Abstract
Squalene epoxidase (SQE) is a crucial regulatory enzyme for the biosynthesis of several important classes of compounds including sterols and triterpenoids. The present paper identified and characterised five SQE genes (GgSQE1 to GgSQE5) from Glycyrrhiza glabra through transcriptome data mining and homology-based cloning, for the first time. The phylogenetic analysis implied their functional divergence. The ORF corresponding to one of the five SQEs, namely, GgSQE1, was cloned and studied for its function in a heterologous system, following transient and stable expressions. The transient expression followed by GgSQE1 encoding protein purification suggested approximately 58.0-kDa protein following the predicted molecular mass of the deduced protein. The gene expression profiling based on qRT-PCR indicated its highest expression (6.4-folds) in the 10-month-old roots. Furthermore, ABA (12.4-folds) and GA3 (2.47) treatments upregulated the expression of GgSQE1 in the shoots after 10 and 12 hours, respectively, which was also reflected in glycyrrhizin accumulation. The inductive effects of ABA and GA3 over GgSQE1 expression were also confirmed through functional analysis of GgSQE1 promoters using GUS fusion construct. Stable constitutive expression of GgSQE1 in Nicotiana tabacum modulated the sterol contents. The study could pave the way for understanding the metabolic flux regulation concerning biosynthesis of related sterols and triterpenoids.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Glycyrrhizin, an oleanane-type triterpenoid saponin, is obtained from the dried roots (stolons) of Glycyrrhiza species (G. uralensis, G. glabra and G. inflate) which is widely recognised as a natural sweetener, flavouring agent and crude drug in Japanese, Chinese and Indian systems of medicine (Shibata et al. 2003). It is the major bioactive ingredient of several herbals, possessing immunomodulatory, antiulcer, anti-allergic and antiviral activities including HIV and severe acute respiratory syndrome (SARS)-associated coronavirus (Asl and Hosseinzadeh 2008; Cinatl et al. 2003; Deng 2001; Fiore et al. 2008; Khan et al. 2013; Menegazzi et al. 2008; Shibata 2000).
According to the centre for the promotion of imports from developing countries (https://www.cbi.eu/market-information/natural-food-additives/liquorice), the market demand for the root is high, with predicted annual global market growth of 4% between 2017 and 2025, which is predominantly met by wild resources. In the USA, the demand for liquorice root is estimated to reach ~US$ 2.9 Bn by 2029, up from ~US$1.9 Bn in 2019 (https://www.transparencymarketresearch.com/licorice-extracts-market.html). For obvious reasons, there have been studies focusing on unravelling the metabolic steps, genes and enzymes involved in the biosynthesis of glycyrrhizin. The core biosynthetic pathway (Supplementary Fig. 1), comprising of early and late stages, has been established for the glycyrrhizin biosynthesis (Ramilowski et al. 2013; Seki et al. 2009; Seki et al. 2008; Seki et al. 2011; Seki et al. 2015). The early steps commence with squalene, derived from the isoprene pool, primarily from the cytosolic mevalonate pathway (MVA) of the cell. Squalene is catalysed by squalene epoxidase (SQE) to form a key intermediate molecule, 2,3-oxidosqualene, which is the precursor of diverse triterpenoid and sterol class of compounds. In G. glabra, 2,3-oxidosqualene is subsequently cyclised to β-amyrin by β-amyrin synthase (bAS), the first committed step for glycyrrhizin biosynthesis (Dubey et al. 2003). Recent studies, involving transcriptome (Ramilowski et al. 2013), draft genome (Mochida et al. 2017) sequencing and in vitro enzyme assays (Xu et al. 2016), have created a substantial knowledge base of genes involved in glycyrrhizin biosynthesis. Overall, squalene synthase, β-amyrin synthase, two cytochrome-P450s (CYP88D6 and CYP72A154) and one UDP-dependent glycosyltransferase (UGT) have been shown to be involved in glycyrrhizin biosynthesis (He et al. 2018; Seki et al. 2009; Seki et al. 2011). However, understanding of the regulation of the pathway is still largely unknown. Also, the critical upstream regulatory branch-point SQE gene has not been cloned and characterised from Glycyrrhiza species, so far.
Squalene epoxidase, a non-cytochrome P450-type monooxygenase, catalyses the first oxygenation and rate-limiting step, converting squalene into oxidosqualene. The product, 2,3-oxidosqualene, is subsequently cyclised by several triterpene synthases to generate an array of triterpenoid skeletons (Phillips et al. 2006). These skeletons, in turn, act as precursors for all the known angiospermic cyclic triterpenoid and phytosterol class of compounds including membrane sterols, brassinosteroid phytohormones and non-steroidal triterpenoids (Brueckner and Tissier 2013). Squalene epoxidase is a pivotal enzyme for studying in context of substrate flux distribution towards biosynthesis of functionally and structurally diverse triterpenoids and sterols. Literature divulged several plants with multiple squalene epoxidases (Laranjeira et al. 2007). Multiple predicted squalene epoxidase (SQE) enzymes had been biochemically characterised from Medicago truncatula (Suzuki et al. 2002), Brassica napus (Schafer et al. 1999), Populus trichocarpa, Oryza sativa (Rasbery et al. 2007), Arabidopsis (Rasbery 2007) and several other plant species. In Arabidopsis, for example, six SQE (1 to 6) proteins have been identified and functionally characterised. Out of the six, only three Arabidopsis SQEs (SQE 1 to 3) displayed epoxidase activity in the functional assay, and the rest (SQE4 to 6) had no illustrative activity, suggesting the functional divergence of SQE homologs. The null sqe1 mutants in Arabidopsis displayed pleiotropic growth defects and accumulation of squalene. The NCBI database reports 426 squalene epoxidases (https://www.ncbi.nlm.nih.gov) from plants with 49 submissions from Fabaceae family covering only nine genera. Among the Fabaceae family, maximum submissions are reported from Glycine max (10) followed by Vigna angularis (9), while no gene is reported from the genus Glycyrrhiza.
More than 200 triterpenoids have been reported from liquorice species (Gupta et al. 2013; Wang et al. 2013). Despite its pivotal role as a branch-point enzyme in the structural and functional diversification of triterpenoid, the SQE genes have not been studied for their function in G. glabra. In the current study, therefore, we carried out transcriptome-wide identification of SQE genes and characterisation of GgSQE1 in G. glabra, a commercially important plant species. In the present study, the GgSQE1 has been studied in details including its phylogeny, regulation and functional attributes. Our study, herein, develops a solid foundation for carrying out further investigations pertaining to the regulatory role of all the SQEs in triterpenoid biosynthesis, in general, and glycyrrhizin biosynthesis, in particular.
Material and methods
Plant material
Three months old in vitro-grown G. glabra plants were used for the gene cloning and elicitor-based experiments. Glycyrrhiza glabra plant was subcultured and multiplied in the tissue culture lab, hardening unit and the experimental farms of the CSIR-IIIM Lab following the published protocol (Gupta et al. 2013). Temporal and spatial analyses were performed using in vitro-regenerated plantlets, plants grown under glass house conditions and field-grown plants. For elicitation studies, in vitro plants, cultured in liquid MS medium (3% sucrose supplemented with 0.05 μM of NAA), as described earlier from our lab (Gupta et al. 2013; Khan et al. 2016), were used. Each set of treatment and control was employed in triplicates and harvested on defined time. The sample was divided into root and shoot and immediately frozen into liquid nitrogen for expression studies and chemical analysis. For transient expression, Nicotiana benthamiana seeds were germinated in autoclaved soil mix and grown in a growth chamber at 23 °C under 16 h: 8-h day-night photoperiod for 6 weeks. The in vitro-grown N. tabacum var. petit havana plants were used for stable transformation experiment.
Full-length cloning of GgSQE1 gene
RNA isolation and cDNA synthesis
Total RNA from the root and shoot samples (control, transformed and treated plants) were extracted using PureLink RNA mini kit (Ambion, USA) following the manufacturer’s instructions. First-strand cDNA was synthesised by reverse transcribing total RNA (2 μg) in a final reaction volume of 20 μl using Superscript III first-strand cDNA synthesis kit (Invitrogen, USA) according to the manufacturer’s instructions. cDNA for the real-time expression analysis, elicitation experiments and functional analysis was synthesised from RNA extracted from G. glabra, N. tabacum and N. benthamiana plants following the instructions of the kit.
