Abstract
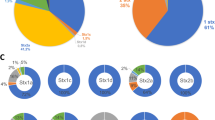
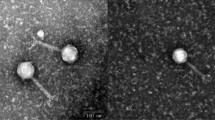
We report the complete genome sequencing of two Escherichia coli T5-related bacteriophages, DT57C and DT571/2, isolated from the same specimen of horse feces. These two isolates share 96 % nucleotide sequence identity and can thus be considered representatives of the same novel species within the genus T5likevirus. The observed variation in the ltfA gene of these phages, resulting from a recent recombination event, may explain the observed host-range differences, suggesting that a modular mechanism makes a significant contribution to the short-term evolution (or adaptation) of T5-like phage genomes in the intestinal ecosystem. Comparison of our isolates to their closest relative, coliphage T5, revealed high overall synteny of the genomes and high conservation of the sequences of almost all structural proteins as well as of the other proteins with identified functions. At the same time, numerous alterations and non-orthologous replacements of non-structural protein genes (mostly of those with unknown functions) as well as substantial differences in tail fiber locus organization support the conclusion that DT57C and DT571/2 form a species-level group clearly distinct from bacteriophage T5.

Similar content being viewed by others
References
Waller AS, Yamada T, Kristensen DM, Kultima JR, Sunagawa S, Koonin EV, Bork P (2014) Classification and quantification of bacteriophage taxa in human gut metagenomes. ISME J 8(7):1551–1552. doi:10.1038/ismej.2014.66
Niu YD, Stanford K, Kropinski AM, Ackermann HW, Johnson RP, She YM, Ahmed R, Villegas A, McAllister TA (2012) Genomic, proteomic and physiological characterization of a T5-like bacteriophage for control of Shiga toxin-producing Escherichia coli O157:H7. PLoS one 7(4):e34585. doi:10.1371/journal.pone.0034585
Letarov A, Kulikov E (2009) The bacteriophages in human- and animal body-associated microbial communities. J Appl Microbiol 107(1):1–13. doi:10.1111/j.1365-2672.2009.04143.x
Letarov A (2012) Bacteriophages as a part of the human microbiome. In: Hyman P, Abedon ST (eds) Bacteriophages in health and disease. Advances in molecular and cellular microbiology, vol 24. CABI Press, Wallingford, pp 6–20
Zivanovic Y, Confalonieri F, Ponchon L, Lurz R, Chami M, Flayhan A, Renouard M, Huet A, Decottignies P, Davidson AR, Breyton C, Boulanger P (2014) Insights into bacteriophage T5 structure from analysis of its morphogenesis genes and protein components. J Virol 88(2):1162–1174. doi:10.1128/JVI.02262-13
Sayers J (2006) Bacteriophage T5. In: Calendar R (ed) The bacteriophages, 2nd edn. Oxford University Press, Oxford, pp 268–276
Knirel YA, Prokhorov NS, Shashkov AS, Ovchinnikova OG, Zdorovenko EL, Liu B, Kostryukova ES, Larin AK, Golomidova AK, Letarov AV (2015) Variations in O-antigen biosynthesis and O-acetylation associated with altered phage sensitivity in Escherichia coli 4s. J Bacteriol 197(5):905–912. doi:10.1128/JB.02398-14
Golomidova A, Kulikov E, Isaeva A, Manykin A, Letarov A (2007) The diversity of coliphages and coliforms in horse feces reveals a complex pattern of ecological interactions. Appl Environ Microbiol 73:5975–5981. doi:10.1128/AEM.01145-07
Kulikov E, Kropinski AM, Golomidova A, Lingohr E, Govorun V, Serebryakova M, Prokhorov N, Letarova M, Manykin A, Strotskaya A, Letarov A (2012) Isolation and characterization of a novel indigenous intestinal N4-related coliphage vB_EcoP_G7C. Virology 426(2):93–99. doi:10.1016/j.virol.2012.01.027
Zdorovenko EL, Golomidova AK, Prokhorov NS, Shashkov AS, Wang L, Letarov AV, Knirel YA (2015) Structure of the O-polysaccharide of Escherichia coli O87. Carbohydr Res 412:15–18. doi:10.1016/j.carres.2015.04.014
Ong ZY, Wiradharma N, Yang YY (2014) Strategies employed in the design and optimization of synthetic antimicrobial peptide amphiphiles with enhanced therapeutic potentials. Adv Drug Deliv Rev 78:28–45. doi:10.1016/j.addr.2014.10.013
Mondigler M, Ayoub AT, Heller KJ (2006) The DNA region of phage BF23 encoding receptor binding protein and receptor blocking lipoprotein lacks homology to the corresponding region of closely related phage T5. J Basic Microbiol 46(2):116–125. doi:10.1002/jobm.200510047
Acknowledgments
We would like to acknowledge Dr. S. T. Abedon (The Ohio State University, USA) for the critical reading and linguistic correction of the manuscript. The work of the laboratory in INMI RAS is supported by RSCF grant #15-15-00134 (comparative sequence analysis, morphology analysis). R.C.G.-F. work is supported by Swiss National Science Foundation grant #31003A_146284.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Rights and permissions
About this article
Cite this article
Golomidova, A.K., Kulikov, E.E., Prokhorov, N.S. et al. Complete genome sequences of T5-related Escherichia coli bacteriophages DT57C and DT571/2 isolated from horse feces. Arch Virol 160, 3133–3137 (2015). https://doi.org/10.1007/s00705-015-2582-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-015-2582-0