Abstract
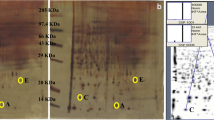
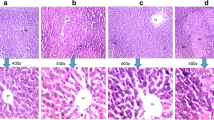
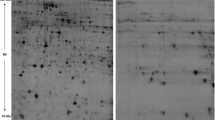
Hepatocellular carcinoma (HCC) is one of the most common cancers associated with high mortality rate. Understanding of events leading to HCC pathophysiology is essential for its better management. We earlier reported development of a novel rodent model by administrating chemical carcinogens, DEN, and 2-AAF for study of HCC at very early stage. 2D-Electrophoresis analysis of total serum proteins identified several differentially expressed proteins in animals undergoing tumorigenesis. MALDI-TOF-MS/MS analyses were performed to characterize the differentially expressed proteins. Further real-time PCR analyses were taken place to quantify the transcript expression for the identified proteins at HCC initiation and tumor stages. Considering protein-protein interactions among the experimentally identified proteins and their interacting neighbors, a protein network has been analyzed that provided further insight into molecular events taking place during HCC development. Histological changes confirmed HCC initiation and hepatotumorigenesis at 1 and 4 months post carcinogen treatment, respectively. Four differentially expressed proteins were identified which were further characterized as regulator of G protein signaling 1 (RGS1), sepiapterin reductase (SPR), similar to zinc finger and BTB domain–containing protein 21 isoform X2 (ZNF295), and a hypothetical protein CXorf58 homolog. Quantification of transcripts for these proteins revealed elevation in their expression both at initiation and tumorigenesis stages. The study deciphers the regulatory role of these proteins during HCC progression.






Similar content being viewed by others
References
Baas A, Boudeau J, Sapkota GP, Smit L, Medema R, Morrice NA, Alessi DR, Clevers HC (2003) Activation of the tumour suppressor kinase LKB1 by the STE20-like pseudokinase STRAD. EMBO J 22:3062–3072
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254. https://doi.org/10.1016/0003-2697(76)90527-3
Faraj Shaglouf LH, Ranjpour M, Wajid S, Jain SK (2020) Elevated expression of cellular SYNE1, MMP10, and GTPase1 and their regulatory role in hepatocellular carcinoma progression. Protoplasma 257:157–167. https://doi.org/10.1007/s00709-019-01423-w
Feinmesser RL, Wicks SJ, Taverner CJ, Chantry A (1999) Ca2+/calmodulin-dependent kinase II phosphorylates the epidermal growth factor receptor on multiple sites in the cytoplasmic tail and serine 744 within the kinase domain to regulate signal generation. J Biol Chem 274:16168–16173
Franz M, Rodriguez H, Lopes C, Zuberi K, Montojo J, Bader GD, Morris Q (2018) GeneMANIA update 2018. Nucleic Acids Res 46:W60–w64. https://doi.org/10.1093/nar/gky311
Fujimoto K, Takahashi SY, Katoh S (2002) Mutational analysis of sites in sepiapterin reductase phosphorylated by Ca2+/calmodulin-dependent protein kinase II. Biochim Biophys Acta 1594:191–198
Fuks F, Burgers WA, Godin N, Kasai M, Kouzarides T (2001) Dnmt3a binds deacetylases and is recruited by a sequence-specific repressor to silence transcription. EMBO J 20:2536–2544. https://doi.org/10.1093/emboj/20.10.2536
Gao H, Yu Z, Bi D, Jiang L, Cui Y, Sun J, Ma R (2007) Akt/PKB interacts with the histone H3 methyltransferase SETDB1 and coordinates to silence gene expression. Mol Cell Biochem 305:35–44. https://doi.org/10.1007/s11010-007-9525-3
Haruki H, Hovius R, Pedersen MG, Johnsson K (2016) Tetrahydrobiopterin biosynthesis as a potential target of the kynurenine pathway metabolite xanthurenic acid. J Biol Chem 291:652–657. https://doi.org/10.1074/jbc.C115.680488
Hurst JH, Hooks SB (2009) Regulator of G protein signaling (RGS) proteins in cancer biology. Biochem Pharmacol 78:1289–1297. https://doi.org/10.1016/j.bcp.2009.06.028
Jen J, Wang Y-C (2016) Zinc finger proteins in cancer progression. J Biomed Sci 23:53. https://doi.org/10.1186/s12929-016-0269-9
Katare DP, Malik S, Mani RJ, Ranjpour M, Jain SK (2018) Novel mutations in transthyretin gene associated with hepatocellular carcinoma. Mol Carcinog 57:70–77. https://doi.org/10.1002/mc.22732
Kusaba H, Ghosh P, Derin R, Buchholz M, Sasaki C, Madara K, Longo DL (2005) Interleukin-12-induced interferon-gamma production by human peripheral blood T cells is regulated by mammalian target of rapamycin (mTOR). J Biol Chem 280:1037–1043. https://doi.org/10.1074/jbc.M405204200
Li H, Rauch T, Chen ZX, Szabo PE, Riggs AD, Pfeifer GP (2006) The histone methyltransferase SETDB1 and the DNA methyltransferase DNMT3A interact directly and localize to promoters silenced in cancer cells. J Biol Chem 281:19489–19500. https://doi.org/10.1074/jbc.M513249200
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)). Method Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Malik S, Bhatnagar S, Chaudhary N, Katare DP, Jain S (2013a) DEN+ 2-AAF-induced multistep hepatotumorigenesis in Wistar rats: supportive evidence and insights. Protoplasma 250:175–183
