Abstract
Pandemic influenza A virus (H1N1) 2009 poses a serious public-health challenge worldwide. To characterize the neutralizing epitopes of this virus, we generated a panel of eight monoclonal antibodies (mAbs) against the HA of the A/California/07/2009 virus. The antibodies were specific for the 2009 pdm H1N1 HA, as the antibodies displayed HA-specific ELISA, hemagglutination inhibition (HAI) and neutralization activity. One mAb (mAb12) showed significantly higher HAI and neutralizing titers than the other mAbs. We mapped the antigenic epitopes of the HA by characterizing escape mutants of a 2009 H1N1 vaccine strain (NYMC X-179A). The amino acid changes suggested that these eight mAbs recognized HA antigenic epitopes located in the Sa, Sb, Ca1 and Ca2 sites. Passive immunization with mAbs showed that mAb12 displayed more efficient neutralizing activity in vivo than the other mAbs. mAb12 was also found to be protective, both prophylactically and therapeutically, against a lethal viral challenge in mice. In addition, a single injection of 10 mg/kg mAb12 outperformed a 5-day course of treatment with oseltamivir (10 mg/kg/day by gavage) with respect to both prophylaxis and treatment of lethal viral infection. Taken together, our results showed that mouse-origin mAbs displayed neutralizing effectiveness in vitro and in vivo. One mAb in particular (mAb12) recognized an epitope within the Sb site and demonstrated outstanding neutralizing effectiveness.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The 2009 flu pandemic, which began in Mexico and was caused by a new variant of H1N1 swine influenza virus, was a global threat to public health [1]. A(H1N1)pdm09 virus spread rapidly around the world after the initial outbreak. When the World Health Organization (WHO) declared a pandemic, a total of 74 countries and territories had reported laboratory-confirmed infections. A(H1N1)pdm09 caused more than 18,000 deaths worldwide between April 2009 and August 2010 [2]. A(H1N1)pdm09 virus is a novel influenza virus, and its genome originates from a combination of North American and Eurasian swine influenza virus lineages [3–5]. A(H1N1)pdm09 viruses can evolve in humans, and genetic analysis has demonstrated that A(H1N1)pdm09 virus has diversified into seven discrete global clades [6]. Furthermore, co-circulation of multiple strains belonging to different clades has been reported [7–10]. Mutations that are associated with antigenic changes can occur in the A(H1N1)pdm09 hemagglutinin (HA). Recently, Igarashi et al. reported that amino acid substitutions in an antigenic epitope have already been found in variants of the A(H1N1)pdm09 virus [11]. Thus, we can speculate that further antigenic mutations will appear in the future, following the seasonal circulation of the A(H1N1)pdm09 virus in humans. Therefore, mapping monoclonal antibody (mAb) epitopes is a necessary step towards understanding antigenic drift in humans.
The influenza A virus genome is composed of eight segments of negative-sense RNA that encode up to 12 proteins. The HA protein is one of two major surface glycoproteins present in the envelope of influenza A virus. Influenza A viruses of 16 (H1-H16) HA subtypes have been identified in wild birds, and a possibly H17-like subtype was found in bats [12, 13]. Only three subtypes (H1, H2 and H3) have been recognized to establish pandemics in the human population. The HA protein is responsible for binding to host-cell receptors and for viral entry and is therefore a primary target of neutralizing antibodies [14]. Thus, HA-specific neutralizing mAbs can bind antigenic epitopes and prevent binding of virions to host cells. This characteristic of mAbs enables the identification of antigenic sites in HA, through the selection of escape mutants. Classical studies using mAbs have identified antigenic epitopes of multiple subtypes of HA, such as H1, H2, H3 and H5 [15–18]. The H1 HA molecules have five distinct antigenic sites: Sa, Sb, Ca1, Ca2 and Cb [15, 19, 20]. Identification of antigenic regions in A(H1N1)pdm09 virus that are capable of eliciting neutralizing antibodies is crucial for the understanding of the epidemiology of this disease.
Antibody immunotherapy is an effective therapeutic strategy and is increasingly used to treat numerous human infectious diseases [21, 22]. Treatment with convalescent-phase blood and serum products resulted in the distinct improvement of clinical outcomes in severely ill influenza patients during the 1918 influenza pandemic, as well as in H5N1 and A(H1N1)pdm09 virus infection in humans [23–25]. In animal models, specific mAbs can confer prophylactic and therapeutic protection against influenza virus infection [26–29]. These observations indicate that the passive administration of antibodies could be a potentially useful treatment and prophylactic option for severe courses of disease.
In the present study, we generated a panel of eight mAbs that specifically recognized the HA of A(H1N1)pdm09. We performed epitope mapping and determined the neutralizing properties and protective efficacy of these mAbs. We also compared the prophylactic and therapeutic efficacies of the mAb12 with an antiviral drug, oseltamivir, in a mouse model of viral infection.
