Abstract
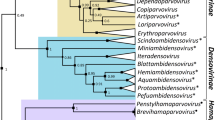
Here, we report a novel mycotombus-like mycovirus, tentatively named "Phoma matteucciicola RNA virus 2" (PmRV2), derived from the phytopathogenic fungus Phoma matteucciicola strain HNQH1. The complete PmRV2 genome is comprised of a positive-sense single-stranded RNA (+ssRNA) of 3,460 nucleotides (nt) with a GC content of 56.71%. Sequence analysis of PmRV2 indicated the presence of two noncontiguous open reading frames (ORFs) encoding a hypothetical protein and an RNA-dependent RNA polymerase (RdRp), respectively. PmRV2 contains a metal-binding ‘GDN’ triplet in motif C of RdRp, while most +ssRNA mycoviruses contained a ‘GDD’ motif in the same region. A BLASTp search showed that the RdRp amino acid sequence of PmRV2 was most closely related to the RdRp of Macrophomina phaseolina umbra-like virus 1 (50.72% identity) and Erysiphe necator umbra-like virus 2 (EnUlV2, 44.84% identity). Phylogenetic analysis indicated that PmRV2 grouped together with EnUlV2 within the recently proposed family "Mycotombusviridae".


Similar content being viewed by others
References
Hollings M (1962) Viruses associated with a die-back disease of cultivated mushroom. Nature 196:962–965
Ghabrial SA, Castón JR, Jiang D, Nibert ML, Suzuki N (2015) 50-plus years of fungal viruses. Virology 479–480:356–368
Kondo H, Hisano S, Chiba S, Maruyama K, Andika IB, Toyoda K, Fujimori F, Suzuki N (2016) Sequence and phylogenetic analyses of novel totivirus-like double-stranded RNAs from field-collected powdery mildew fungi. Virus Res 213:353–364
Marzano SL, Domier LL (2016) Novel mycoviruses discovered from metatranscriptomics survey of soybean phyllosphere phytobiomes. Virus Res 213:332–342
Kondo H, Botella L, Suzuki N (2022) Mycovirus diversity and evolution revealed/inferred from recent studies. Annu Rev Phytopathol 60:14.1-14.30
Zhou J, Wang Y, Liang X, Xie C, Liu W, Miao W, Kang Z, Zheng L (2020) Molecular characterization of a novel ourmia-like virus infecting Phoma matteucciicola. Viruses 12:231
Donaire L, Pagán I, Ayllón M (2016) Characterization of Botrytis cinerea negative-stranded RNA virus 1, a new mycovirus related to plant viruses, and a reconstruction of host pattern evolution in negative-sense ssRNA viruses. Virology 499:212–218
Sato Y, Shamsi W, Jamal A, Bhatti MF, Kondo H, Suzuki N (2020) Hadaka virus 1: a capsidless eleven-segmented positive-sense single-stranded RNA virus from a phytopathogenic fungus, Fusarium oxysporum. MBio 11:e00450-520
Yu X, Li B, Fu Y, Jiang D, Ghabrial SA, Li G, Peng Y, Xie J, Cheng J, Huang J, Yi X (2010) A geminivirus-related DNA mycovirus that confers hypovirulence to a plant pathogenic fungus. Proc Natl Acad Sci USA 107:8387–8392
Zhou J, Hu X, Liang X, Wang Y, Xie C, Zheng L (2021) Complete genome sequence of a novel mycovirus from Phoma matteucciicola. Arch Virol 166:317–320
Zheng L, Shu C, Zhang M, Yang M, Zhou E (2019) Molecular characterization of a novel endornavirus conferring hypovirulence in rice sheath blight fungus Rhizoctonia solani AG-1 IA strain GD-2. Viruses 11:178
Anagnostakis SL (1982) Biological control of chestnut blight. Science 215:466–471
Rigling D, Prospero S (2018) Cryphonectria parasitica, the causal agent of chestnut blight: invasion history, population biology and disease control. Mol Plant Pathol 19:7–20
Zheng F, Xu G, Zhou J, Xie C, Cui H, Miao W, Kang Z, Zheng L (2019) Complete genomic sequence and organization of a novel mycovirus from Phoma matteucciicola strain LG915. Arch Virol 164:2209–2213
Zheng F, Ma R, Xu G, Zheng F, Ding X, Xie C (2018) Leaf blight on Curcuma wenyujin caused by Phoma matteucciicola in China. Plant Dis 102:2042
Morris TJ, Dodds JA (1979) Isolation and analysis of double stranded RNA from virus-infected plant and fungal tissue. Phytopathology 69:854–858
Lin Y, Zhou J, Zhou X, Shuai S, Zhou R, An H, Fang S, Zhang S, Deng Q (2020) A novel narnavirus from the plant-pathogenic fungus Magnaporthe oryzae. Arch Virol 165:1235–1240
Zuker M (2003) Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res 31:3406–3415
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549
Vázquez AL, Alonso JM, Parra F (2000) Mutation analysis of the GDD sequence motif of a calicivirus RNAdependent RNA polymerase. J Virol 74:3888–3891
Funding
This study was financially supported by Hainan Province Key R&D Project (No. ZDYF2022XDNY242), the National Natural Science Foundation of China (No. 32260648), the Scientific Research Foundation for Advanced Talents, Hainan University (No. KYQD(ZR)1873), the Leading Talent Project of Main Discipline Leaders in Jiangxi Province (No. 20213BCJL22046), and the Key Projects of Jiangxi Natural Science Foundation (No. 20202ACBL205006).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with animals or human participants performed by any of the authors.
Additional information
Handling Editor: Nobuhiro Suzuki.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhou, S., Chen, D., Fu, Y. et al. Characterization of a novel mycotombus-like virus from the plant-pathogenic fungus Phoma matteucciicola. Arch Virol 168, 103 (2023). https://doi.org/10.1007/s00705-023-05714-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00705-023-05714-9