Malik S, Bhatnagar S, Chaudhary N, Katare DP, Jain SK (2013b) Elevated expression of complement C3 protein in chemically induced hepatotumorogenesis in Wistar rats: a correlative proteomics and histopathological study. Exp Toxicol Pathol 65(6):767–773. https://doi.org/10.1016/j.etp.2012.11.003
Mossanen JC, Kohlhepp M, Wehr A, Krenkel O, Liepelt A, Roeth AA, Möckel D, Heymann F, Lammers T, Gassler N, Hermann J, Jankowski J, Neumann UP, Luedde T, Trautwein C, Tacke F (2019) CXCR6 inhibits hepatocarcinogenesis by promoting natural killer T- and CD4+ T-cell-dependent control of senescence. Gastroenterology 156(6):1877–1889.e4. https://doi.org/10.1053/j.gastro.2019.01.2472019.01.247
Ogunwobi OO et al (2019) Mechanisms of hepatocellular carcinoma progression. World J Gastroenterol 25:2279–2293. https://doi.org/10.3748/wjg.v25.i19.2279
Ranjpour M, Katare DP, Wajid S, Jain SK (2018a) HCC specific protein network involving interactions of EGFR with A-Raf and transthyretin: experimental analysis and computational biology correlates. Anti Cancer Agents Med Chem. https://doi.org/10.2174/1871520618666180507141632
Ranjpour M, Wajid S, Jain SK (2018b) Elevated expression of A-Raf and FA2H in hepatocellular carcinoma is associated with lipid metabolism dysregulation and cancer progression. Anti Cancer Agents Med Chem. https://doi.org/10.2174/1871520618666181015142810
Ranjpour M, Wajid S, Jain SK (2020) Elevated expression of cytosolic phospholipase A2 delta is associated with lipid metabolism dysregulation during hepatocellular carcinoma. Progression Cell J 22:17–22. https://doi.org/10.22074/cellj.2020.6527
Rentsch CA et al (2006) Differential expression of TGFβ-stimulated clone 22 in normal prostate and prostate cancer. Int J Cancer 118:899–906
Roh J et al (2016) RGS1 expression is associated with poor prognosis in multiple myeloma. J Clin Pathol 70:jclinpath-2016. https://doi.org/10.1136/jclinpath-2016-203713
Sarbassov DD, Guertin DA, Ali SM, Sabatini DM (2005) Phosphorylation and regulation of Akt/PKB by the rictor-mTOR complex. Science 307:1098–1101. https://doi.org/10.1126/science.1106148
Sethakorn N, Dulin NO (2013) RGS expression in cancer: oncomining the cancer microarray data. J Recept Signal Transduct Res 33:166–171. https://doi.org/10.3109/10799893.2013.773450
Shao H, Cheng HY, Cook RG, Tweardy DJ (2003) Identification and characterization of signal transducer and activator of transcription 3 recruitment sites within the epidermal growth factor receptor. Cancer Res 63:3923–3930
Tsai CC et al (2006) Gene expression analysis of human hepatocellular carcinoma by using full-length cDNA library. J Biomed Sci 13:241–249. https://doi.org/10.1007/s11373-005-9062-6
van Meer S, de Man RA, Siersema PD, van Erpecum KJ (2013) Surveillance for hepatocellular carcinoma in chronic liver disease: evidence and controversies. World J Gastroenterol 19:6744–6756. https://doi.org/10.3748/wjg.v19.i40.6744
Wang Q et al (2011) Zinc finger protein ZBTB20 expression is increased in hepatocellular carcinoma and associated with poor prognosis. BMC Cancer 11:271
Wu Y et al (2020) Sepiapterin reductase promotes hepatocellular carcinoma progression via FoxO3a/Bim signaling in a nonenzymatic manner. Cell Death Dis 11:248–248. https://doi.org/10.1038/s41419-020-2471-7
Yoo DR, Chong SA, Nam MJ (2009) Proteome profiling of arsenic trioxide-treated human hepatic cancer cells. Cancer Genomics Proteomics 6:269–274
Yoon CH et al (2012) Crucial role of TSC-22 in preventing the proteasomal degradation of p53 in cervical cancer. PLoS One 7:e42006. https://doi.org/10.1371/journal.pone.0042006
Zhen C, Zhu C, Chen H, Xiong Y, Tan J, Chen D, Li J (2017) Systematic analysis of molecular mechanisms for HCC metastasis via text mining approach. Oncotarget 8:13909–13916. https://doi.org/10.18632/oncotarget.14692
Funding
The work was supported by a MRP grant from the University Grant Commission (UGC) to S. K. Jain [Grant # 40-155/2011 (SR)]. M. R. is grateful to ICCR, Govt. of India, for their financial support [Grant # 2-261/2013-14/ISD-II].
Author information
Authors and Affiliations
Contributions
All authors have contributed equally and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval
Experiments were conducted following the guidelines of the “Committee for the Purpose of Control and Supervision of Experiments on Animals” (approval number granted by the ethics committee of Jamia Hamdard University is 908/CPCSEA), Government of India. The study was performed in accordance with the ethical standards.
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Handling Editor: Jörn Bullerdiek
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Ranjpour, M., Wajid, S. & Jain, S.K. Elevated expression of sepiapterin reductase, regulator of G protein signaling 1, hypothetical protein CXorf58 homolog, and zinc finger and BTB domain–containing protein 21 isoform X2 is associated with progression of hepatocellular carcinoma. Protoplasma 258, 1133–1143 (2021). https://doi.org/10.1007/s00709-021-01632-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00709-021-01632-2