Cloning of full-length GgSQE1gene
Degenerate primers (Supplementary Table 1) designed from the conserved sequences of SQE genes of related plant species (Supplementary Table 2) were used for the amplification of the core cDNA fragment of GgSQE1gene under following conditions: 35 cycles of 95 °C for 1 min, 45 °C for the 30s and 72 °C for 1 min followed by a final extension of 72 °C for 7 min. A 537-bp DNA fragment was amplified and cloned into a cloning vector (pJET1.2, Fermentas, USA) and subjected to nucleotide sequencing. This fragment was subsequently used for designing gene-specific primers (GSP) for RACE-PCR and amplification of full-length gene using SMARTer RACE cDNA amplification kit (Clonetech, USA) as per the manufacturer’s instructions. The initial 5’and 3’RACE-PCR reactions were performed using respective outer gene-specific primers (GgSQEP1 and GgSQEP3) with UPM primer (Universal Primer A Mix). Primary PCR product was used as a template for nested 5’ and 3’RACE-PCR reactions with respective nested primers (GgSQEP2 and GgSQEP4) and NUP primer (Nested Universal Primer A Mix). Both initial and nested PCR reactions were performed under the following conditions: 4 min at 94 °C, 35 cycles (30s at 94 °C, 30s at 60 °C, 2 min at 72 °C) and 10 min at 72 °C. The amplified fragments of 5’ and 3’RACE were cloned into a cloning vector (pJET1.2) and transformed into DH5α cells (Invitrogen, USA). Based on the antibiotic screening and PCR analysis, positive clones were selected and sequenced using ABI 3130xl genetic analyser (Applied Biosystems, USA). The amplified sequence of full-length GgSQE1gene (1590 bp) was cloned, sequenced and submitted to NCBI (MG763680). Primers used for full-length gene cloning and real-time and semi-quantitative expression analyses of GgSQE1gene are listed in Supplementary Table 1.
Isolation and cloning of GgSQE1 promoter
For the isolation of the upstream region of the GgSQE1, Genome Walker Universal Kit (Clonetech, USA) was used as per the instructions in the kit. Briefly, the genomic DNA was isolated using CTAB method and digested with four sets of restriction enzymes (DraI, EcoRI, PvuII and StuI), independently to get four genomic libraries (D1, D2, D3, D4). Primary PCR was performed using appropriately diluted genomic libraries as a template with adaptor primer (AP1) and outer gene-specific primer (GSP1) as forward and reverse primers, respectively. The secondary PCR was done using the nested primer (AP2 and GSP2) pairs. The amplified product obtained from the D2 library was purified from agarose gel using the QIAquick spin kit (Qiagen) and further cloned into pTZ57R/T cloning vector (Fermentas) and transformed into DH5α cells. The plasmid was isolated from confirmed DH5α transformants and sequenced.
In silico sequence analysis and homology searches and statistical analysis
The sequence data were trimmed using VecScreen software (https://www.ncbi.nlm.nih.gov/tools/vecscreen/) and aligned by BLAST and ClustalW program. The presence of putative cis-regulatory elements was detected by PLACE (http://www.dna.affrc.go.jp/PLACE/signalscan.html) and PlantCARE (plantcare/html) scan tools. The nucleotide sequence was assembled and subjected to BLAST (https://blast.ncbi.nlm.nih.gov) analysis to confirm the gene identity and translated using EXPASY translate tool (http://www.expasy.ch/tools/dna.html) and the properties of deduced amino acid sequences were estimated using ProtParam (http://www.expasy.ch/tools/protparam.html). Structurally and functionally essential regions were identified in the deduced protein sequence by SMART tool. Secondary structure was determined by SOPMA (http://npsa-pbil.ibcp.fr) program. Hydrophobicity analysis was done using Kyte-Doolittle (http://gcat.davidson.edu/DGPB/kd/kyte-doolittle.htlm) and TMHMM (http://www.cbs.dtu.dk/services/) web tools. 3D protein structure elucidation was done by Pyre2. Conserved residue analysis of SQE1 protein was performed using ConSurf and ConSeq web servers (http://consurf. tau.ac.il/overview.htm) (Berezin et al. 2004). The phylogenetic tree was constructed using MEGA6.0 software (Koichiro Tamura et al. 2013). For the identification of other homologs of GgSQE1, the homology searches were carried out against an in-house transcriptome resource of G. glabra (BioProject ID: PRJNA664636) as well as against the transcriptome database of G. uralensis (Mochida et al. 2017) (http://ngs-data-archive.psc.riken.jp/Gur-genome/download.pl). The qRT-PCR data was compared and analysed using one-way ANOVA and two-way ANOVA (for treatments) in GraphPad Prism 8 software. Values were expressed as means ± SD representing three independent biological samples, each with three technical replicates. Differences were scored as statistical significance with P value ranging between < 0.0001 to < 0.05 levels. For chemical analysis, one-way ANOVA was used.
Real-time expression analysis of GgSQE1 gene
The expression analysis of all the five identified GgSQE gene homologs was carried out in different tissues of the regenerated plant, and GgSQE1 was also investigated after ABA and GA3 treatments in the in vitro plants at different time points (0, 4, 8, 10, 12 and 16 hours) through qRT-PCR (Step One Plus Real-Time PCR System, Applied Biosystems, USA) using gene-specific primers (Supplementary Table 1). Also, the relative expression analysis of GgSQE1 in the infiltrated and mock-treated control leaves of N. benthamiana plants was conducted on Step One Plus Real-Time PCR System (Applied Biosystems, USA). Real-time PCR reactions were performed using cDNA (1 μl), Premix Ex Taq (Takara, Otsu, Japan) (5 μl) and 2 pmol of forward and reverse primers each, in a final volume of 10 μl. Cycling conditions optimised were 95 °C for 30 s, 95 °C for 3 s followed by 40 cycles of 58 °C for 1 min and dissociation stage at 95 °C for 15 s, 60 °C for 30 s and 95 °C for 15 s. The specificity of each primer pair was verified by analysing the melting curve of PCR products. β-actin was kept as an internal control. Relative quantification analysis was done by the ΔΔCT method (Schmittgen and Livak 2008). Three biological replicates for each sample were used for real-time PCR analysis, and three technical replicates were analysed for each biological replicate.
Heterologous expression of GgSQE1 in Nicotiana species
Gene and promoter construct and Agrobacterium infiltration for heterologous expression
The GgSQE1 gene was subcloned into two expression vectors pBI121 and pEAQ-HT (Sainsbury et al. 2009) for stable and transient expressions, respectively. For stable expression, SacI and XbaI sites were introduced in the gene and subcloned into the plasmid pBI121 to get pBI121-GgSQE1 gene construct (Supplementary Figure 2a), while for transient expression, it was subcloned into the pEAQ-HT vector (Supplementary Figure 2b) using AgeI and SmaI restriction sites in frame with a C-terminus histidine (6xHIS) affinity tag (Supplementary Table 1). For promoter cloning, the identified upstream fragment (750 bp) was subcloned into pBI121 vector at Hind III and BamHI sites replacing the CaMV35S promoter with GgSQE1 promoter (Supplementary Figure 2c). The pBI121 vector, two gene constructs (pBI121-GgSQE1 and pEAQ-HT-GgSQE1) and promoter construct (pBI121-GgSQE1P:GUS) were individually transformed into A. tumefaciens strain GV3101 by the freeze-thaw method (Weigel and Glazebrook 2006).
Heterologous expression
A single colony of transformed Agrobacterium (GV3101) cell was inoculated in LB medium (5 ml) with appropriate antibiotics and grown for 24 hours at 28 °C under shaking conditions at 180 rpm. The inoculum (1%) from primary culture was used for the secondary culture, which was incubated for 12 hours under similar conditions. The culture was pelleted and washed thrice using infiltration buffer containing 1-M MgCl2, 1-M MES pH 5.6 and 200-mM acetosyringone and finally re-suspended into the infiltration buffer. The mix was kept at room temperature for 3 hours. Subsequently, the suspension (OD~0.6) was infiltrated into the leaves of N. benthamiana plants (4–5 weeks old) by pressing needleless syringe against the abaxial side of the leaf. The labelled transformed plants were kept in growth chamber until further analysis. Infiltrated leaf tissue was collected after every 2 days post-infiltration for checking the transcript levels of the transformed gene using RT-PCR. The leaves were harvested 5 days post-infiltration (dpi) for recombinant protein extraction and analysis.
For the stable and constitutive expression of the GgSQE1, N. tabacum was transformed with pBI121-GgSQE1 vector through Agrobacterium-mediated transformation following the leaf disk method (Horsch et al. 1989). The N. tabacum plants were also transformed with empty vector pBI121, following the same method. The regenerated shoots under the antibiotic selection pressure (kanamycin) were rooted and confirmed for their transgenic nature through PCR with gene-specific primer set. The transgenic lines in T1 generation along with wild-type N. tabacum plants were grown in the growth chamber and chemically analysed for sterols.
