Abstract
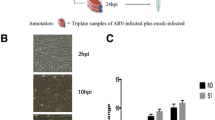
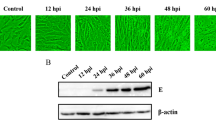
Infection with recombinant avian leukosis virus (ALV) has previously been linked to malignancies and immunosuppression. However, the processes behind the unique pathophysiology of recombinant ALV are poorly understood. In this study, we analyzed gene expression patterns in chicken fibroblast cells (CEFs) infected with the recombinant ALV isolate GX14FF03 and used the RNA-seq technique to perform a complete analysis of the transcribed mRNAs. A total of 907 significant differentially expressed genes (SDEGs) were identified. Among these SDEGs, the most significantly upregulated gene was interleukin 8-like 1 (IL8L1), while the most significantly downregulated gene was fibroblast growth factor 16 (FGF16). The 907 SDGEs were highly enriched (p < 0.05) for 252 Gene Ontology (GO) terms, including 197 BP, 3 CC, and 52 MF. According to KEGG data analysis, SDEGs are implicated in eight significant pathways (p < 0.05). Furthermore, protein-protein interaction (PPI) network analysis revealed that IL8L1 interacts with 17 genes. These findings shed light on the molecular mechanisms involved in recombinant ALV infection by showing the mRNA expression profile in CEFs infected with GX14FF03 virus.




Similar content being viewed by others
Availability of data and materials
The datasets used and/or analyzed in the current study are available from the corresponding author upon reasonable request.
Abbreviations
- ALV:
-
Avian leukosis virus
- CEFs:
-
Chicken fibroblast cells
- SDEGs:
-
Significant differentially expressed genes
- IL8L1:
-
Interleukin 8-like 1
- FGF16:
-
Fibroblast growth factor 16
- GO:
-
Gene Ontology
- DMEM:
-
Dulbecco’s modified Eagle medium
- FBS:
-
Fetal bovine serum
- TCID50 :
-
50% Tissue culture infectious dose
- PPI:
-
Protein-protein interaction
- qPCR:
-
Real-time quantitative PCR
- BP:
-
Biological process
- CC:
-
Cellular component
- MF:
-
Molecular function
References
Deng QM, Li M, He CW, Lu QE, Gao YL, Li QH, Shi MY, Wang PK, Wei P (2021) Genetic diversity of avian leukosis virus subgroup J (ALV-J): toward a unified phylogenetic classification and nomenclature system. Virus Evol 7:veab037
Li XY, Yu Y, Ma MG, Chang FF, Muhammad F, Yu MM, Ren CQ, Bao YL, Zhang Z, Liu AJ, Qing P, Gao L, Qi XL, Li K, Liu CJ, Zhang YP, Cui HY, Wang XM, Gao YL (2021) Molecular characteristic and pathogenicity analysis of a novel multiple recombinant ALV-K strain. Vet Microbiol 260:109184
Wang PK, Lin LL, Shi MY, Li HJ, Gu ZM, Li M, Gao YL, Teng H, Mo ML, Wei TC, Wei P (2020) Vertical transmission of ALV from ALV-J positive parents caused severe immunosuppression and significantly reduced Marek’s disease vaccine efficacy in three-yellow chickens. Vet Microbiol 244:108683
Li HJ, Wang PK, Lin LL, Shi MY, Gu ZM, Huang T, Mo ML, Wei TC, Zhang HM, Wei P (2019) The emergence of the infection of subgroup J avian leucosis virus escalated the tumour incidence in commercial Yellow chickens in Southern China in recent years. Transbound Emerg Dis 66:312–316
Li TF, Xie J, Yao XH, Zhang J, Li CP, Ren D, Li LY, Xie Q, Shao HX, Qin A, Ye JQ (2021) The tyrosine phosphatase SHP-2 dephosphorylated by ALV-J via its Env efficiently promotes ALV-J replication. Virulence 12:1721–1731
Hang BL, Sang JJ, Qin AJ, Qian K, Shao HX, Mei M, Ye JQ (2014) Transcription analysis of the response of chicken bursa of Fabricius to avian leukosis virus subgroup J strain JS09GY3. Virus Res 188:8–14
Zhou JR, Liu JH, Li HM, Zhao Y, Cheng ZQ, Hou YM, Guo HJ (2020) Regulatory effects of chicken TRIM25 on the replication of ALV-A and the MDA5-mediated type I interferon response. Vet Res 51(51):145
Wu XP, Zeng YK, Lu RB, An YJ, Yu SY, Zhao JR, Wu YJ, Wu BC, Wang QX, Huang YF (2018) Transcription analysis of the interaction between chicken thymus and recombinant avian leukosis virus isolate FJ15HT0. Virus Res 244:147–152
Wang PK, Yang YL, Lin LL, Li HJ, Wei P (2017) Complete genome sequencing and characterization revealed a recombinant subgroup B isolate of avian leukosis virus with a subgroup J-like U3 region. Virus Genes 53:927–930
Li QH, Wang PK, Li M, Lin LL, Shi MY, Li HJ, Deng QM, Huang T, Mo ML, Wei TC, Wei P (2020) Recombinant subgroup B avian leukosis virus combined with the subgroup J env gene significantly increases its pathogenicity. Vet Microbiol 250:108862
Denny P, Feuermann M, Hill DP, Lovering RC, Plun-Favreau H, Roncaglia P (2018) Exploring autophagy with gene ontology. Autophagy 14:419–436