Material and methods
Virus preparation
Strain NYMC X-179A [A/reassortant/NYMC X-179A (California/07/2009 x NYMC X-157) (H1N1)] was generated by New York Medical College and supplied by the Centers for Disease Control and Prevention (USA). This strain is a pandemic vaccine strain recommended for use in vaccine development [30]. Other influenza virus strains, A/PR/8/34 (H1N1) and A/New Caledonia/20/1999 (H1N1), were propagated in the allantoic cavities of 10-day-old embryonated chicken eggs at 37 °C for 48 h and stored at −80 °C until use. The seed virus was grown in embryonated eggs, and the 50 % tissue culture infectious dose (TCID50) was determined in Madin-Darby canine kidney (MDCK) cells according to Reed-Muench method [31]. The recombinant virus rgH1N12009/PR8, which possesses the HA and NA of A/California/04/2009 and the six remaining gene segments from a mouse-adapted A/PR/8/34, was generated as described previously [32]. The 50 % mouse lethal dose (MLD50) of rgH1N12009/PR8 was determined, and the result was calculated by the Reed-Muench method.
Generation of mAbs
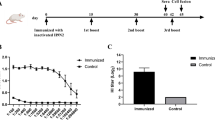
In vivo electroporation was carried out according to a method described previously [33]. Briefly, adult female BALB/c mice (6–8 weeks old) were immunized intramuscularly with two doses of 50 μg plasmid DNA encoding the full-length HA gene of the 2009 H1N1 strain A/California/04/2009. After injection in the right quadriceps muscle, electric pulses were delivered using an electric pulse generator (ECM830; BTX, San Diego, CA). Three pulses of 100 V each, followed by three pulses of the opposite polarity, were delivered to each injection site at a rate of one pulse per second [33]. Each pulse lasted for 50 ms. After the second immunization of DNA vaccine, mice were immunized intraperitoneally one time with 2 μg inactivated NYMC X-179A vaccine (manufactured by Shanghai Institute of Biological Products). Hybridization of splenocytes from these mice with Sp2/0 myeloma cells was then performed. Hybridoma supernatants were used for screening of mAbs for reactivity by enzyme-linked immunosorbent assay (ELISA). Fourteen hybridomas were used to obtain mAb-containing mouse ascites fluids. RDE-treated ascites fluids containing antiviral antibodies were selected and used for measurement of microneutralization. The mAbs were then purified from ascites by affinity chromatography using an IgG purification kit (Pierce) before use. The isotypes of the mAbs were determined with mouse monoclonal antibody isotyping reagents by indirect ELISA according to the manufacturer’s procedure (IsoQuick™ Strips for Mouse Monoclonal Isotyping, Sigma-Aldrich, St. Louis, MO, USA).
ELISA
ELISA was performed using a 96-well plate (EIA plate, Costar, Cambridge, MA, USA) with reagents consisting of, first, inactivated vaccine of NYMC X-179A, A/PR/8/34 or A/New Caledonia/20/1999(diluted in 10 μg/ml), manufactured by Shanghai Institute of Biological Products (SIBP); second, serial dilutions of monoclonal antibodies; third, goat anti-mouse IgG Ab (γ-chain specific) (Southern Biotechnology Associates, Inc., USA) conjugated with biotin; fourth, streptavidin conjugated with alkaline phosphatase (Southern Biotechnology Associates, Inc., USA); and finally, p-nitrophenyl phosphate. The amount of chromogen produced was measured based on absorbance at 410 nm and 630 nm using an ELISA reader (Genios, Tecan).
HAI and MN assays
Hemagglutination inhibition (HAI) was performed to screen for H1-specific mAbs as described elsewhere with some modifications [34]. Purified mAbs were serially diluted (twofold) in V-bottom 96-well plates. Four HA units of viral antigen was incubated with the mAbs at room temperature for 1 h, followed by the addition of 0.5 % chicken red blood cells and incubation at room temperature for 30 minutes. The HAI titres were determined as the reciprocals of the highest dilutions that completely inhibited hemagglutination.
Cell-based microneutralization (MN) assays were performed as described previously [35]. Viruses were incubated with mAb at room temperature for one hour, with rocking. The mixture containing virus and mAb was then transferred to the wells of a 96-well plate containing a confluent layer of MDCK cells and incubated for 48 hours at 37 °C. Individual determinations were performed in triplicate. Endpoints were determined by hemagglutination assay. The microneutralizing titer was defined as the highest dilution of antibody at which all of the culture wells were negative for infection.
Selection of escape mutants
Escape mutants were selected by culturing NYMC X-179A in embryonated chicken eggs in the presence of mAb as described previously [36]. Briefly, viruses (10 HAU/ml) were incubated with purified mAb (1000 HAIU/ml) for 1 h, and the mixtures were then inoculated into the allantoic cavities of 9-day-old specific-pathogen-free (SPF) embryonated chicken eggs. The virus yield was used for limiting-dilution cloning in embryonated chicken eggs.
Viral RNA was extracted by lysing the viruses with TRIzol LS Reagent (Life Technologies, Inc.). The RNA was reverse transcribed into single-stranded DNA with M-MuLV reverse transcriptase (New England Biolabs). The HA gene was amplified using a Phusion™ High-Fidelity PCR Kit (New England Biolabs), with segment-specific primers [37]. The PCR products were purified using a Cycle-Pure Kit and a Gel Extraction Kit (Omega Bio-Tek, USA), and the fragments were cloned into pGEM-T Easy Vector. The fragments were sequenced and edited using DNASTAR sequence analysis software (DNASTAR Inc.).
RDE avidity assay
As described previously [38], a 10 % (vol/vol) chicken erythrocyte solution was treated with various concentrations (1 U, 100 mU, 50 mU, 25 mU, 10 mU, 5 mU, and 0 U) of receptor-destroying enzyme (RDE, Denka Seiken, Japan) for 1 h at 37 °C. The cells were washed, diluted to 2 % (vol/vol) in PBS, and added to 96-well V-bottomed plates containing 4 HA units of the mutant and wt viruses. The plates were gently shaken, and the results were recorded after incubation for 30 min at room temperature.