Protein purification and western blotting
The recombinant protein with C-terminal 6x-HIS tag was extracted from the infiltrated leaves 5 days post-infiltration (dpi) by flash-freezing them in liquid nitrogen. Leaves were crushed to obtain fine powder using a pre-cooled motor pestle. The recombinant protein was purified, as described earlier (de Souza 2015). Briefly, pre-grind leaf tissue was homogenised in three volumes of extraction buffer containing 25-mM TRIS, 150-mM NaCl, β-mercaptoethanol, 1% Triton X-100, 2% PVP, 10% glycerol, 0.1% TWEEN-20 and 1x protease inhibitor cocktail from CST (Cell Signaling Technologies) to the final volume, and centrifuged at 20,000×g for 5 min at 4 °C. This was followed by purification using Nickel NTA beads. The pre-washed Nickel NTA beads (100 μl) were added to crude protein extract, and the mixture was incubated for 2 h at 4 °C for efficient binding of tagged proteins to the beads. Imidazole (10 μM) was added to the washing buffer to minimise non-specific binding of the histidine-rich protein. Finally, the bound protein was eluted using elution buffer (extraction buffer containing imidazole concentration ranging between 25 and 200 mM), at 4 °C, and was loaded on SDS Gel (10%) as per the standard protocol, which was further confirmed by western blotting using anti-HIS antibody (Supplementary Figure 3).
Assay of the promoter activity
The leaves of N. benthamiana plants were Agro-infiltrated with the constructs (empty pBI121 vector and pBI121-GgSQE1-Prom-GUS), followed by ABA and GA3 (100 μM each) treatments. Abscisic acid was prepared in aqueous solution while GA3 was prepared in aqua-ethanolic solution with 0.05% Tween 20 and sprayed over the plant. Mock treatment was given with autoclaved distilled water and aqua-ethanolic solution to the respective plant. Promoter infiltrated and elicited leaves were harvested after 16 hours along with the mock-treated controls. All the samples were analysed qualitatively using GUS assay and quantitatively using MUG assays. GUS assays were performed on transiently transformed leaves of N. benthamiana with pBI121-GgSQE1-Prom-GUS and pBI121 empty vector. The substrate, X-gluc (5-bromo-4-chloro-3-indolyl-β-glucuronide), was prepared in 20-mM sodium phosphate buffer (pH 7.2) containing 0.1% Triton X-100, 10-mM EDTA, 50-mM potassium ferricyanide and 50-mM potassium ferrocyanide. Infiltrated leaves were cut into small discs and incubated with the substrate overnight at 37 °C following the published protocol (Janssen and Gardner 1990). Subsequently, the tissues were cleared with serial transfers in 70% alcohol to remove excess blue colour, for photography. The relative expression intensity was quantified using MUG assay. For MUG assay, the Agro-infiltrated leaves were homogenised in 1-ml extraction buffer containing 10-mM EDTA, 50-mM NaH2PO4 (PH7), 0.1% sodium laurylsarcosine, 10-mM β-mercaptoethanol and 0.1% Triton X-100. The homogenate was centrifuged at 12,000 rpm for 15 min at 4 °C before the supernatant was collected. The protein concentrations were measured by the Bradford method using bovine serum albumin (BSA) as a standard. Fluorometric enzymatic assay was carried out by adding 50 μl of pre-warmed (37 °C) protein extract to 200-μl pre-warmed (37 °C) MUG assay buffer. After incubation at 37 °C for 120 minutes, 10 μl from the reaction mix was added to 190 μl of stop buffer (200-mM sodium carbonate) in a microtitre plate to stop the reaction. Fluorescence was measured using a microplate spectro-fluorometer (1420 Multilabel Counter, PerkinElmer, Finland) at the excitation wavelength of 365 nm and the emission wavelength of 455 nm. All the experiments were performed in triplicates. Untransformed plant and plant having only pBI121construct were used as controls.
Chemical analysis
Extraction
The extraction of phytosterols was done following the protocol published by Xiang et al. (Li, et al. 2016) including the acid hydrolysis and alkaline saponification steps with little modifications. Finally, the dried plant extract generated from the equal weight (1 g) of the starting material was dissolved in methanol before subjecting to GC-MS analysis with an injection volume of 1 μl. The procedure for glycyrrhizin extraction from the in vitro-regenerated G. glabra plants was essentially similar to the earlier published reports (Gupta et al. 2013; Khan et al. 2016).
GC-MS analysis of the transformed N. tabacum plant
GC-MS analysis of the samples was performed on an Agilent 7890A gas chromatograph coupled to an Agilent 5975C inert XL MSD mass spectrometer with triple-axis detector and a MassHunter Workstation software (USA) using a DB-5 (30 m × 0.25 mm i.d. × 0.25 μm film thickness). Helium was used as carrier gas at a flow rate of 0.5 mL/min. Temperature programming was 50 °C hold for 1 minute, 50–250 °C at a rate of 50C/min, and held for 5 min. Mass spectra were recorded in electron impact (EI) mode with ionisation energy of 70 eV and a scan rate of 0.5 s/scan. The scan range was 50–600 amu. Inlet and transfer line temperature were set 250 °C. The split ratio used was 50:1. Component identification was achieved by Wiley and NIST libraries. Compounds were also identified by peak enrichment on co-injection with available authentic standards. Peak area percentages (%) were obtained electronically from the EIC response without the use of correction factors.
LC–MS/MS analysis
The lyophilized ABA- and GA3-treated aerial tissues and the untreated control of the in vitro-regenerated G. glabra plants were extracted and analysed following the published protocol (Khan et al. 2016). Briefly, analytical chromatographic separations of the extract were carried out on a chromolith performance RP-18e column (100 × 4.6 mm, Merck, Germany) protected by a chromolith guard column of the same company. The optimised parameters were column temperature at 30 °C; flow rate at 0.4 ml/min and the sample injection volume was 10 μl. The solvent system optimised was a linear gradient of acetonitrile/water and formic acid. The mobile phase was programmed at acetonitrile 40% for 4 min, 50% for 6 min, 60% for 15 min, 70% for 5 min, 80% for 5 min and from 80 to 40% in 15 min. The detection, identification and quantification of glycyrrhizin were performed on an Agilent 6410 LC/MS–MS (Agilent Technologies, USA) triple-stage quadrupole mass spectrometer equipped with the electrospray ionisation (ESI) interface and liquid chromatography was performed on an Agilent 1260 Infinity (Agilent, USA) quaternary pump equipped with an autosampler, column heater and online degasser. ESI positive mode with single ion monitoring was chosen for the quantification of investigated compounds. All the standards used in the present study were procured from Sigma-Aldrich (USA).
Results
Identification, cloning and characterisation of squalene epoxidase homologs
A full-length CDS, putatively encoding SQE, named as GgSQE1, was established by employing a homology-based cloning approach involving 5’ and 3’ RACE. A 1590-bp CDS of GgSQE1, thus cloned, was predicted to encode a polypeptide of 529 amino acid residues, which showed maximum sequence homology (89%) with the SQE from Astragalus membranaceus (KJ010819.1) (Table 1).The predicted molecular weight was found to be 58.2 kDa having the theoretical pI of 8.35. The estimated half-life of the protein was calculated to be 30 hours (in vitro) and 20 and 10 hours in in vivo systems of yeast and E. coli, predicting it to be an unstable protein (Gao et al. 2016). Although squalene epoxidases of various origins have been investigated with respect to substrate requirements including cofactors and inhibitors, no structural model for the gene is available (Ruckenstuhl et al. 2007). An earlier report by Schreuder (Schreuder et al. 1989) cited p-hydroxybenzoate hydroxylase (PHBH) of Pseudomonas fluorescens to be the prototype of FAD-dependent hydroxylases and the only enzyme in this class of flavin proteins for which the three-dimensional structure is known. However, in the present study, in silico alignment of the deduced protein sequences with PHBH template (Supplementary Fig. 4) showed very low homology (26%). Hence, this template was not considered for homology modelling. Instead, we decided to identify protein template based on protein folding/threading using Phyre2 software. The software selected oxidoreductase (c2qa2A) template for the 3D structure prediction (Fig. 1) of 376 residues (71% query coverage) with 100% confidence level as the single highest scoring template protein (c2qa2A). The predicted 3D structure was comprised of α-helix and β sheets (Fig. 1a), binding sites (Fig. 1b), conserved (green) and variable (violet) amino acid residues (Fig. 1c) and 39-bp long signal peptide at the N-terminal. The protein showed U-shaped transmembrane topology having both N&C terminals in the extracellular matrix (Fig. 1d). The predicted model showed that the residues present at the centre core of the structure, namely, residues ARG 329, ASN 228, VAL189, ASP 95, VAL 103, PRO358, GLY 104, GLU105, THR 188, CYS225, GLU 93 and ILE 92, were involved in ligand molecule interactions (shown in blue). The predicted structure had a major cytosolic region (26–460 aa), three transmembrane regions (6–25 aa; 461–479 aa; and 487–505 aa) and extra cytosolic region (480–486 aa). Further, the study on docking the protein with various related substrates will help in understanding the enzyme-substrate flexibility for metabolic reprogramming the pathways.