Pian C, Zhang GL, Gao LB, Fan XD, Li F (2020) miR+Pathway: the integration and visualization of miRNA and KEGG pathways. Brief Bioinform 21:699–708
Szklarczyk D, Morris JH, Cook H, Kuhn M, Wyder S, Simonovic M, Santos A, Doncheva NT, Roth A, Bork P, Lars JJ, Christian VM (2017) The STRING database in 2017: quality-controlled protein-protein association networks, made broadly accessible. Nucleic Acids Res 45:D362–D368
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13:2498–2504
Bada M, Hunter L (2008) Identification of OBO nonalignments and its implications for OBO enrichment. Bioinformatics 24:1448–1455
Li TF, Xie J, Liang GD, Ren D, Sun S, Lv L, Xie Q, Shao HX, Gao W, Qin AJ, Ye JQ (2019) Co-infection of vvMDV with multiple subgroups of avian leukosis viruses in indigenous chicken flocks in China. BMC Vet Res 15:288
Machacek M, Saunders H, Zhang Z, Tan EP, Li JB, Li TG, Villar MT, Artigues A, Lydic T, Cork G, Slawson C, Fields PE (2019) OElevated -GlcNAcylation enhances pro-inflammatory Th17 function by altering the intracellular lipid microenvironment. J Biol Chem 294:8973–8990
Gudd CLC, Au L, Triantafyllou E, Shum B, Liu T, Nathwani R, Kumar N, Mukherjee S, Dhar A, Woollard KJ, Yon Y, Pinato DJ, Thursz MR, Goldin RD, Gor ME, Larkin J, Khamri W, Antoniades CG, Truajlic S, Possamai LA (2021) Activation and transcriptional profile of monocytes and CD8 T cells are altered in checkpoint inhibitor-related hepatitis. J Hepatol 75:177–189
Hu XM, Chen SH, Jia CX, Xue SL, Dou CF, Dai ZQ, Xu H, Sun Z, Geng TY, Cui HM (2018) Gene expression profile and long non-coding RNA analysis, using RNA-Seq, in chicken embryonic fibroblast cells infected by avian leukosis virus. J Archiv Virol 163:639–647
Qiu LL, Li ZT, Chang GB, Bi YL, Liu XP, Xu L, Zhang Y, Zhao WM, Xu Q, Chen GH (2017) Discovery of novel long non-coding RNAs induced by subgroup J avian leukosis virus infection in chicken. Dev Comp Immunol 76:292–302
de Oliveira S, Reyes-Aldasoro C, Candel S, Renshaw SA, Mulero V, Calado A (2013) Cxcl8 (IL-8) mediates neutrophil recruitment and behavior in the zebrafish inflammatory response. J Immunol (Baltimore, MD 1950) 190:4349–4359
Jin H, Kong ZM, Mehboob A, Jiang B, Xu J, Cai YH, Liu WX, Hong JB, Li YQ (2020) Transcriptional profiles associated with Marek’s disease virus in bursa and spleen lymphocytes reveal contrasting immune responses during early cytolytic infection. Viruses 12:354
Zheng M, Karki R, Williams EP, Yang D, Fitzpatrick E, Vogel P, Jonsson CB, Kanneganti TD (2021) TLR2 senses the SARS-CoV-2 envelope protein to produce inflammatory cytokines. Nat Immunol 22:829–838
Patra MC, Shah M, Choi SD (2020) Toll-like receptor-induced cytokines as immunotherapeutic targets in cancers and autoimmune diseases. Semin Cancer Biol 64:61–82
Liu D, Qiu QQ, Zhang X, Dai MM, Qin JR, Hao JJ, Liao M, Cao WS (2016) Infection of chicken bone marrow mononuclear cells with subgroup J avian leukosis virus inhibits dendritic cell differentiation and alters cytokine expression. Infect Genet Evol 44:130–136
Acknowledgements
The manuscript was kindly reviewed by Dr. Richard Roberts, Aurora, CO 80014, USA.
Funding
This work was supported by the Shandong Provincial Natural Science Foundation [ZR2019BC047], the Guangxi Special Funding on Science and Technology Research [AA17204057], the Guangxi Program for Modern Agricultural Industry Technical System Construction-Chicken Industry [nycytxgxcxtd-19-03], the Major Basic Program of Natural Science Foundation of Shandong Province, China [ZR2019ZD21], the Taishan Scholars Program of Shandong Province, China [ts20190955].
Author information
Authors and Affiliations
Contributions
Conceptualization: WPK. WJ conceived and designed the experiments. LQH collected the samples and data. WPK, LQH, DQM, and LM performed the experiments; WPK analyzed the data and wrote the manuscript. DQM and LM reviewed and edited the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethical approval
No animal experiments were performed in this study.
Additional information
Handling Editor: Tim Skern.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, P., Li, Q., Wangjing et al. Transcription analysis of chicken embryo fibroblast cells infected with the recombinant avian leukosis virus isolate GX14FF03. Arch Virol 167, 2613–2621 (2022). https://doi.org/10.1007/s00705-022-05597-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-022-05597-2