In vivo protection experiments
All animal experiments were conducted in accordance with ethical procedures and policies approved by the Wuhan Institute of Virology’s Institutional Animal Care and Use Committee. Six- to eight-week-old female BALB/c mice were used for the challenge studies. Mice were anesthetized by intraperitoneal administration of chloroform and challenged with rgH1N12009/PR8. To evaluate the prophylactic efficacy of the mAbs, mice were injected intraperitoneally with specific mAbs. At various times after passive administration (24, 48 or 72 h), mice were challenged with a lethal dose of rgH1N12009/PR8. Animals were observed daily for mortality and morbidity, and body weight was measured for up to 14 days after infection. To determine the therapeutic efficacy of the mAbs, mice were challenged with a lethal dose of rgH1N12009/PR8. At various times after infection (24, 48 or 72 h), mice were treated intraperitoneally with specific mAbs. Animals were observed daily for mortality and morbidity, and body weight was measured for up to 14 days after infection.
Statistical analysis
For survival, probability was calculated using Fisher’s exact test, comparing the rate of survival of mice treated with specific mAbs with those of the control groups.
Results
Generation and characterization of A(H1N1)pdm09 HA-specific mAbs
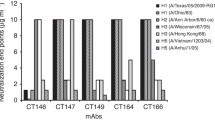
A panel of eight hybridoma clones secreting mAbs to A(H1N1)pdm09 HA was screened. As shown in Table 1, these clones showed strong ELISA binding activity to NYMC X-179A inactivated vaccine, but not to seasonal flu virus A/New Caledonia/20/1999 or A/PR/8/34. HA inhibition (HAI) studies showed that the mAbs were capable of inhibiting HA reactions by NYMC X-179A virus and erythrocytes and that the HAI titer was within the range of 1:32 to 1:2048. The HAI titer of mAb12 was significantly higher than those of the other mAbs. None of the mAbs could inhibit hemagglutination of A/New Caledonia/20/1999 or A/PR/8/34. A microneutralization (MN) test gave a result similar to that obtained using the HAI test, in which the eight mAbs could specifically inhibit the proliferation of NYMC X-179A virus in MDCK cells, and the mAb12 MN titer was significantly higher those of the other mAbs (MN titer 1:32,768). None of the mAbs could neutralize A/New Caledonia/20/1999 or A/PR/8/34. All eight mAbs had potency in HAI and MN, and thus could recognize the HA globular head epitope. Antigen classification results showed that there were five strains with the IgG1 subtype, two strains with IgG2a and one strain with IgG2b. Our results showed that the eight mAbs were A(H1N1)pdm09 HA specific antibodies, and the in vitro virus-neutralizing capability of mAb12 was significantly higher than those of other mAbs.
Epitope mapping of mAbs
Co-culture mixture of virus and an antibody in the allantoic cavity of chick embryos, which can produce escape strains, was used to analyze amino acid mutations to determine the antigenic sites that were recognized by the mAbs [16, 36]. We used this method to obtain mAb escape strains and to map the epitopes of the mAbs. As shown in Table 2 and Fig. 1, the mAb3 escape strain had a G158E mutation located in the Sa antigenic site. MAb1 and mAb10 recognized the same epitope, and both of their escape mutants had an S188N mutation. The escape strain of mAb12 had a Q192E mutation. These mutations were located in the Sb antigenic site. The G173E mutation of the mAb8 escape mutant was located in the Ca1 antigenic site. The K145E and A144E mutations in the site recognized by mAb7 and mAb11 were located in the Ca2 antigenic site. The mAb2 escape strain had a K242E mutation, which was located adjacent to Sa and Ca1 and could represent a unique epitope not identified previously. We then performed an ELISA binding assay to distinguish antigenic mutations and adsorptive mutations. It has been demonstrated that antigenic mutants show significantly lower optical density (OD) values than do the wt parental viruses, while adsorptive mutants show a binding pattern and OD values similar to those of the wt parental viruses [38, 39]. Fig. S1 shows the results from an antibody binding ELISA in which eight viruses with mutations at A144E (mAb11-mt), K145E (mAb7-mt), G158E (mAb3-mt), G173E (mAb8-mt), S188 N (mAb1-mt), Q192E (mAb12-mt) and K242E (mAb2-mt), respectively, demonstrated significantly reduced binding to the corresponding mAb compared to the wt parental virus. This indicated that these escape mutants are antigenic mutants and not adsorptive mutants. This was confirmed by measuring RDE avidity, which showed that all of the mutants had titers similar to that of wt virus (data not shown).
