Abstract
Here, we report a novel bat adenovirus strain isolated from apparently healthy bats of the species Rhinolophus cornutus in Japan. The genome of the isolate was 36,506 bp in length and encoded at least 33 proteins. Phylogenetic analysis of the DNA polymerase amino acid sequence, which provides one demarcation criterion for adenoviral species, indicated that the isolate belongs to the species Bat mastadenovirus C in the genus Mastadenovirus. Most of the encoded proteins shared high sequence similarity with those of known bat adenovirus C strains detected in different species of Rhinolophus, whereas the fiber protein and some E3- and E4-related proteins shared moderate similarity, and only the large E3 protein, which contains several host immune-suppression-related motifs, showed considerably lower similarity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Bats are an important reservoir of many zoonotic viruses [1,2,3,4] including adenoviruses [5, 6]. Adenoviruses are non-enveloped icosahedral viruses with a double-stranded DNA genome that infect a wide range of vertebrate hosts from amphibians and fish to mammals. The family Adenoviridae is divided into six genera (Atadenovirus, Aviadenovirus, Ichtadenovirus, Mastadenovirus, Siadenovirus, and Testadenovirus). Mastadenoviruses infect a variety of mammalian hosts, including bats. Bat mastadenoviruses (BtAdVs) have been classified as members of the species Bat mastadenovirus A-J by the International Committee on Taxonomy of Viruses (ICTV) [7]. BtAdVs are divided into three groups depending on host family classification: group 1, comprised of members of species A, B, G, and J, isolated from Vespertilionidae bats and genetically closely related to canine adenoviruses; group 2, comprised of members of species C from Rhinolophidae bats; and group 3, comprised of members of species D, E, F, H, and I from Miniopteridae and Pteropodidae bats [8].
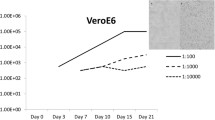
To investigate bat viruses, we collected fecal samples from apparently healthy insectivorous Rhinolophus cornutus bats with permission from the local government in Shizuoka Prefecture of Japan in September 2020. We then inoculated antibiotic-treated fecal suspensions onto African green monkey Vero cells expressing a transmembrane protease, serine 2 (Vero/TMPRSS2), which may support the replication of certain unknown viruses [9]. A clear cytopathic effect was observed in the cells at 7 days post-inoculation. The cell supernatant was passed through a 0.22-μm filter and inoculated onto fresh Vero/TMPRSS2 cells, producing an extensive cytopathic effect (Supplementary Fig. S1). This indicated that the virus had been isolated successfully. To identify the genome type of the isolate, we analyzed its growth in the presence of ribavirin or 5-iodo-2'-deoxyuridine (IUDR), which inhibit the growth of RNA and DNA viruses, respectively. Growth was inhibited with IUDR but not with ribavirin, suggesting that the isolate was a DNA virus. Then, to determine the complete genome sequence of the viral isolate, we grew it in cells for 4 days and purified it from the culture medium by ultracentrifugation with a 20% sucrose cushion. Viral DNA was extracted from the purified virus using a NucleoSpin Tissue Kit (Macherey-Nagel, Duren, Germany). A DNA library was prepared using a TruSeq Nano DNA Low Throughput Library Prep Kit (Illumina, San Diego, CA, USA) and subjected to next-generation sequencing (NovaSeq 6000, Illumina). The sequence dataset was assembled de novo using CLC Genomics Workbench software (v.8, CLC bio, Aarhus, Denmark), and sequence comparisons indicated that the isolate was a BtAdV strain, which was named Rc-kw20. The complete genome of Rc-kw20 is 36,506 nucleotides long with 54% G+C content. The complete nucleotide sequence has been deposited in the DDBJ/GenBank database under the accession number MZ683934.
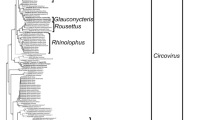
The full genomic nucleotide sequence was analyzed phylogenetically (Fig. 1a). The phylogenetic tree indicated that strain Rc-kw20 was positioned in the same cluster with previous BtAdV-C strains, all of which had been detected in Rhinolophus sinicus bats in China, suggesting that Rc-Kw20 belongs to the species Bat mastadenovirus C in the genus Mastadenovirus. The full genomic nucleotide sequence of Rc-kw20 was 83.8%, 83.9%, and 83.7% identical to BtAdV-C strains, WIV9, WIV10, and WIV11, respectively (GenBank accession numbers KT698853-5).