The homology modelling of GgSQE1. a Three-dimensional models and conserved residue prediction; b 3D structures of GgSQE1 (cartoon display) as predicted by Phyre2; c predicted ligands (shown in green) with their binding sites as predicted by 3D LigandSite web server. The residues involved in ligand molecule interactions are ARG 329, ASN 228, VAL189, ASP 95, VAL 103, PRO358, GLY 104, GLU105, THR 188, CYS225, GLU 93 and ILE 92 (shown in purple) in the core of the structure. Conserved residue analysis of GgSE1 was performed using ConSeq web servers. Residue conservation from variable to conserved is shown on the given scale, i.e. green (1) to violet (9); d 39-bp long signal peptide at the N-terminal and a U-shaped transmembrane topology having both N&C terminals in the extracellular matrix. The predicted structure had a major cytosolic region (26–460 aa), three transmembrane regions (6–25 aa; 461–479 aa; and 487–505 aa) and an extra cytosolic region (480–486 aa)
The GgSQE1 protein sequence was used to identify SQE homologs in the transcriptome of Glycyrrhiza species. Transcriptome-wide examination of Glycyrrhiza species revealed five putative squalene epoxidase gene homologs (Table 1). The identified GgSQE1–GgSQE5 sequences ranged between 1326 and 1599 bp corresponding to amino acid residue ranging between 441 and 532. Considering the size of the deduced protein sequence, it is possible that GgSQE4 represents a partial CDS. Three putative G. glabra epoxidase enzymes (GgSQE2, GgSQE3 and GgSQE4) were 90–96% homologous to the characterised Ononis spinosa SQE enzyme (AUD09558.1), which is involved in seco-triterpenoid biosynthesis (Almeida et al. 2018) (Table 1). While GgSQE1 was found homologs (89%) to Astragalus membranaceus (AHY94896.1 and AKO83630.1) with a role in triterpenoid biosynthesis, GgSQE5 was 76% homologous to Arabidopsis thaliana squalene epoxidase 3 enzyme (AT4G37760) important for embryo development (Laranjeira et al. 2015). The GgSQE1 sequence was BLAST searched to identify SQE proteins from other plant species for multiple sequence alignment and phylogenetic analysis (http://www.ebi.ac.uk/Tools/msa/muscle/). The multiple sequence alignment of all the five GgSQE proteins (Fig. 2) along with the amino acid residues of the known proteins from other plant species (Supplementary Figure 5) showed several conserved domains. The characteristic flavin adenine dinucleotide (FAD) domain and nicotinamide adenine dinucleotide (NAD) binding site occupied most of the protein. The FAD domain is reported to be involved in the biosynthesis processes of the enzymes (Li et al. 2018; Li et al. 2009). As can be seen from Fig. 2, the N-terminal amino acid residues of the SQE proteins belonging to Fabaceae family were more diverse (less homology) than the middle and C-terminals of the protein. The phylogenetic analysis of the protein sequences illustrated the position of the gene with the other known species (Fig. 3). The tree clustered 40 protein sequences broadly into three major groups with members of the same family grouping together. Group I with 12 members was comprised of two subgroups having six members in each group, belonging to Araliaceous and Fabaceae families. Group II had 11 members, all belonging to the Fabaceae family. This group was, however, subdivided into Fabaceae I with four members and Fabaceae II with seven members. Group III had six members all from the Brassicaceae family assembled into two subgroups.
Multiple sequence alignment of the deduced five amino acid sequences identified from the Glycyrrhiza glabra (MG763680 SQE1); Glycyrrhiza glabra (MN593324 SQE2); Glycyrrhiza glabra (MN593325 SQE3); Glycyrrhiza glabra (MN593326 SQE4); and Glycyrrhiza glabra (MN593328 SQE5) with proteins from other plant species present in NCBI: Ononis spinosa (AUD09558.1); Medicago truncatula (KEH39980.1); Astragalus membranaceus (AHY94896.1); Glycine soja (KHN46220.1); and Cajanus cajan (KYP54510.1) identical residues are marked (*) and characteristic residues (FAD and NAD binding) are underlined with black
Phylogenetic tree constructed using the deduced amino acid sequences from Glycyrrhiza glabra (GgSQE1–GgSQE5) along with 42 other SQE proteins. The protein sequence IDs used in the analysis were BAF79915.1 (Euphorbia tirucalli); MG763680 (glabra SQE1); MN593324 (Glycyrrhiza glabra SQE2); MN593325 (Glycyrrhiza glabra SQE3); MN593326 (Glycyrrhiza glabra SQE4); MN593328 (Glycyrrhiza glabra SQE5); ABC94943.1 (Medicago sativa); APD76301.1 (Medicago sativa); EOY15642.1 (Theobroma cacao); KYP54510.1 (Cajanus cajan); KHG06672.1 (Gossypium arboreum); KEH39981.1 (Medicago truncatula); KEH39980.1 (Medicago truncatula); ACQ90301.1 (Gynostemma pentaphyllum); AFZ93645.1 (Euphorbia lathyris); KYP34787.1 (Cajanus cajan); KEH39980.1 (Medicago truncatula); ALD84329.1 (Morus alba); AFI33134.2 (Eriobotrya japonica); ARA73618.1 (Lepidium apetalum); AT1G58440.1 (Arabidopsis thaliana ATSQE1); AT2G22830 (Arabidopsis thaliana ATSQE2); AT4G37760.1 (Arabidopsis thaliana ATSQE3); AT5G24140.1 (Arabidopsis thaliana ATSQE4); AT5G24150.1 (Arabidopsis thaliana ATSQE5); AT5G24160.1 (Arabidopsis thaliana ATSQE6); AB923908.1 (Botryococcus braunii SQE-I); AB986538.1 (Botryococcus braunii SQE-2); ADC32655.1 (Aralia elata); GU574803.1 (Withania somnifera); AGK62446.1 (Panax quinquefolius); AJV26446.1 (Panax ginseng); ACJ24907.2 (Panax ginseng); AFZ93645.1 (Euphorbia lathyris); ABE60738.1 (Panax notoginseng); BAD15330.1 (Panax ginseng); AHY94896.1 (Astragalus membranaceus); AUD09558.1 (Ononis spinosa); KHN17192.1 (Glycine soja); ACJ05633.1 (Nigella sativa); EOX97273.1 (Theobroma cacao); OAY63657.1 (Ananas comosus). The evolutionary history was inferred using the maximum likelihood method based on the JTT matrix-based model. Evolutionary analyses were conducted in MEGA6 (Koichiro Tamura et al. 2013)
Spatial and temporal expression analysis of squalene epoxidase genes
The tissue-specific expression profiling of all the five identified GgSQE gene homologs (GgSQE1–GgSQE5) was performed using qRT-PCR in the leaf, stem and root tissues of the field-grown plant and the in vitro-regenerated plants (Fig. 4a–e). GgSQE1 expression (Fig. 4a) was the highest in roots of both the in vitro (22.0-folds) and the field plants (16.2-folds) as compared to the control (field stem). Further, the expressions of other GgSQE homologs (GgSQE2–GgSQE5) were highly reduced in all the tissues of the plants under investigation (Fig. 4b–e). The expression profile of GgSQE2 was highly downregulated in all the tissues studied from in vitro and field plants (0.1–0.3), while GgSQE3 had similar expression (1.0–0.8) in all the tissues of the field plant and in vitro root tissue (1.1), while it was highly downregulated in the in vitro leaf (0.1) and stem (0.2) tissues. The expression of GgSQE4 corresponded to GgSQE3 expression pattern, while GgSQE5 was seen to be slightly upregulated (1.1), only in the in vitro root tissue (Fig. 4e). The time-based expression dynamics of GgSQE1 in the in vitro-regenerated tissues of G. glabra (Supplementary Figure 6) revealed upregulation of GgSQE1 in the root tissues of 10 (6.3-folds) and 12 months (5.1-fold) old plantlet, while it was found to be largely unchanged in the in vitro shoot tissues (0.8- to 1.2-folds) under investigation.
Real-time expression analysis of five squalene epoxidase homologs (GgSQE1–GgSQE5) in different tissues (leaf, stem and root) of the field and in vitro-regenerated Glycyrrhiza glabra plants. a GgSQE1; b GgSQE2; c GgSQE3; d GgSQE4; e GgSQE5. The data was compared and analysed using one-way ANOVA in GraphPad Prism 8 software. Values are expressed as means ± SD representing three independent biological samples, each with three technical replicates. Differences were scored as statistical significance at ***P < 0.0001, **P < 0.001 and *P < 0.05. Asterisks indicate the comparison of expression levels of SQE homologs in the field stem with the other tissues of in vitro and field plant
Transcriptional regulation of GgSQE1 gene
In order to understand the transcriptional regulation of GgSQE1 gene, its 750-bp upstream region was cloned and sequenced. The cloned region was predicted to display the presence of various cis-regulatory elements including ABA- and GA-responsive elements (ABRE and P-Box), the binding site for MYB proteins, WRKY71, transcriptional activators like ARR1 and ARR2, etc. (Table 2). The presence of these regulatory sequences points towards their regulatory role on GgSQE1 expression. The promoter activity was investigated by subcloning GgSQE1 promoter into pBI121 plasmid, replacing CaMV35S promoter of the wild-type plasmid and evaluated for GUS expression.