Positions of amino acid changes in the antigenic sites of HA of A/California/07/2009 (H1N1) influenza virus. The HA structure was obtained from the Protein Data Bank (accession number 3LZG), and images were created using PyMOL 0.99. Amino acid positions are designated according to mature H3 numbering. Antigenic sites are indicated as colored surfaces as follows: Sa, red; Sb, green; Ca1, yellow; Ca2, magenta; and Cb, cyan
We also analyzed the HAI activity of the eight mAbs with the escape mutant strains. As shown in Table 3, the inhibition titer of the mAbs for mAb3-mt was significantly lower than that of the parental strain, which indicated the mAb3-mt mutation significantly affected the recognition of HA by the mAbs. The inhibition titer of mAb11 with other escape strains (excluding mAb3-mt) was comparable to that of the parent strain, which indicated that there was little impact of the Sa and Sb antigenic site mutations on the binding of mAb11 to A144 of Ca2. Similarly, the inhibition titer of mAb12 with mAb7-mt, mAb8-mt and mAb11-mt was unaffected, which indicated that the mutation of Ca1- and Ca2-related sites of the virus had little impact on the binding of mAb12 to the Sb antigenic site Q192. In addition, the titers of mAb1, mAb2, mAb10 and mAb12 were significantly decreased for three other strains of mAb-mt, which indicated that mutations in the Sa or Sb antigenic sites had a significant mutual impact.
Incidence of amino acid substitutions among recent H1N1 isolates
To determine whether the same mAb escape strain mutation had emerged in field strains, we selected nucleotide sequences of 1631 virus strains (946 in 2009, 347 in 2010 and 338 in 2011) from the GenBank database and analyzed the mutations of the corresponding sites. As shown in Table 4, mutations of K145E, G173E, Q192E and K242E were not observed in field strains. Both the G158E and S188 N mutations appeared in 2009 and disappeared in 2011, which indicated that these mutations occurred in quasispecies of A(H1N1)pdm09. Only the A144E mutation screened by mAb11 occurred once every year. This suggests that although several mAb escape mutants were found in field strains, the incidence of the mAb-selected mutations in field strains was relatively low.
Therapeutic protection of mAbs against challenge in mice
The in vitro study results demonstrated that the eight mAbs displayed neutralizing efficacy against the virus, so we further analyzed their neutralizing capability in vivo. Mice were infected with 5 MLD50 of rgH1N12009/PR8 virus, followed by intraperitoneal injection with 10 mg/kg or 2 mg/kg of mAbs 24 hours after infection. The weight loss and survival rate of the mice were monitored over 2 weeks. As shown in Fig. 2, the therapeutic effect of the mAbs was dose dependent, with higher doses resulting in a higher survival rate. A high dose of mAb2 or mAb12 could provide 100 % protection, and except for mAb7, the other mAbs could provide a partial protective effect, while all mice in the control group died within 11 days of infection. The neutralizing capability of mAb12 in vitro was significantly greater than that of the other mAbs, and correspondingly, the in vivo therapeutic effect of mAb12 was greater than the other mAbs. In the 10 mg/kg treatment group, the infected mice experienced only a 6 % weight loss. Even at a low dose of 2 mg/kg, mAb12 could provide 100 % protection of mice. These results showed that mAb12, which recognized the Sb antigenic site, exerted an outstanding neutralizing effect in vivo.
Survival rate and weight loss of mice infected with rgH1N12009/PR8. Mice were challenged intranasally with 5 MLD50 of rgH1N12009/PR8 and then received 10 mg or 2 mg per kg body weight of mAbs intraperitoneally 24 h after challenge. Mice were monitored daily for 14 days for survival and body weight loss. The control group was injected with PBS after infection
Prophylactic and therapeutic efficacy of mAb12 against viral infection
In this study, we further analyzed the prophylactic and therapeutic effects of mAb12 against viral infection in mice. In the prophylactic experiment, each group of mice (n = 5) received an intraperitoneal injection of 10 mg/kg mAb12 at 72, 48 and 24 hours before infection, respectively. Mice injected with PBS were used as a control group. Mice were then infected with 10 MLD50 of lethal virus, and survival and weight loss were observed for 2 weeks. As shown in Fig. 3, all mAb12-treated mice survived and displayed a gradual increase in weight over the two-week observation period. In contrast, all of the control mice died following infection and experienced a 40 % weight loss before death. In the therapeutic experiment, each group of mice (n = 5) was infected with 10 MLD50 of virus and received intraperitoneal injection of 10 mg/kg mAb12 at 24, 48 and 72 hours after infection. Mice injected with PBS were used as a control group. As shown in Fig. 3, all mAb12-treated mice survived but experienced weight loss to varying degrees. Mice treated after 24 hours of infection experienced 20 % weight loss after 6 days but gradually recovered, and their weight returned to pre-infection levels 2 weeks after infection. Mice receiving mAb12 48 hours and 72 hours after infection lost up to 35 % of their weight nine days after infection, which they then gradually recovered. Mice receiving mAb12 after 48 hours had a relatively faster weight recovery rate than mice receiving mAb12 treatment after 72 hours. The weight loss of mice in the control group reached 40 % before death. These results demonstrated that mAb12 has prophylactic and therapeutic activity on lethal viral infections in vivo, and the prophylactic efficacy was greater than its therapeutic efficacy.
