contributions
SA and IB conceived, planned and oversaw the experiments. TK, ŞD and AO carried out the experiments on the bioinformatic analysis, AO and SŞ carried out experiments on qRT-PCR analysis. SA and IB analyzed and integrated the datasets and drafted the manuscript. SA and IB critically read and contributed to improve.
Corresponding authors
Ethics declarations
Ethics approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Consent to participate and for publication
Not applicable.
Conflict of interest
The authors declare no competing interests.
Additional information
Handling Editor: Néstor Carrillo.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Okay, A., Kırlıoğlu, T., Durdu, Y.Ş. et al. Omics approaches to understand the MADS-box gene family in common bean (Phaseolus vulgaris L.) against drought stress. Protoplasma 261, 709–724 (2024). https://doi.org/10.1007/s00709-024-01928-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00709-024-01928-z