Abstract
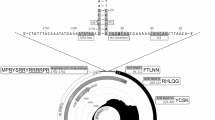
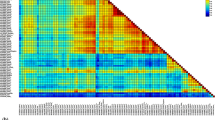
The subfamily Parvovirinae within the family Parvoviridae consists of viruses that can infect a wide range of vertebrate hosts and cause effects ranging from severe disease to asymptomatic infection. In the present study, high-throughput sequencing (HTS) was utilized to analyze samples obtained from an abortion outbreak in a sheep flock to identify a putative viral etiology. A highly divergent nearly complete parvovirid genome sequence, approximately 4.9 kb in length, was determined. The nonstructural protein (NS1) amino acid (aa) sequence of this virus shared less than 30% identity with those of other copiparvoviruses and less than 22% identity with those of members of other genera in the subfamily Parvovirinae. Phylogenetically, this virus, which we have provisionally named "sheep copiparvovirus 1", formed a cluster with copiparvovirus sequences and should be classified as a member of a new species in the genus Copiparvovirus.

Similar content being viewed by others
Availability of data and materials
The genome sequence and predicted amino acid sequence described here are available in the GenBank database under the accession number MT875453.
References
Cotmore SF, Agbandje-McKenna M, Canuti M et al (2019) ICTV Report Consortium, 2019, ICTV Virus Taxonomy Profile: Parvoviridae. J Gen Virol 100:367–368
Allander T, Emerson SU, Engle RE, Purcell RH, Bukh J (2011) A virus discovery method incorporating DNase treatment and its application to the identification of two bovine parvovirus species. Proc Natl Acad Sci USA 98:11609–11614
Cheung AK, Wu G, Wang D, Bayles DO, Lager KM, Vincent AL (2010) Identification and molecular cloning of a novel porcine parvovirus. Arch Virol. 155:801–806
Weber MN, Mosena ACS, da Silva MS et al (2020) Virome of crab-eating (Cerdocyon thous) and pampas foxes (Lycalopex gymnocercus) from southern Brazil and Uruguay. Infect Genet Evol 85:104421
Cotmore SF, Tattersall P (2014) Parvoviruses: small does not mean simple. Annu Rev Virol 1:517–537
Koonin EV (1993) A common set of conserved motifs in a vast variety of putative nucleic acid-dependent ATPases including MCM proteins involved in the initiation of eukaryotic DNA replication. Nucleic Acids Res 21:2541–2547
Zádori Z, Szelei J, Lacoste M et al (2001) A viral phospholipase A2 is required for parvovirus infectivity. Dev Cell 1:291–302
Acknowledgements
We are grateful to Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), Fundação de Amparo à Pesquisa do Estado do Rio Grande do Sul (FAPERGS), Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES) and Propesq/UFRGS for financial support to this study
Funding
Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES)-Finance Code 001, Fundação de Amparo à Pesquisa do Rio Grande do Sul (FAPERGS), and Pró-reitoria de Pesquisa da Universidade Federal do Rio Grande do Sul (Propesq/UFRGS).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Additional information
Handling Editor: Ana Cristina Bratanich.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Mosena, A.C.S., da Silva, M.S., Lorenzett, M.P. et al. A new highly divergent copiparvovirus in sheep. Arch Virol 166, 1517–1520 (2021). https://doi.org/10.1007/s00705-021-05020-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-021-05020-2