Prophylactic and therapeutic protection against rgH1N12009/PR8 by mAb12. To test for prophylactic efficacy, mice were passively immunized with mAb12 (10 mg/kg) or with PBS and were subjected to viral challenge with rgH1N12009/PR8 (10 MLD50) 24, 48 and 72 h later. Mice were monitored daily for 14 days for survival and body weight loss. To test for therapeutic efficacy, mAb12 was administered 24, 48 and 72 h after viral challenge, and the mice were monitored as described above
Comparison of prophylactic and therapeutic efficacy of mAb12 with oseltamivir
Oseltamivir is an important drug for the clinical prevention and treatment of influenza virus infection, and in this experiment, we compared the in vivo effectiveness of mAb12 with oseltamivir to prevent or treat against infection with rgH1N12009/PR8 virus. To compare the prophylactic efficacy, one group of mice (n = 6) received a single intraperitoneal injection of 10 mg/kg mAb12 one day before viral challenge, and a second group (n = 6) received a daily gavage of 10 mg/kg oseltamivir for 5 consecutive days, starting one day before infection. The control group was injected with PBS before infection. All mice were then infected with 10 MLD50 of lethal virus, and survival and weight loss were monitored daily for 2 weeks. As shown in Fig. 4, all mAb12-treated mice survived (100 %, 6/6), almost no weight loss was observed after infection, and the body weight of treated mice increased gradually over the two-week observation period. Oseltamivir-treated mice had a survival rate of 33 % (2/6), and the mice suffered continuous weight loss after infection, which reached a maximum at day 12 (about 40 %), after which they gradually recovered. In the control group, all mice died after 9 days of infection, and the weight loss reached 40 % before death. In the therapeutic experiments, mice were infected with 10 MLD50 of lethal virus. Three days after infection, the mice developed clinical symptoms such as piloerection, shortness of breath, and slow movement. At this time, one group of mice (n = 6) received a single intraperitoneal injection of 10 mg/kg mAb12, and a second group received a once daily gavage with 10 mg/kg of oseltamivir for 5 consecutive days. The control group was injected with PBS. As shown in Fig. 4, although mAb12-treated mice suffered progressive weight loss after infection, which reached a maximum (about 30 %) at day 8, all mice survived, and their body weight gradually recovered after day 8. Oseltamivir-treated mice suffered progressive weight loss after infection, from which they did not recover even after drug administration, and the mice all died within 12 days when the weight loss reached its maximum of 40 %. Mice in the control group all died within 9 days after infection and suffered a continuing loss of weight after infection. This result showed that mAb12 is more effective in the prevention and treatment of viral infections in vivo than oseltamivir.
Prophylactic and therapeutic efficacy of mAb12 and oseltamivir against lethal challenge with rgH1N12009/PR8. To test for prophylactic efficacy, mice received either a single intraperitoneal injection of 10 mg/kg mAb12 one day before challenge or 10 mg/kg oseltamivir orally by gavage for 5 days starting 1 day before the virus challenge. The control group was injected with PBS after infection. Mice were monitored daily for 14 days for survival and body weight loss. To test for therapeutic efficacy, mice received either a single intraperitoneal injection of 10 mg/kg mAb12 on day 3 after infection or 10 mg/kg oseltamivir orally by gavage for 5 days from day 3 after infection onward. The control group was injected with PBS after infection. Mice were monitored daily for 14 days for survival and body weight loss
Discussion
The use of mAbs for the screening of mutant escape strains can be employed to locate antigenic sites on influenza virus HA [15, 16, 19, 20, 36]. Since the A(H1N1)pdm09 virus pandemic, studies have used mAbs to determine HA antigenic epitopes, and analysis of results using mouse mAbs has demonstrated that most antibodies could recognize antigenic epitopes in Sa and Sb. A study by Krause [40] and Manicassamy [41] confirmed that three amino acids, K157, G158 and K166, in the Sa antigenic site were related to HA antigenicity. In a recent study, Rudneva et al. prepared five mAbs and identified the recognition sites, amino acids N129, K156, G158, and N159, present in the Sa antigenic site, and D190 in the Sb antigenic site [42]. In human studies carried out to identify binding epitopes, human mAbs demonstrated that Sa was an immunodominant antigenic site of A(H1N1)pdm09 virus [40, 41]. In comparison, the eight mAbs prepared in this study recognized a greater diversity of epitopes, which were located within Sa, Sb, Ca1 and Ca2, probably due to the use of a prime-boost strategy, (two DNA immunizations with one boost of inactivated vaccine). The prime-boost immunization program in animals can improve the immune response levels to antigen [43–45], or even induce neutralizing antibodies to the HA2 stalk region [46], so we speculated that a prime-boost protocol may provide a greater stimulus opportunity for the less exposed regions of HA. In addition, the majority of mAbs recognized the Sb and Ca2 antigenic sites (5/8), and only one mAb recognized the Sa antigenic site. The results of this study were slightly different from those of previous studies, indicating that further studies on the A(H1N1)pdm09 HA immunodominant antigenic sites are required.
Virus mutations in the antigenic sites may result in decreasing neutralizing antibody titers, or even antigenic drift. In previous studies, Strengell et al. found that mutations of N125D and/or N156K (N129D or N159K numbering according to Protein Data Bank (PDB) ID code 3lzg) in the A(H1N1)pdm09 HA antigenic site could reduce the virus-neutralizing capability of antiserum [47]. Here, we analyzed the HAI activity of the mAbs with the mutant escape strains and found that the titers of all eight mAbs that inhibited the mAb3 escape mutant (G158E) decreased significantly. Thus, we speculated that the mAb3 escape mutation might lead to antigenic drift of the virus. In addition, the binding of mAbs to Ca1 and Ca2 antigenic epitopes was less likely to be affected by mutation of the Sa and Sb antigenic epitopes, although mutations of Sa or Sb had a significant mutual influence, similar to what has been reported previously for Ca1 or Ca2 sites [16, 42]. An explanation for this could be that the antigenic mutations resulted in conformational changes that not only affected the binding of mAbs to their recognized antigenic epitopes but also influenced the interaction of the adjacent antigenic epitopes and their antibodies.
