Abstract
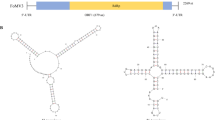
A novel mitovirus, tentatively designated as “Fusarium oxysporum mitovirus 2” (FoMV2), was isolated from the pathogenic Fusarium oxysporum f. sp. ginseng strain 0414 infecting Panax ginseng. The complete genome of FoMV2 is 2388 nt in length with a GC content of 30.57%. It contains a large open reading frame (ORF) encoding a putative RNA-dependent RNA polymerase (RdRp) of 713 amino acids with a molecular weight of 83.05 kDa. The sequence identity between FoMV2 and Botrytis cinerea mitovirus 8 and Fusarium verticillioides mitovirus 1 was 87.94% and 77.85%, respectively. Phylogenetic analysis showed that FoMV2 belongs to the genus Unuamitovirus in the family Mitoviridae. To the best of our knowledge, this is the first report of an unuamitovirus isolated from F. oxysporum f. sp. ginseng causing ginseng root rot.



Similar content being viewed by others
Data availability
The complete genome sequence of FoMV2 has been deposited in the NCBI database with the accession number OR714797.
References
Paez-Espino D, Eloe-Fadrosh EA, Pavlopoulos GA, Thomas AD, Huntemann M, Mikhailova N, Rubin E, Ivanova NN, Kyrpides NC (2016) Uncovering Earth’s virome. Nature 536(7617):425–430. https://doi.org/10.1038/nature19094
Xie JT, Jiang DH (2014) New insights into mycoviruses and exploration for the biological control of crop fungal diseases. Annu Rev Phytopathol 52(1):45–68. https://doi.org/10.1146/annurev-phyto-102313-050222
Ghabrial SA (1998) Origin, adaptation and evolutionary pathways of fungal viruses. Virus Genes 16(1):119–131. https://doi.org/10.1023/a:1007966229595
Lian ZQ, Das S, Luo JX, Andika IB, Sun L (2021) Complete genome sequence of a novel ourmia-like mycovirus infecting the phytopathogenic fungus Botryosphaeria dothidea. Arch Virol 166(12):3461–3465. https://doi.org/10.1007/S00705-021-05221-9
Hollings M (1962) Viruses associated with a die-back disease of cultivated mushroom. Nature 196(4858):962–965. https://doi.org/10.1038/196962a0
Ghabrial SA, Castón JR, Jiang DH, Nibert ML, Suzuki N (2015) 50-plus years of fungal viruses. Virology 479–480:356–368. https://doi.org/10.1016/j.virol.2015.02.034
Mu F, Xie JT, Cheng SF, You MP, Barbetti MJ, Jia JC, Wang QQ, Cheng JS, Fu YP, Chen T, Jiang DH (2017) Virome characterization of a collection of S. sclerotiorum from Australia. Front Microbiol 8:2540. https://doi.org/10.3389/fmicb.2017.02540
Yu X, Li B, Fu YP, Jiang DH, Ghabrial SA, Li GQ, Peng YL, Xie JT, Cheng JS, Huang JB, Yi XH (2010) A geminivirus-related DNA mycovirus that confers hypovirulence to a plant pathogenic fungus. Proc Natl Acad Sci USA 107(18):8387–8392. https://doi.org/10.1073/pnas.0913535107
Wang YF, Zhao H, Cao JY, Yin XM, Guo YS, Guo LH, Wu HY, Zhang M (2022) Characterization of a novel mycovirus from the phytopathogenic fungus Botryosphaeria dothidea. Viruses 14(2):331–340. https://doi.org/10.3390/v14020331
Nuss DL (1992) Biological control of chestnut blight: an example of virus-mediated attenuation of fungal pathogenesis. Microbiol Rev 56(4):561–576. https://doi.org/10.1128/MR.56.4.561-576.1992
Li SW, Li YT, Hu CH, Han CG, Zhou T, Zhao C, Wu XH (2020) Full genome sequence of a new mitovirus from the phytopathogenic fungus Rhizoctonia solani. Arch Virol 165(7):1719–1723. https://doi.org/10.1007/s00705-020-04664-w
Zerbini FM, Siddell SG, Lefkowitz EJ, Mushegian AR, Adriaenssens EM, Alfenas-Zerbini P, Dempsey DM, Dutilh BE, García ML, Hendrickson RC, Junglen S, Krupovic M, Kuhn JH, Lambert AJ, Łobocka M, Oksanen HM, Robertson DL, Rubino L, Sabanadzovic S, Simmonds P, Smith DB, Suzuki N, Van Doorslaer K, Vandamme AM, Varsani A (2023) Changes to virus taxonomy and the ICTV statutes ratified by the International Committee on Taxonomy of Viruses. Arch Virol 168(7):175. https://doi.org/10.1007/S00705-023-05797-4
Xu ZY, Wu SS, Liu LJ, Cheng JS, Fu YP, Jiang DH, Xie JT (2014) A mitovirus related to plant mitochondrial gene confers hypovirulence on the phytopathogenic fungus Sclerotinia sclerotiorum. Virus Res 197:127–136. https://doi.org/10.1016/j.virusres.2014.12.023