Histochemical analysis
The untreated untransformed (Fig. 5a), untreated empty vector transformed (pBI121-35S-GUS) (Fig. 5b) and untreated SQE1 promoter–transformed (GgSQE1-Prom-GUS) (Fig. 5c) N. benthamiana plants were subjected to ABA and GA3 treatments to understand the transcriptional regulation. The enhanced GUS activity (darkly stained) was seen in the treated leaves (Fig. 5d, e) as compared to the three control leaves (Fig. 5a, b and c) indicating augmented GUS activity in the leaves driven by enhanced promoter activity under the influence of both the hormones. The promoter activity was further quantified using MUG assay (Fig. 5f). The relative MU value showed 3.6-fold and 1.6-folds enhanced GgSQE1 promoter activity under ABA and GA3 treatments, respectively, as compared to the control plants (Fig. 5f). These results prompted us to study the expression of GgSQE1 gene following ABA and GA3 (Fig. 6a) treatments in G. glabra. The RT-PCR–based expression analysis of GgSQE1 under the ABA influence showed maximum expression (12.4-folds) in shoot at 10 hours, while minimum was observed in 4 hours (1.7-fold) as compared to the untreated control shoot. The GA3 treatment, however, was maximum (2.5-folds) at 12 hours, while minimum (1.1-folds) was seen after 10 hours (Fig. 6a).
Histochemical analysis of GUS activity. For qualitative analysis, Nicotiana benthamiana leaves were infiltrated with two controls a distilled water; b empty vector (pBI121-35S-GUS) and c GgSQE1 promoter construct (GgSQE1-Prom-GUS). The GgSQE1-Prom–transformed N. benthamiana leaves were subjected to two hormonal treatments, d ABA and e GA3 (100 μM each). f MUG-based quantitative analysis was performed in Nicotiana benthamiana leaves infiltrated with distilled water (NC) empty vector (pBI121); untreated GgSQE1 promoter construct (SQE1-UT) and GgSQE1 promoter construct treated with ABA (SQE1-ABA) and GA3 (SQE1-GA3). Values are means ± SE of three independent biological replicates each with three technical replicates. SE is represented by bars. Obtained values were compared and analysed with one-way ANOVA using GraphPad Prism 8 software. Differences were scored as statistical significance at ***P < 0.0001, **P < 0.001 and *P < 0.05. Asterisks indicate the comparison of GgSQE1 promotor activity level in treated and untreated Nicotiana benthamiana leaves
Influence of hormones on the gene expression and glycyrrhizin accumulation. a Quantitative real-time expression analysis of squalene epoxidase 1 under the influence of ABA and GA3 hormones in the shoot tissues at 0, 4, 8, 10, 12 and 16 hours post-treatments; b LC-MS–based analysis of glycyrrhizin in the hormone-treated shoot tissues of G. glabra. The data were compared and analysed using two-way ANOVA in GraphPad Prism 8 software. Values are expressed as means ± SD representing three independent biological samples, each with three technical replicates. Differences were scored as statistical significance at *** P < 0.0001, **P < 0.001 and *P < 0.05. Asterisks indicate difference in expression levels of SQE1 in control and treated plants
Functional characterisation of GgSQE1 gene
In order to study the in planta function of the identified GgSQE1, we carried out its heterologous expression in N. benthamiana and N. tabacum. The effectiveness of syringe infiltration of Agrobacterium into plant tissue was optimised using green fluorescent proteins (GFP) in N. benthamiana. The N. benthamiana leaves were infiltrated with Agrobacteria containing pEAQ-HT-GFP construct and expression was observed over the entire leaf area under UV light (395 nm) starting from 2 days post-infection (dpi) to 5 dpi where it was found maximum. No green fluorescence was observed from leaves infiltrated with dH2O, indicating that the fluorescence was specific to the GFP gene and was not the result of background fluorescence from the leaves (Fig. 7a, b). Firstly, to confirm whether the GgSQE1 expression leads to the biosynthesis of the protein of the expected size, transient expression of the open reading frame corresponding to the GgSQE1 along with a C-terminal His tag was carried out in N. benthamiana. The expression vector (Supplementary Figure 2b) used in the present study has been demonstrated to be highly efficient (Fig. 7c) in large-scale production of recombinant protein in N. benthamiana (Sainsbury et al. 2009). Following Agro-infiltration, the transcript level of GgSQE1 was examined at 0, 3, 5, 8, 10 and 12 days post-infiltration (Fig. 7d). The results showed maximum accumulation of transcripts between 3 and 5 days post-infection.
Transient heterologous expression of GgSQE1 gene in Nicotiana benthamiana. a Un-infiltrated control leaves; b leaves infiltrated with pEAQ-HT-GFP vector. qRT-PCR analysis of c transient expression analysis of GgSQE1 gene in untransformed Nicotiana benthamiana plant (UT) and transformed plant (GgSQE1); and d transient expression of GgSQE1 gene on different days (0, 3, 5, 8, 10 and 12) post-infiltration in the confirmed Nicotiana benthamiana transformant
To further investigate the functional aspect of GgSQE1, stable tobacco transgenic lines overexpressing GgSQE1 gene were developed following Agrobacterium-mediated transformation (Fig. 8). The putative transformants were screened on kanamycin (100 μg/ml) containing MS media along with control. The antibiotic-resistant transformants were reconfirmed by PCR using gene-specific primers and real-time primers (Supplementary Figure 7).
Heterologous stable transformation of GgSQE1 gene from Glycyrrhiza glabra into Nicotiana tabacum mediated by Agrobacterium tumefaciens (GV3101) harbouring pBI121 plasmid (pBI121-GgSQE1). The first row (Fig. 8a–d) represents the pictures of different stages of transformed plant and second row has empty vector–transformed plant (Fig. 8e–h). a Callus induction from Agrobacterium tumefaciens (GV3101) containing pBI121-GgSQE1–transformed leaf disc; b putative transgenic shoots cultured on MS medium containing kanamycin (100 μg/ml ) and cefotaxime (250 μg/ml ); c acclimatised transformed tobacco plantlet; d flowering stage of regenerated transformed plant; e empty vector–transformed plants; f empty vector control plants cultured on MS medium containing kanamycin (100 μg/ml ) and cefotaxime (250 μg/ml ); g acclimatised empty vector–transformed tobacco plantlet; and h empty vector–transformed control plants
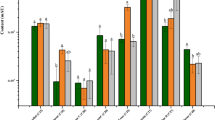
The impact of overexpression of the GgSQE1 gene on the downstream sterol pathway in N. tabacum was analysed through GC-MS. The plants raised from the seeds of the T1 generation were screened for the sterol contents. The seed-raised plants showed wide variation in the analysed constituents. However, a significant modulation in the content of the downstream sterols was observed in the transformed plants. The relative percentage of the selected six lines is represented in Fig. 9. The positive effect of GgSQE1 overexpression was evident by the presence of α-tocopherol (vitamin E) while it was absent in the control and empty vector transformed plants. The higher levels of few sterols in the control plant may be expected as they are the segregating population of the seed-derived plants. The EIC spectra of the sample and control along with individual identified peaks of the compounds are given in Supplementary Figure 8.
Histogram representing the quantitative data of the phytol, cholesterol, campesterol, stigmasterol, D-α-tocopherol, stigmast-5-en-3-ol, cycloartenol, ergosterol and sitosterols. In wild-type plant, empty vector–transformed plant and GgSQE1-transformed plants. The contents of phytosterols in transformed plants were compared to those in the empty vector–transformed plants. The data was compared and analysed using one-way ANOVA in GraphPad Prism 8 software. Histogram represents values which are means ± SE of three independent biological replicates, each with three technical replicates. SE is represented by bars. Differences were scored as statistical significance at **P < 0.01 and *P < 0.05 levels. Asterisks show the difference between transformed and empty vector–transformed plants
Discussion
Squalene epoxidase (SQE) catalyses the stereo-specific conversion of squalene to 2,3-oxidosqualene, which, in turn, serves as a precursor of several structurally and functionally different sterols and triterpenoids, including glycyrrhizin. In recent years, the SQE-encoding genes from many organisms have been isolated and characterised, including those of pathogenic fungi, plants, mice, rats and humans (Ruckenstuhl et al. 2005). In plants, the presence of multiple SQE genes and their phylogenetic diversity suggest their diverse and new evolutionary roles. Experiments conducted in A. thaliana revealed multiple copies of non-redundant SQE genes (Rasbery 2007). Studies have demonstrated varied and differential expression of squalene epoxidase gene paralogs from several plant species, indicating their regulatory/diverse function (Laranjeira et al. 2007). The study demonstrated constitutive expression of two SQEs (SQE1 and SQE3) in A. thaliana, while SQE2 and SQE4 could be induced under certain conditions. The squalene epoxidase 1 (AtSQE1) predominantly catalysed enzymatic step of the sterol biosynthetic pathway, which could be functionally complemented by AtSQE3 homolog. However, AtSQE3 predominantly functions in embryo development (Laranjeira et al. 2007; Suzuki et al. 2002). Despite the crucial role of SQE, the genes encoding SQE have not been characterised in Glycyrrhiza species in detail so for. In the present study, therefore, we have carried out identification, cloning and functional characterisation of SQE genes from G. glabra. The study demonstrated differential expression of squalene epoxidase gene and its homologs in different tissues (Fig. 4a–e) and under the influence of ABA and GA3 hormones (Fig. 6a). The data showed constitutive expression of GgSQE1 gene under normal conditions in both the roots and shoot tissues of the plant, while the four homologs (GgSQE2–GgSQE5) were highly repressed in all the tissues. The roughly conserved expression pattern of GgSQE25 genes can be attributed to their conserved evolutionary relatedness as all these genes were found to be clustered together within the phylogenetic tree. GgSQE1, on the other hand, was subgrouped into Fabaceae A, clustering with Astragalus membranaceus which has a role in astragaloside biosynthesis (Kim et al. 2014). Further study of the homologs will help in understanding their role in different physiological processes occurring in G. glabra plant.