We analyzed the mutations present in mAbs-induced escape mutants in a field strain and found that the incidence of mAb-selected mutations in field strains is relatively low. The G158E mutation, which was recognized by mAb3, was previously reported to be present in the viral RNA quasispecies extracted from lung tissue of an A(H1N1)pdm09 virus-infected patient [48] and was observed in a human mAb escape mutant [38]. However, after 2011, the G158E mutation was not found in field strains. Similar observations were noted for the S188 N mutation (escape mutation selected by mAb1 or mAb10) [49]. Why the majority of mAb-selected mutations did not spread through the populations could be that the mutation in HA had a negative impact on the virus. Indeed, previous studies have shown that an HA mutation in an H5 or H9 mouse-adapted mAb escape strain could reduce the virulence of the virus in mice [50–52]. In addition, Rudneva et al. analyzed the binding capacity of A(H1N1)pdm09 mAb escape strains to the receptor and found that most of the mutations that occurred in HA caused a reduction in receptor-binding capacity (both alpha-2-3- and alpha 2-6-sialic receptors) [42]. Thus, we can conclude that mutations occurring in antigenic sites of A(H1N1)pdm09 virus affected the prevalence of the mutated virus.
Passive immunization may provide an alternative for the treatment of acute infectious diseases. In this study, we prepared eight mAbs that recognized Sa, Sb, Ca1 and Ca2 antigenic sites. Animal experiments showed that the therapeutic effects of mAb treatment in mice may differ, as treatment with mAb12, which recognizes Q192 within the Sb antigenic site, showed the most prominent effects. MAb12 treatment provided complete protection of mice against infection and reduced weight loss after infection. Even low doses could provide full protection of mice. MAb2, which recognizes K242 (adjacent to Sa and Ca1, an unique epitope not identified previously), completely protected infected mice at high doses, although its effects on weight loss were not as potent as mAb12 treatment. MAbs that recognized the other antigenic sites, including Sb, Ca1 and Ca2, only provided partial protection, while mAb7, which recognizes K145 of Ca2, was unable to provide any protection for infected mice. Our results may be helpful in the design of effective vaccines such as epitope-based peptide vaccines. In addition, we compared the in vivo ability of mAb12 to prevent or treat viral infections in mice and found that it had greater efficacy for prophylaxis compared with treatment, demonstrating reduced clinical symptoms and weight loss in treated mice. These results suggest the importance of passive immunization before a virus epidemic, especially in high-risk groups such as children, the elderly, people who are immune deficient, or those with occupational exposure. Since the use of mAbs against influenza virus is in the process of receiving approval for commercial use [53], it still needs to be determined through clinical trials whether passive immunization should be widely applied in high-risk groups or rather as an individual approach. However, another clinically used mAb can provide a reference for us: palivizumab. Palivizumab is a humanized mAb that is used for prevention of respiratory syncytial virus (RSV) infection in children at high risk for severe disease [54]. Clinical investigation has revealed that children who received palivizumab at monthly intervals had 55 % fewer RSV-related hospitalizations than those who did not [54]. Although the mAb against RSV is the only commercial mAb approved for prevention of a viral disease, combining the course of prophylactic administration of palivizumab in clinical use, our results in the animal model strengthen the concept of prevention of viral infection through passive immunization.
The neuraminidase inhibitor oseltamivir is a clinical anti-flu drug, which is effective for both influenza virus type A and B. However, oseltamivir-resistant virus strains have been observed in clinical situations that affect the antiviral effects of the drug [55]. Therefore, it is necessary to develop new antiviral drugs, and as such, mAbs may be considered an ideal alternative. A previous study comparing the effects of oseltamivir with human mAb CR6261 in mice for the prevention and treatment of H5N1 or H1N1 infections showed that CR6261 was more potent than oseltamivir [56]. In the current study, one single injection of mAb12 was more effective in the prevention and treatment of viral infections in mice when compared with a continuous administration of oseltamivir over 5 days, confirming the previous study. Although no clinical comparisons have been made between mAbs and drugs in controlling influenza virus infections, passive immunization with convalescent serum could control viral infection better than oseltamivir treatment. Passive immune convalescent serum treatment for patients with severe infection rapidly reduced the viral load and eventually helped the patients recover, in contrast to the continuous administration of oseltamivir, which did not reduce the viral load [24]. This may be due to delayed oseltamivir treatment, since it has been reported that oseltamivir suppresses viral load more effectively when given early [57, 58]. In conclusion, these findings suggest that antibodies may provide better prevention and treatment against viral infections than anti-flu drugs.
Although mouse mAbs have demonstrated good antiviral effects in animal models, they should be humanized to prevent immunity to the mouse mAbs and the production of human anti-mouse antibodies in patients. One major disadvantage of mAb treatment is that influenza virus can produce escape strains through mutation, resulting in the loss of recognition of antigenic sites and thus the antiviral effect of antibodies. Therefore, it will be important to develop a humanized mAb with broad-spectrum recognition of antigenic sites for the treatment of the 2009 pdmH1N1 flu virus.