Genetic analysis of bat mastadenoviruses. Phylogenetic trees based on (a) full-length nucleotide sequences, (b) DNA polymerase amino acid sequences, and (c) hexon amino acid sequences were constructed by the neighbor-joining method with 1,000 bootstrap replicates using MEGA X software. Bootstrap values are indicated as percentages at the nodes. Bars indicate evolutionary distance in units of substitutions per site. All bat mastadenoviruses are underlined, and the novel isolate identified in this study is shown in bold red. (d) Gene map of BatAdV-Rc-kw20 in which genes are shown as grey arrows and ITR sequences are shown as black rectangles
Open reading frames (ORFs) were predicted using SnapGene Viewer software (SnapGene v.5; GSL Biotech, San Diego, CA, USA) (Table 1), and 33 putative ORFs were identified. Comparisons of the DNA polymerase amino acid sequences of Rc-kw20 and other BtAdV-C strains, which provide one demarcation criterion for adenoviral species [10], showed that they were nearly 90% identical (Table 1). Phylogenetic analysis confirmed that the Rc-kw20 strain was positioned in the BtAdV-C cluster (Fig. 1b), and this was supported by analysis using hexon protein amino acid sequences (Fig. 1c), confirming that Rc-Kw20 is a BtAdV-C strain and that Rhinolophus cornutus should be included as a susceptible host for group 2 BtAdV.
We also generated a genomic map, which included the 54-bp-long inverted terminal repeat (ITR) sequences (Fig. 1d). Twenty of the 33 proteins encoded by Rc-kw20 shared high sequence similarity (more than 80% identity) with those of previously characterized BtAdV-C strains, whereas the fiber protein and several E3- and E4-related proteins shared moderate similarity (ranging from 48.0% to 67.2% sequence identity), and only the large E3 (E3l) protein, which contains host immune-suppression-related motifs [11], showed considerably lower similarity (31.8% to 34.6% sequence identity) (Table 1). Possible functional differences involving proteins with lower sequence similarity might determine the host specificity of BtAdV-C strains.
Recent studies have indicated that Rhinolophus spp. bats may represent an ancestral host for severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), which causes COVID-19. Indeed, related bat sarbecoviruses have been detected in Rhinolophus affinis, R. sinicus, R. pusillus, R. malayanus, R. acuminatus, and R. shameli in Asian countries [12,13,14,15,16,17]. In addition, we detected a bat sarbecovirus in R. cornutus in Japan [18]. Further studies are needed to assess the zoonotic potential of bat viruses inhabiting these Rhinolophus species, including BtAdV-C.
References
Lau SK, Woo PC, Li KS, Huang Y, Tsoi HW, Wong BH, Wong SS, Leung SY, Chan KH, Yuen KY (2005) Severe acute respiratory syndrome coronavirus-like virus in Chinese horseshoe bats. Proc Natl Acad Sci USA 102:14040–14045. https://doi.org/10.1073/pnas.0506735102
Epstein JH, Anthony SJ, Islam A, Kilpatrick AM, Ali Khan S, Balkey MD, Ross N, Smith I, Zambrana-Torrelio C, Tao Y, Islam A, Quan PL, Olival KJ, Khan MSU, Gurley ES, Hossein MJ, Field HE, Fielder MD, Briese T, Rahman M, Broder CC, Crameri G, Wang LF, Luby SP, Lipkin WI, Daszak P (2020) Nipah virus dynamics in bats and implications for spillover to humans. Proc Natl Acad Sci USA 117:29190–29201. https://doi.org/10.1073/pnas.2000429117
Johnson N, Vos A, Freuling C, Tordo N, Fooks AR, Müller T (2010) Human rabies due to lyssavirus infection of bat origin. Vet Microbiol 142:151–159. https://doi.org/10.1016/j.vetmic.2010.02.001
Kuzmin IV, Niezgoda M, Franka R, Agwanda B, Markotter W, Breiman RF, Shieh WJ, Zaki SR, Rupprecht CE (2010) Marburg virus in fruit bat, Kenya. Emerg Infect Dis 16:352–354. https://doi.org/10.3201/eid1602.091269
Kobayashi T, Matsugo H, Maruyama J, Kamiki H, Takada A, Maeda K, Takenaka-Uema A, Tohya Y, Murakami S, Horimoto T (2019) Characterization of a novel species of adenovirus from Japanese microbat and role of CXADR as its entry factor. Sci Rep 9:573. https://doi.org/10.1038/s41598-018-37224-z
Borkenhagen LK, Fieldhouse JK, Seto D, Gray GC (2019) Are adenoviruses zoonotic? A systematic review of the evidence. Emerg Microbes Infect 8:1679–1687. https://doi.org/10.1080/22221751.2019.1690953