Furthermore, the dynamics of GgSQE1 transcripts in the month-wise in vitro-regenerated tissues of G. glabra demonstrated its differential expression (Supplementary Figure 6). The root tissues of 10- and 12-month-old in vitro-grown plant upregulated higher expression, while in shoot tissues, its expression was found to be largely unchanged. The higher expression of GgSQE1 in roots, with advancing age, can be related to higher secondary metabolites accumulation, including sterols, flavonoids and terpenoids in mature plant (Hayashi 2007; Hayashi et al. 2003). Additionally, to understand the transcriptional regulation of GgSQE1 gene, its upstream region was cloned, sequenced and functionally characterised using GUS staining and quantified by MUG assay. Transcriptionally fused GgSQE1 promoter with GUS reporter gene was transiently expressed in N. benthamiana leaves and subjected to ABA and GA3 treatments, based on the presence of various cis-regulatory elements present in the upstream region of the gene. Hormone-treated leaves showed higher GUS activity compared to the untreated N. benthamiana leaves which motivated us to study the expression of GgSQE1 gene following ABA and GA3 treatments in G. glabra plant. Quantitative expression analysis of GgSQE1 gene in the aerial tissues of the treated plants showed enhanced expression under both the treatments; however, the effect was found to be more pronounced under ABA (12.4-folds in 10 hours) than the GA3 (2.5-folds in 12 hours) treatments. The results were completely in line with glycyrrhizin accumulation pattern in the shoot tissues subjected to hormone treatments (Fig. 6b), indicating a positive correlation between GgSQE1 expression and glycyrrhizin accumulation. Earlier studies on G. uralensis had also demonstrated detection of enhanced glycyrrhizic acid, triterpene saponins and flavonoid compounds (Qiao et al. 2017) under the influence of GA3 and ABA (Li et al. 2016).
For the functional characterisation, in planta heterologous expression of GgSQE1 gene was carried out transiently in N. benthamiana and stably in N. tabacum plants. The GgSQE1 expression and recombinant protein were confirmed by RT-PCR (Fig. 7) and western blot (Supplementary Figure 3b) analyses in N. benthamiana, while stable GgSQE1 overexpressing tobacco transgenic lines were analysed to see its impact on downstream sterol pathway in N. tabacum. GC-MS analysis of the T1 generation of the selected lines showed accumulation of α-tocopherol in the selected transformed lines (Fig. 9). The enhanced accumulation of phytosterols, especially the presence of α-tocopherol in N. tabacum, suggested the involvement of GgSQE1 in sterol biosynthesis.
Several earlier studies on SQE enzyme have demonstrated its role in oxidosqualene production in the roots and reproductive tissues of Arabidopsis (Rasbery 2007), saponin biosynthesis (Li et al. 2009) and combinatorial biosynthesis. Also the studies pertaining to the effect of inhibiting SQE gene have highlighted their influence on plant secondary metabolites (Cattel et al. 1992; Wentzinger et al. 2002). In the present study, the involvement of GgSQE1 was demonstrated in the downstream sterol biosynthesis in Nicotiana species and terpenoid biosynthesis in G. glabra plant. Our conclusion was based on (1) enhanced upregulated expression of only GgSQE1 in the roots (site of glycyrrhizin biosynthesis) in both the field and in vitro plants studied; while all the other homologs (GgSQE25) were highly repressed in the tissues investigated (Fig. 4); (2) ABA-induced enhanced promoter activity (Fig. 5) corresponding to upregulated GgSQE1 gene expression (Fig. 6a) corroborating with the enhanced accumulation of glycyrrhizin in the ABA-treated G. glabra samples (Fig. 6b); and (3) enhanced accumulation of phytosterols in the stable transformants of N. tabacum plants as compared to the empty vector transformed plant (Fig. 9). The present work on the identification of full-length five SQE genes from Glycyrrhiza species and cloning and characterisation of GgSQE1 gene and promoter will form the basis for the future studies on regulation and metabolic engineering of the downstream pathways, as squalene epoxidase is the key enzyme involved in several pathways. The identification of the GgSQE homologs will open up avenues in understanding their role in plant growth and development.
Conclusion
Cloning characterisation and heterologous expression of squalene epoxidase genes from Glycyrrhiza species are significant as 2,3-oxidosqualene is the upstream precursor metabolite for several industrial and pharmaceutically important molecules. These products have immense chemical diversity ranging from triterpenoids (glycyrrhizin), phytosterols (brassinosteroids), lupane (betulinic acid) to vitamins. Identification of five SQE genes from Glycyrrhiza species will throw light on their role in the regulation of various pathways in the life cycle of the plant. The enhanced activity under abscisic acid and GA3 of the cloned upstream region of GgSQE1 will be useful in future studies on pathway manipulations. Further, GgSQE1 was seen to be expressed more in root with the age of the plant, suggesting higher secondary metabolite production in root with the advancing age of G. glabra plant. The study will help in understanding the regulatory role of SQE gene in the secondary metabolite pathways, including glycyrrhizin biosynthesis and channelisation of the precursor for enhanced target compound.
References
Almeida A, Dong L, Khakimov B, Bassard J-E, Moses T, Lota F, Goossens A, Appendino G, Bak S (2018) A single oxidosqualene cyclase produces the seco-triterpenoid α-onocerin. Plant Physiol 176:1469–1484
Asl MN, Hosseinzadeh H (2008) Review of pharmacological effects of Glycyrrhiza sp. and its bioactive compounds. Phytother Res 22:709–724
Berezin C, Glaser F, Rosenberg J, Paz I, Pupko T, Fariselli P, Casadio R, Ben-Tal N (2004) ConSeq: the identification of functionally and structurally important residues in protein sequences. Bioinformatics 20:1322–1324
Brueckner K, Tissier A (2013) High-level diterpene production by transient expression in Nicotiana benthamiana. Plant Methods 9:46/41–46/10 10 pp
Cattel L, Ceruti M, Balliano G, Viola F (1992) 2,3-Oxidosqualene cyclase and squalene epoxidase as target enzymes for the development of new sterol biosynthesis inhibitors. ACS Symp Ser 497:174–191
Cinatl J, Morgenstern B, Bauer G, Chandra P, Rabenau H, Doerr HW (2003) Glycyrrhizin, an active component of liquorice roots, and replication of SARS-associated coronavirus. Lancet 361:2045–2046
de Souza A (2015) Expression and Partial Purification of His-tagged Proteins in a Plant System. Bio-protocol 5:e1572
Deng S (2001) A chinese medicinal preparation for the treatment of cancer comprising herba chelidonii, radix angelicae sinensis, radix ginseng, radix euphorbiae fischerianae, herba ecliptae, rhizoma curculiginis, pericarpium papaveris, folium artemisiae argyi, radix glycyrrhizae, herba euphorbiae helioscopiae, spica prunellae. In:
Dubey V, Bhalla R, Luthra R (2003) An overview of the non-mevalonate pathway for terpenoid biosynthesis in plants. J Biosci 28:637–646
Fiore C, Eisenhut M, Krausse R, Ragazzi E, Pellati D, Armanini D, Bielenberg J (2008) Antiviral effects of Glycyrrhiza species. Phytother Res 22:141–148
Gao K, Xu J-s, Sun J, Xu Y-h, J-h W, Sui C (2016) Molecular cloning and expression of squalene epoxidase from medicinal plant, Bupleurum chinense. Chin Herb Med 8:67–74
Gupta S, Pandotra P, Gupta AP, Verma MK, Ahuja A, Vishwakarma RA (2013) Direct rhizogenesis, in vitro stolon proliferation and high-throughput regeneration of plantlets in Glycyrrhiza glabra. Acta Physiol Plant 35:2699–2705
Hayashi H (2007) Field surveys and comparative analysis of Glycyrrhiza plants. Foods Food Ingredients J Jpn 212:357–364
Hayashi H, Hattori S, Inoue K, Sarsenbaev K, Ito M, Honda G (2003) Field survey of Glycyrrhiza plants in Central Asia (1). Characterization of G. uralensis, G. glabra and the putative intermediate collected in Kazakhstan. Biol Pharm Bull 26:867–871