References
Swine influenza A (H1N1) infection in two children—Southern California. Morb Mortal Wkly Rep 2009 March–April 58
World Health Organization (2010) Pandemic (H1N1) 2009, update 112. World Health Organization, Geneva, Switzerland
Garten RJ, Davis CT, Russell CA, Shu B, Lindstrom S, Balish A, Sessions WM, Xu X, Skepner E, Deyde V, Okomo-Adhiambo M, Gubareva L, Barnes J, Smith CB, Emery SL, Hillman MJ, Rivailler P, Smagala J, de Graaf M, Burke DF, Fouchier RA, Pappas C, Alpuche-Aranda CM, Lopez-Gatell H, Olivera H, Lopez I, Myers CA, Faix D, Blair PJ, Yu C, Keene KM, Dotson PD Jr, Boxrud D, Sambol AR, Abid SH, St George K, Bannerman T, Moore AL, Stringer DJ, Blevins P, Demmler-Harrison GJ, Ginsberg M, Kriner P, Waterman S, Smole S, Guevara HF, Belongia EA, Clark PA, Beatrice ST, Donis R, Katz J, Finelli L, Bridges CB, Shaw M, Jernigan DB, Uyeki TM, Smith DJ, Klimov AI, Cox NJ (2009) Antigenic and genetic characteristics of swine-origin 2009 A(H1N1) influenza viruses circulating in humans. Science 325(5937):197–201
Smith GJ, Vijaykrishna D, Bahl J, Lycett SJ, Worobey M, Pybus OG, Ma SK, Cheung CL, Raghwani J, Bhatt S, Peiris JS, Guan Y, Rambaut A (2009) Origins and evolutionary genomics of the 2009 swine-origin H1N1 influenza A epidemic. Nature 459(7250):1122–1125
Dawood FS, Jain S, Finelli L, Shaw MW, Lindstrom S, Garten RJ, Gubareva LV, Xu X, Bridges CB, Uyeki TM (2009) Emergence of a novel swine-origin influenza A (H1N1) virus in humans. N Engl J Med 360(25):2605–2615
Nelson M, Spiro D, Wentworth D, Beck E, Fan J, Ghedin E, Halpin R, Bera J, Hine E, Proudfoot K, Stockwell T, Lin X, Griesemer S, Kumar S, Bose M, Viboud C, Holmes E, Henrickson K (2009) The early diversification of influenza A/H1N1pdm. PLoS Curr 1:RRN1126
Makkoch J, Suwannakarn K, Payungporn S, Prachayangprecha S, Cheiocharnsin T, Linsuwanon P, Theamboonlers A, Poovorawan Y (2012) Whole Genome Characterization, Phylogenetic and Genome Signature Analysis of Human Pandemic H1N1 Virus in Thailand, 2009–2012. PLoS One 7(12):e51275
Potdar VA, Chadha MS, Jadhav SM, Mullick J, Cherian SS, Mishra AC (2010) Genetic characterization of the influenza A pandemic (H1N1) 2009 virus isolates from India. PLoS One 5(3):e9693
Barrero PR, Viegas M, Valinotto LE, Mistchenko AS (2011) Genetic and phylogenetic analyses of influenza A H1N1pdm virus in Buenos Aires, Argentina. J Virol 85(2):1058–1066
Ilyicheva T, Susloparov I, Durymanov A, Romanovskaya A, Sharshov K, Kurskaya O, Ignashkina M, Shestopalov A (2011) Influenza A/H1N1pdm virus in Russian Asia in 2009–2010. Infect Genet Evol 11(8):2107–2112
Igarashi M, Ito K, Yoshida R, Tomabechi D, Kida H, Takada A (2010) Predicting the antigenic structure of the pandemic (H1N1) 2009 influenza virus hemagglutinin. PLoS One 5(1):e8553
Tong S, Li Y, Rivailler P, Conrardy C, Castillo DA, Chen LM, Recuenco S, Ellison JA, Davis CT, York IA, Turmelle AS, Moran D, Rogers S, Shi M, Tao Y, Weil MR, Tang K, Rowe LA, Sammons S, Xu X, Frace M, Lindblade KA, Cox NJ, Anderson LJ, Rupprecht CE, Donis RO (2012) A distinct lineage of influenza A virus from bats. Proc Natl Acad Sci 109(11):4269–4274
Fouchier RA, Munster V, Wallensten A, Bestebroer TM, Herfst S, Smith D, Rimmelzwaan GF, Olsen B, Osterhaus AD (2005) Characterization of a novel influenza A virus hemagglutinin subtype (H16) obtained from black-headed gulls. J Virol 79(5):2814–2822
Skehel JJ, Wiley DC (2000) Receptor binding and membrane fusion in virus entry: the influenza hemagglutinin. Annu Rev Biochem 69:531–569
Caton AJ, Brownlee GG, Yewdell JW, Gerhard W (1982) The antigenic structure of the influenza virus A/PR/8/34 hemagglutinin (H1 subtype). Cell 31(2 Pt 1):417–427
Kaverin NV, Rudneva IA, Govorkova EA, Timofeeva TA, Shilov AA, Kochergin-Nikitsky KS, Krylov PS, Webster RG (2007) Epitope mapping of the hemagglutinin molecule of a highly pathogenic H5N1 influenza virus by using monoclonal antibodies. J Virol 81(23):12911–12917
Tsuchiya E, Sugawara K, Hongo S, Matsuzaki Y, Muraki Y, Li ZN, Nakamura K (2001) Antigenic structure of the haemagglutinin of human influenza A/H2N2 virus. J Gen Virol 82(Pt 10):2475–2484
Underwood PA (1984) An antigenic map of the haemagglutinin of the influenza Hong Kong subtype (H3N2), constructed using mouse monoclonal antibodies. Mol Immunol 21(7):663–671
