Abstract
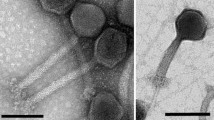
In this study, we isolated a mycobacteriophage infecting Mycobacterium smegmatis mc2155 from a soil sample collected in Shandong Province in China. This phage was named Shandong1. It is a member of the family Siphoviridae with an isometric head and a long tail. Its genome was found to be 60,618 bp long with 67.46% G + C content and 96 putative protein-coding genes. No tRNA-encoding genes were identified. Comparative genomics analysis showed that the mycobacteriophage Shandong1 should be considered a member of a new species in mycobacteriophage cluster K.


Similar content being viewed by others
References
Organization WH (2016) Global tuberculosis report 2016
Liu J, Dehbi M, Moeck G, Arhin F, Bauda P, Bergeron D, Callejo M, Ferretti V, Ha N, Kwan T (2004) Antimicrobial drug discovery through bacteriophage genomics. Nat Biotechnol 22(2):185–191
Chung IY, Jang HJ, Bae HW, Cho YH (2014) A phage protein that inhibits the bacterial ATPase required for type IV pilus assembly. P Natl Acad Sci USA 111(31):11503–11508
Molshanski-Mor S, Yosef I, Kiro R, Edgar R, Manor M, Gershovits M, Laserson M, Pupko T, Qimron U (2014) Revealing bacterial targets of growth inhibitors encoded by bacteriophage T7. P Natl Acad Sci USA 111(52):18715–18720
Fan X, Yan J, Xie L, Zeng L, Young RF, Xie J (2015) Genomic and proteomic features of mycobacteriophage SWU1 isolated from China soil. Gene 561(1):45–53
Delcher AL, Bratke KA, Powers EC, Salzberg SL (2007) Identifying bacterial genes and endosymbiont DNA with Glimmer. Bioinformatics 23(6):673–679
Schattner P, Brooks AN, Lowe TM (2005) The tRNAscan-SE, snoscan and snoGPS web servers for the detection of tRNAs and snoRNAs. Nucleic Acids Res 33(Web Server issue):W686–W689
Dong TG, Dong S, Catalano C, Moore R, Liang X, Mekalanos JJ (2015) Generation of reactive oxygen species by lethal attacks from competing microbes. Proc Natl Acad Sci 112(7):2181–2186
Samaddar S, Grewal RK, Sinha S, Ghosh S, Roy S, Gupta SKD (2016) Dynamics of Mycobacteriophage-mycobacterial host interaction: evidence for secondary mechanisms for host lethality. Appl Environ Microbiol 82(1):124–133
Pope WH, Ferreira CM, Jacobs-Sera D, Benjamin RC, Davis AJ, DeJong RJ, Elgin SC, Guilfoile FR, Forsyth MH, Harris AD, Harvey SE, Hughes LE, Hynes PM, Jackson AS, Jalal MD, MacMurray EA, Manley CM, McDonough MJ, Mosier JL, Osterbann LJ, Rabinowitz HS, Rhyan CN, Russell DA, Saha MS, Shaffer CD, Simon SE, Sims EF, Tovar IG, Weisser EG, Wertz JT, Weston-Hafer KA, Williamson KE, Zhang B, Cresawn SG, Jain P, Piuri M, Jacobs WR Jr, Hendrix RW, Hatfull GF (2011) Cluster K mycobacteriophages: insights into the evolutionary origins of mycobacteriophage TM4. PLoS One 6(10):e26750
Hatfull GF, Jacobs-Sera D, Lawrence JG, Pope WH, Russell DA, Ko CC, Weber RJ, Patel MC, Germane KL, Edgar RH (2010) Comparative genomic analysis of 60 mycobacteriophage genomes: genome clustering, gene acquisition, and gene size. J Mol Biol 397(1):119–143
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
This study was funded by the National Natural Science Foundation (Grant number 31600148, 81371851, 81071316, 81271882), the National Key R&D Project (Grant number 2016YFC0502304), the Shandong Excellent Young Scientist Award Fund (Grant number BS2014YY031), and the Foundation of the University of Jinan (Grant number XBS1519, XKY1633, XBS1442).
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Fan, X., Gao, X., Wang, Q. et al. Complete genome sequence analysis of the novel mycobacteriophage Shandong1. Arch Virol 162, 3903–3905 (2017). https://doi.org/10.1007/s00705-017-3534-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-017-3534-7