He J, Chen K, Hu Z-m, Li K, Song W, Yu L-y, Leung C-H, Ma D-L, Qiao X, Ye M (2018) UGT73F17, a new glycosyltransferase from Glycyrrhiza uralensis, catalyzes the regiospecific glycosylation of pentacyclic triterpenoids. Chem Commun 54:8594–8597
Horsch RB, Fry J, Hoffmann N, Neidermeyer J, Rogers SG, Fraley RT (1989) Leaf disc transformation. Plant molecular biology manual. Springer, In, pp 63–71
Janssen B-J, Gardner RC (1990) Localized transient expression of GUS in leaf discs following cocultivation with Agrobacterium. Plant Mol Biol 14:61–72
Khan R, Khan AQ, Lateef A, Rehman MU, Tahir M, Ali F, Hamiza OO, Sultana S (2013) Glycyrrhizic acid suppresses the development of precancerous lesions via regulating the hyperproliferation, inflammation, angiogenesis and apoptosis in the colon of Wistar rats. PLoS One 8:e56020
Khan S, Pandotra P, Manzoor MM, Kushwaha M, Sharma R, Jain S, Ahuja A, Amancha V, Bhushan S, Guru SK, Gupta AP, Vishwakarma R, Gupta S (2016) Terpenoid and flavonoid spectrum of in vitro cultures of Glycyrrhiza glabra revealed high chemical heterogeneity: platform to understand biosynthesis. Plant Cell. Tissue Organ Cult (PCTOC) 124:507–516
Kim YB, Thwe AA, Li X, Tuan PA, Lee S, Lee JW, Arasu MV, Al-Dhabi NA, Park SU (2014) Accumulation of Astragalosides and Related Gene Expression in Different Organs of Astragalus Membranaceus Bge. var Mongholicus (Bge.). Molecules 19:10922–10935
Koichiro Tamura GS, Peterson D, Filipski A, Kumar S (2013) MEGA6: Molecular Evolutionary Genetics Analysis version 6.0. Mol Biol Evol 30:2725–2729
Laranjeira S, Amorim-Silva V, Esteban A, Arro M, Ferrer A, Tavares Rui M, Botella Miguel A, Rosado A, Azevedo H (2007) Arabidopsis Squalene Epoxidase 3 (SQE3) Complements SQE1 and Is Important for Embryo Development and Bulk Squalene Epoxidase Activity. Mol Plant 8:1090–1102
Laranjeira S, Amorim-Silva V, Esteban A, Arro M, Ferrer A, Tavares Rui M, Botella Miguel A, Rosado A, Azevedo H (2015) Arabidopsis Squalene Epoxidase 3 (SQE3) Complements SQE1 and Is Important for Embryo Development and Bulk Squalene Epoxidase Activity. Mol Plant 8:1090–1102
Li Z, Liang-Qiong MA, Feng XU, Yan X-J, Zhang W-W, Lipinski M, Scholz M, Pieper K, Fischer R, Pruefer D, Mueller KJ (2009) A squalene epoxidase from Nigella sativa participates in saponin biosynthesis and mediates terbinafine resistance in yeast. Cent Eur J Biol 4:163–169
Li Y-P, Yu C-x, Qiao J, Zang Y-M, Xiang Y, Ren G-X, Wang L, Zhang X-Y, Liu C-S (2016) Effect of exogenous phytohormones treatment on glycyrrhizic acid accumulation and preliminary exploration of the chemical control network based on glycyrrhizic acid in root of Glycyrrhiza uralensis. Rev Bras 26:490–496
Li Z, Liang-Qiong MA, Feng XU, Xiao-Juan YAN, Zhang W-W (2018) Cloning and Expression Analysis of a Squalene Epoxidase Gene from Ginkgo biloba. Not Bot Horti Agrobo 46:39–44
Menegazzi M, Di Paola R, Mazzon E, Genovese T, Crisafulli C, Dal Bosco M, Zou Z, Suzuki H, Cuzzocrea S (2008) Glycyrrhizin attenuates the development of carrageenan-induced lung injury in mice. Pharmacol Res 58:22–31
Mochida K, Sakurai T, Seki H, Yoshida T, Takahagi K, Sawai S, Uchiyama H, Muranaka T, Saito K (2017) Draft genome assembly and annotation of Glycyrrhiza uralensis, a medicinal legume. Plant J 89:181–194
Phillips D, Rasbery J, Bartel B, Matsuda S (2006) Biosynthetic diversity in plant triterpene cyclization. Curr Opin Plant Biol 9:305–314
Qiao J, Luo Z, Li Y, Ren G, Liu C, Ma X (2017) Effect of abscisic acid on accumulation of five active components in root of Glycyrrhiza uralensis. Molecules 22:1982
Ramilowski JA, Sawai S, Seki H, Mochida K, Yoshida T, Sakurai T, Muranaka T, Saito K, Daub CO (2013) Glycyrrhiza uralensis transcriptome landscape and study of phytochemicals. Plant Cell Physiol 54:697–710
Rasbery JM (2007) Genetic and biochemical characterization of the squalene epoxidase gene family in Arabidopsis thaliana. In, p 164 pp.
Rasbery J, Shan H, LeClair RJ, Norman M, Matsuda SPT, Bartel B (2007) Arabidopsis thaliana Squalene Epoxidase 1 is Essential for Root and Seed Development. JBC 282:17002–17013
Ruckenstuhl C, Eidenberger A, Lang S, Turnowsky F (2005) Single amino acid exchanges in FAD-binding domains of squalene epoxidase of Saccharomyces cerevisiae lead to either loss of functionality or terbinafine sensitivity. Biochem Soc Trans 33:1197–1201
Ruckenstuhl C, Lang S, Poschenel A, Eidenberger A, Baral PK, Kohut P, Hapala I, Gruber K, Turnowsky F (2007) Characterization of Squalene Epoxidase of Saccharomyces cerevisiae by Applying Terbinafine-Sensitive Variants. Antimicrob Agents Chemother 51:275–284
Sainsbury F, Thuenemann EC, Lomonossoff GP (2009) pEAQ: versatile expression vectors for easy and quick transient expression of heterologous proteins in plants. Plant Biotechnol J 7:682–693
Schafer UA, Reed DW, Hunter DG, Yao K, Weninger AM, Tsang EW, Reaney MJ, MacKenzie SL, Covello PS (1999) An example of intron junctional sliding in the gene families encoding squalene monooxygenase homologues in Arabidopsis thaliana and Brassica napus. Plant Mol Biol 39:721–728
Schmittgen TD, Livak KJ (2008) Analyzing real-time PCR data by the comparative CT method. Nat Protoc 3:1101–1108
Schreuder HA, Prick PAJ, Wierenga RK, Vriend G, Wilson KS, Hol WGJ, Drenth J (1989) Crystal structure of the p-hydroxybenzoate hydroxylasesubstrate complex refined at 1.9 Ã resolution: Analysis of the enzyme-substrate and enzyme-product complexes. J Mol Biol 208:679–696
Seki H, Ohyama K, Sawai S, Mizutani M, Ohnishi T, Sudo H, Akashi T, Aoki T, Saito K, Muranaka T (2008) Licorice beta-amyrin 11-oxidase, a cytochrome P450 with a key role in the biosynthesis of the triterpene sweetener glycyrrhizin. Proc Natl Acad Sci U S A 105:14204–14209
Seki H, Ohyama K, Muranaka T (2009) Discovery of glycyrrhizin-biosynthetic gene in licorice, a medicinal plant. Success in production of biosynthetic intermediates by recombinant yeast. Kagaku to Seibutsu 47:84–86
Seki H, Sawai S, Ohyama K, Mizutani M, Ohnishi T, Sudo H, Fukushima EO, Akashi T, Aoki T, Saito K, Muranaka T (2011) Triterpene functional genomics in licorice for identification of CYP72A154 involved in the biosynthesis of glycyrrhizin. Plant Cell 23:4112–4123
Seki H, Tamura K, Muranaka T (2015) P450s and UGTs: key players in the structural diversity of triterpenoid saponins. Plant Cell Physiol 56:1463–1471
Shibata S (2000) A drug over the millennia: pharmacognosy, chemistry, and pharmacology of licorice. Yakugaku Zasshi 120:849–862
Shibata S, Kondo K, Terabayashi S (2003) Identification of the licorice root stored in shosoin based on the sequences of internal transcribed spacer (ITS) on nrDNA and the chemotaxonomic consideration. Proc Jpn Acad, Ser B 79:176–180
Suzuki H, Achnine L, Xu R, Matsuda SPT, Dixon RA (2002) A genomics approach to the early stages of triterpene saponin biosynthesis in Medicago trunculata. Plant J 32:1033–1048
Wang J, Zhang J, Gao W, Wang Q, Yin S, Liu H, Man S (2013) Identification of triterpenoids and flavonoids, step-wise aeration treatment as well as antioxidant capacity of Glycyrrhiza uralensis Fisch. cell. Ind Crop Prod 49:675–681
Weigel D, Glazebrook J (2006) Transformation of Agrobacterium using the freeze-thaw method. CSH Protoc 2006
Wentzinger L, Bach T, Hartmann M (2002) Inhibition of Squalene Synthase and Squalene Epoxidase in Tobacco Cells Triggers an Up-Regulation of 3-Hydroxy-3-Methylglutaryl Coenzyme A Reductase. Plant Physiol 130:334–346
Xu G, Cai W, Gao W, Liu C (2016) A novel glucuronosyltransferase has an unprecedented ability to catalyse continuous two-step glucuronosylation of glycyrrhetinic acid to yield glycyrrhizin. New Phytol 212:123–135
Acknowledgements
The authors acknowledge the grant (SERB/SB/SO/PS/90/2013) from the Science and Engineering Research Board, India, for funding the study. Malik Muzafar Manzoor and Pankaj Pandotra acknowledge Senior Research fellowship from Council for Scientific and Industrial Research and Pooja Goyal acknowledges Junior Research fellowship from University Grant Commission, India, and Guru Nanak Dev University, Amritsar for PhD registration. We thank Prof. George P Lomonossoff (Department of Biological Chemistry, John Innes Centre, Norwich, UK) for the kind gift of pEAQ-HT vector.