Walker PJ, Siddell SG, Lefkowitz EJ, Mushegian AR, Adriaenssens EM, Dempsey DM, Dutilh BE, Harrach B, Harrison RL, Hendrickson RC, Junglen S, Knowles NJ, Kropinski AM, Krupovic M, Kuhn JH, Nibert M, Orton RJ, Rubino L, Sabanadzovic S, Simmonds P, Smith DB, Varsani A, Zerbini FM, Davison AJ (2020) Changes to virus taxonomy and the statutes ratified by the International Committee on Taxonomy of Viruses (2020). Arch Virol 165:2737–2748. https://doi.org/10.1007/s00705-020-04752-x
Ogawa H, Kajihara M, Nao N, Shigeno A, Fujikura D, Hang’ombe BM, Mweene AS, Mutemwa A, Squarre D, Yamada M, Higashi H, Sawa H, Takada A (2017) Characterization of a novel bat adenovirus isolated from straw-colored fruit bat (Eidolon helvum). Viruses 9:371. https://doi.org/10.3390/v9120371
Shirogane Y, Takeda M, Iwasaki M, Ishiguro N, Takeuchi H, Nakatsu Y, Tahara M, Kikuta H, Yanagi Y (2008) Efficient multiplication of human metapneumovirus in Vero cells expressing the transmembrane serine protease TMPRSS2. J Virol 82:8942–8946. https://doi.org/10.1128/JVI.00676-08
Benko M, Harrach B, Both GW, Russell WC, Adair BM, Adam E, deJong JC, Hess M, Johnson M, Kajon A, Kidd AH, Lehmkuhl HD, Li Q, Mautner V, Pring-Akerblom P, Wadell G (2005) Adenoviridae. In: Fauquet CM, Mayo MA, Maniloff J, Desselberger U, Ball LA (eds) Virus taxonomy. Eighth report of the international committee on the taxonomy of viruses. Elsevier Academic Press, San Diego, pp 213–228
Tan B, Yang XL, Ge XY, Peng C, Zhang YZ, Zhang LB, Shi ZL (2016) Novel bat adenoviruses with an extremely large E3 gene. J Gen Virol 97:1625–1635. https://doi.org/10.1099/jgv.0.000470
Li LL, Wang JL, Ma XU, Sun XM, Li JS, Yang XF, Shi WF, Duan ZJ (2021) A novel SARS-CoV-2 related coronavirus with complex recombination isolated from bats in Yunnan province, China. Emerg Microbes Infect 10:1683–1690. https://doi.org/10.1080/22221751.2021.1964925
Hassanin A, Tu VT, Curaudeau M, Csorba G (2021) Inferring the ecological niche of bat viruses closely related to SARS-CoV-2 using phylogeographic analyses of Rhinolophus species. Sci Rep 11:14276. https://doi.org/10.1038/s41598-021-93738-z
Wacharapluesadee S, Tan CW, Maneeorn P, Duengkae P, Zhu F, Joyjinda Y, Kaewpom T, Chia WN, Ampoot W, Lim BL, Worachotsueptrakun K, Chen VC, Sirichan N, Ruchisrisarod C, Rodpan A, Noradechanon K, Phaichana T, Jantarat N, Thongnumchaima B, Tu C, Crameri G, Stokes MM, Hemachudha T, Wang LF (2021) Evidence for SARS-CoV-2 related coronaviruses circulating in bats and pangolins in Southeast Asia. Nat Commun 12:972. https://doi.org/10.1038/s41467-021-21240-1
Zhou P, Yang XL, Wang XG, Hu B, Zhang L, Zhang W, Si HR, Zhu Y, Li B, Huang CL, Chen HD, Chen J, Luo Y, Guo H, Jiang RD, Liu MQ, Chen Y, Shen XR, Wang X, Zheng XS, Zhao K, Chen QJ, Deng F, Liu LL, Yan B, Zhan FX, Wang YY, Xiao GF, Shi ZL (2020) A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 579:270–273. https://doi.org/10.1038/s41586-020-2012-7
Zhou H, Chen X, Hu T, Li J, Song H, Liu Y, Wang P, Liu D, Yang J, Holmes EC, Hughes AC, Bi Y, Shi W (2020) A novel bat coronavirus closely related to SARS-CoV-2 contains natural insertions at the S1/S2 cleavage site of the spike protein. Curr Biol 30:2196-2203.e3. https://doi.org/10.1016/j.cub.2020.05.023
Lau SKP, Luk HKH, Wong ACP, Li KSM, Zhu L, He Z, Fung J, Chan TTY, Fung KSC, Woo PCY (2020) Possible bat origin of severe acute respiratory syndrome coronavirus 2. Emerg Infect Dis 26:1542–1547. https://doi.org/10.3201/eid2607.200092
Murakami S, Kitamura T, Suzuki J, Sato R, Aoi T, Fujii M, Matsugo H, Kamiki H, Ishida H, Takenaka-Uema A, Shimojima M, Horimoto T (2020) Detection and characterization of bat sarbecovirus phylogenetically related to SARS-CoV-2, Japan. Emerg Infect Dis 26:3025–3029. https://doi.org/10.3201/eid2612.203386
Funding
This work was supported by the Japan Agency for Medical Research and Development (AMED) under Grant numbers JP21fk0108602 and JP21fk0108615.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethical approval
This study did not involve experiments with human participants performed by any of the authors. Experiments with bat samples were approved by the Animal Experiment Committee of the Graduate School of Agricultural and Life Sciences at the University of Tokyo (approval number P21-058).
Additional information
Handling Editor: T. K. Frey.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Katayama, M., Murakami, S., Matsugo, H. et al. Complete genome sequence of a novel bat mastadenovirus C strain isolated from Rhinolophus cornutus in Japan. Arch Virol 167, 979–982 (2022). https://doi.org/10.1007/s00705-021-05357-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-021-05357-8