Wu MD, Zhang L, Li GQ, Jiang DH, Ghabrial SA (2010) Genome characterization of a debilitation-associated mitovirus infecting the phytopathogenic fungus Botrytis cinerea. Virology 406:117–126. https://doi.org/10.1016/j.virol.2010.07.010
Li PF, Bhattacharjee P, Wang SC, Zhang LH, Ahmed I, Guo LH (2019) Mycoviruses in Fusarium species: an update. Front Cell Infect Microbiol 9:257–267. https://doi.org/10.3389/fcimb.2019.00257
Cho WK, Yu J, Lee KM, Son M, Min K, Lee YW, Kim KH (2012) Genome-wide expression profiling shows transcriptional reprogramming in Fusarium graminearum by Fusarium graminearum virus 1-DK21 infection. BMC Genomics 13:173–188. https://doi.org/10.1186/1471-2164-13-173
Yu J, Kim KH (2020) Exploration of the interactions between mycoviruses and Fusarium graminearum. Adv Virus Res 106:123–144. https://doi.org/10.1016/bs.aivir.2020.01.004
Wang SC, Kondo H, Liu L, Guo LH, Qiu DW (2013) A novel virus in the family Hypoviridae from the plant pathogenic fungus Fusarium graminearum. Virus Res 174(1–2):69–77. https://doi.org/10.1016/j.virusres.2013.03.002
Li PF, Zhang HL, Chen XG, Qiu DW, Guo LH (2015) Molecular characterization of a novel hypovirus from the plant pathogenic fungus Fusarium graminearum. Virology 481:151–160. https://doi.org/10.1016/j.virol.2015.02.047
Wang SC, Ruan SJ, Zhang MM, Nie JH, Nzabanita C, Guo LH (2022) Interference of small RNAs in Fusarium graminearum through FgGMTV1 infection. J Fungi 8(12):1237. https://doi.org/10.3390/JOF8121237
Darissa O, Adam G, Schäfer W (2012) A dsRNA mycovirus causes hypovirulence of Fusarium graminearum to wheat and maize. Eur J Plant Pathol 134(1):181–189. https://doi.org/10.1007/s10658-012-9977-5
Lemus-Minor CG, Cañizares MC, García-Pedrajas MD, Pérez-Artés E (2018) Fusarium oxysporum f. sp. dianthi virus 1 accumulation is correlated with changes in virulence and other phenotypic traits of its fungal host. Phytopathology 108(8):957–963. https://doi.org/10.1094/PHYTO-06-17-0200-R
Wang J, Feng S, Lu BH, Yang LN, Wang X, Zhang YJ, Gao J (2022) Fusarium oxysporum f. sp. ginseng, a new forma specialis causing Fusarium root rot of Panax ginseng. Phytopathol Mediterr 61(3):417-429. https://doi.org/10.36253/phyto-13723
Scotto-Lavino E, Du GW, Frohman MA (2006) 5’ end cDNA amplifification using classic RACE. Nat Protoc 1(6):2555–2562. https://doi.org/10.1038/nprot.2006.480
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23(21):2947–2948. https://doi.org/10.1093/bioinformatics/btm404
Nicholas KB (1997) GeneDoc: analysis and visualization of genetic variation, EMBNEW. Embnew News 4:14. https://doi.org/10.11118/actaun201361041061
Liu H, Liu M, Zhu HJ, Zhong J, Liao XL, Zhou Q (2020) Molecular characterization of a novel mitovirus from the plant-pathogenic fungus Botryosphaeria dothidea. Arch Virol 166(2):633–637. https://doi.org/10.1007/s00705-020-04886-y
Jacquat AG, Ulla SB, Debat HJ, Muñoz-Adalia EJ, Theumer MG, Pedrajas MDG, Dambolena JS (2022) An in silico analysis revealed a novel evolutionary lineage of putative mitoviruses. Environ Microbiol 24(12):6463–6475. https://doi.org/10.1111/1462-2920.16202
Acknowledgments
This research work was supported by the Scientific and Technological Development Project of Jilin Province (20210204047YY).
Funding
The funding has been received from Jilin Provincial Scientific and Technological Development Program with Grant no. 20210204047YY.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals.
Additional information
Handling Editor: Nobuhiro Suzuki.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ma, K., Cai, L., Wang, R. et al. Complete genome sequence of a novel mitovirus isolated from the fungus Fusarium oxysporum f. sp. ginseng causing ginseng root rot. Arch Virol 169, 53 (2024). https://doi.org/10.1007/s00705-024-05962-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00705-024-05962-3