Funding
MM and PG acknowledge CSIR and UGC for the respective fellowships. This work was supported by the Science and Engineering Research Board, Department of Science and Technology, Government of India (SERB/SB/SO/PS/90/2013). University of Jammu and Guru Nanak Dev University, Amritsar, are also acknowledged.
Author information
Authors and Affiliations
Contributions
Malik Muzafar Manzoor: characterisation and transient expression and RT-PCR studies; Pooja Goyal: in silico analysis and stable heterologous expression, hormone treatment and RT-PCR studies; Pankaj Pandotra: gene cloning; Mohd Saleem Dar and Mohd Jamal Dar: protein purification; Ajai P Gupta: chemical analysis; Ashok Ahuja: tissue culture system; Suphla Gupta and Prashant Misra: conceptualisation, supervision and manuscript preparation; Ram Vishwakarma and Manoj.K.Dhar: providing the necessary facilities.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Handling Editor: Handling Editor: Peter Nick
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Supplementary Table 1
List of primer sequences used in the present study (DOCX 16 kb)
Supplementary Table 2
List of plant species aligned for designing degenerate primers of squalene epoxidase from Glycyrrhiza glabra (DOCX 15 kb)
Supplementary Fig. 1
A schematic representation of glycyrrhizin biosynthetic pathway in Glycyrrhiza glabra. One-step catalytic reactions are indicated with solid arrows, and multistep catalytic reactions are indicated with dashed arrows. MVA pathway, mevalonate pathway; MEP pathway, plastid-localised 2-C-methyl-derythritol 4-phosphate pathway; IPP, isopentenyl diphosphate; DMAPP, dimethyl allyl diphosphate; FPP, farnesyl pyrophosphate; IDI, isopentenyl diphosphate isomerase; FPS, farnesyl pyrophosphate synthase; SQS, squalene synthase; SQE, squalene epoxidase; bAS, β-amyrin synthase; CYP88D6, β-amyrin-11-oxidase; CYP72A154,11-oxo-beta-amyrin-30-oxidase ; UGAT, UDP-glycosyltransferase (PNG 2464 kb)
Supplementary Fig. 2
The schematic representation of expression cassette used in the present study. Expression vector for heterologous stable GgSQE1 gene expression (a); heterologous transient GgSQE1 gene expression (b); and promoter activity (c) (PNG 922 kb)
Supplementary Fig. 3
SDS protein gel of cell lysate and purified protein. a (M) protein marker; (L) lysate; (P) pellet; (F) flow-through; (W1-W3) washing; (E1-E3) Elutions (Imidazole concentrations ranging from 100 to 200 mM); b western blot of the purified protein heterologously expressed in Nicotiana benthamiana (PNG 8351 kb)
Supplementary Fig. 4
Multiple sequence alignment of p-hydroxybenzoate hydroxylase (PHBH) of Pseudomonas fluorescens with squalene epoxidase 1 gene identified from Glycyrrhiza glabra (PNG 335 kb)
Supplementary Fig. 5
Multiple sequence alignment of the squalene epoxidase protein from various plant species including the five homologs identified from Glycyrrhiza glabra in the present study using MUSCLE (http://www.ebi.ac.uk/Tools/msa/muscle/). The protein sequence IDs used in the analysis were MG763680 (Glycyrrhiza glabra SQE1); MN593324 (Glycyrrhiza glabra SQE2); MN593325 (Glycyrrhiza glabra SQE3); MN593326 (Glycyrrhiza glabra SQE4); MN593328 (Glycyrrhiza glabra SQE5); ABC94943.1 (Medicago sativa) APD76301.1 (Medicago sativa); KYP54510.1 (Cajanus cajan); KHG06672.1 (Gossypium arboreum); KEH39981.1 (Medicago truncatula); KEH39980.1 (Medicago truncatula); ACQ90301.1 (Gynostemma pentaphyllum); AFZ93645.1 (Euphorbia lathyris); KYP34787.1 (Cajanus cajan); KEH39980.1 (Medicago truncatula); ALD84329.1 (Morus alba); AFI33134.2 (Eriobotrya japonica); ARA73618.1 (Lepidium apetalum); AT1G58440.1 (Arabidopsis thaliana ATSQE1); AT2G22830 (Arabidopsis thaliana ATSQE2); AT4G37760.1 (Arabidopsis thaliana ATSQE3); AT5G24140.1 (Arabidopsis thaliana ATSQE4); AT5G24150.1 (Arabidopsis thaliana ATSQE5); AT5G24160.1 (Arabidopsis thaliana ATSQE6); AB923908.1 (Botryococcus braunii SQE-I); AB986538.1 (Botryococcus braunii SQE-2); ADC32655.1 (Aralia elata); GU574803.1 (Aralia elata); GU574803.1 (Withania somnifera); AGK62446.1 (Panax quinquefolius); AJV26446.1 (Panax ginseng); ACJ24907.2 (Panax ginseng); ABE60738.1 (Panax notoginseng); BAD15330.1 (Panax quinquefolius); AHY94896.1 (Astragalus membranaceus); AUD09558.1 (Ononis spinosa); KHN36995.1 (Glycine soja); KHN17192.1 (Glycine soja); ACJ05633.1 (Nigella sativa); EOX97273.1 (Theobroma cacao); OAY63657.1 (Ananas comosus); BAF79915.1 (Euphorbia tirucalli) (PDF 2175 kb)
Supplementary Fig. 6
Represented photograph of the a field plant; b in vitro-regenerated Glycyrrhiza glabra plant and c qRT-PCR expression analysis of GgSQE1 gene studied in different months (0, 2, 4, 8, 10 and 12) of the regenerated Glycyrrhiza glabra plant. The data were compared and analysed using two-way ANOVA in GraphPad Prism 8 software. Values are expressed as means ± SD representing three independent biological samples, each with three technical replicates. Differences were scored as statistical significance at *** P < 0.0001, **P < 0.001 and *P < 0.05. Asterisks indicate difference in expression levels of SQE1 in root and shoot tissues of 2-month-old plant, with the respective tissues of different months (PNG 1301 kb)
Supplementary Fig.7
a Confirmation PCR for the stable integration of GgSQE1 gene in Nicotiana tabacum lines using full-length gene-specific primers. (M) 1Kb DNA ladder; (L1–L3) transformed lines showing amplified (1.5 kb) integrated GgSQE1 gene; (L4) empty vector–transformed line; (UT) untransformed line. The confirmed transgenic lines were also subjected to b semi-quantitative PCR analysis. (M) 100-bp ladder; (L1–L3) transformed lines; (L4) empty vector–transformed line; (UT) untransformed line and c actin which was used for normalisation (PNG 1115 kb)
Supplementary Fig. 8
Representative GC-MS overlay chromatogram (a) of the empty vector–transformed and GgSQE1-transformed plants and (b) peak of each identified compounds (PNG 7226 kb)
Rights and permissions
About this article
Cite this article
Manzoor, M.M., Goyal, P., Pandotra, P. et al. Transcriptome-wide identification of squalene epoxidase genes from Glycyrrhiza glabra L.: expression analysis and heterologous expression of GgSQE1 suggest important role in terpenoid biosynthesis. Protoplasma 258, 991–1007 (2021). https://doi.org/10.1007/s00709-021-01616-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00709-021-01616-2