Brownlee GG, Fodor E (2001) The predicted antigenicity of the haemagglutinin of the 1918 Spanish influenza pandemic suggests an avian origin. Philos Trans R Soc Lond B Biol Sci 356(1416):1871–1876
Luoh SM, McGregor MW, Hinshaw VS (1992) Hemagglutinin mutations related to antigenic variation in H1 swine influenza viruses. J Virol 66(2):1066–1073
Marasco WA, Sui J (2007) The growth and potential of human antiviral monoclonal antibody therapeutics. Nat Biotechnol 25(12):1421–1434
Sawyer LA (2000) Antibodies for the prevention and treatment of viral diseases. Antiviral Res 47(2):57–77
Luke TC, Kilbane EM, Jackson JL, Hoffman SL (2006) Meta-analysis: convalescent blood products for Spanish influenza pneumonia: a future H5N1 treatment? Ann Intern Med 145(8):599–609
Zhou B, Zhong N, Guan Y (2007) Treatment with convalescent plasma for influenza A (H5N1) infection. N Engl J Med 357(14):1450–1451
Hung IF, To KK, Lee CK, Lee KL, Chan K, Yan WW, Liu R, Watt CL, Chan WM, Lai KY, Koo CK, Buckley T, Chow FL, Wong KK, Chan HS, Ching CK, Tang BS, Lau CC, Li IW, Liu SH, Chan KH, Lin CK, Yuen KY (2011) Convalescent plasma treatment reduced mortality in patients with severe pandemic influenza A (H1N1) 2009 virus infection. Clin Infect Dis 52(4):447–456
Chen Y, Qin K, Wu WL, Li G, Zhang J, Du H, Ng MH, Shih JW, Peiris JS, Guan Y, Chen H, Xia N (2009) Broad cross-protection against H5N1 avian influenza virus infection by means of monoclonal antibodies that map to conserved viral epitopes. J Infect Dis 199(1):49–58
Renegar KB, Small PA Jr, Boykins LG, Wright PF (2004) Role of IgA versus IgG in the control of influenza viral infection in the murine respiratory tract. J Immunol 173(3):1978–1986
Smirnov YA, Lipatov AS, Gitelman AK, Claas EC, Osterhaus AD (2000) Prevention and treatment of bronchopneumonia in mice caused by mouse-adapted variant of avian H5N2 influenza A virus using monoclonal antibody against conserved epitope in the HA stem region. Arch Virol 145(8):1733–1741
Palladino G, Mozdzanowska K, Washko G, Gerhard W (1995) Virus-neutralizing antibodies of immunoglobulin G (IgG) but not of IgM or IgA isotypes can cure influenza virus pneumonia in SCID mice. J Virol 69(4):2075–2081
Clark TW, Pareek M, Hoschler K, Dillon H, Nicholson KG, Groth N, Stephenson I (2009) Trial of 2009 influenza A (H1N1) monovalent MF59-adjuvanted vaccine. N Engl J Med 361(25):2424–2435
Reed LJ, Muench H (1938) A simple method of estimating fifty per cent endpoints. Am J Hyg 27:493–497
Chen Q, Huang S, Chen J, Zhang S, Chen Z (2013) NA proteins of influenza A viruses H1N1/2009, H5N1, and H9N2 show differential effects on infection initiation, virus release, and cell-cell fusion. PLoS one 8(1):e54334
Chen J, Fang F, Li X, Chang H, Chen Z (2005) Protection against influenza virus infection in BALB/c mice immunized with a single dose of neuraminidase-expressing DNAs by electroporation. Vaccine 23(34):4322–4328
Tabynov K, Kydyrbayev Z, Sansyzbay A, Khairullin B, Ryskeldinova S, Assanzhanova N, Kozhamkulov Y, Inkarbekov D (2012) Immunogenic and protective properties of the first kazakhstan vaccine against pandemic influenza A (H1N1) pdm09 in Ferrets. Virol Sin 27(6):344–351
Mozdzanowska K, Furchner M, Washko G, Mozdzanowski J, Gerhard W (1997) A pulmonary influenza virus infection in SCID mice can be cured by treatment with hemagglutinin-specific antibodies that display very low virus-neutralizing activity in vitro. J Virol 71(6):4347–4355
Webster RG, Laver WG (1980) Determination of the number of nonoverlapping antigenic areas on Hong Kong (H3N2) influenza virus hemagglutinin with monoclonal antibodies and the selection of variants with potential epidemiological significance. Virology 104(1):139–148
Chen J, Fang F, Yang Z, Liu X, Zhang H, Zhang Z, Zhang X, Chen Z (2009) Characterization of highly pathogenic H5N1 avian influenza viruses isolated from poultry markets in central China. Virus Res 146(1–2):19–28
O’Donnell CD, Vogel L, Wright A, Das SR, Wrammert J, Li GM, McCausland M, Zheng NY, Yewdell JW, Ahmed R, Wilson PC, Subbarao K (2012) Antibody pressure by a human monoclonal antibody targeting the 2009 pandemic H1N1 virus hemagglutinin drives the emergence of a virus with increased virulence in mice. mBio 3(3):e00120–12
Hensley SE, Das SR, Bailey AL, Schmidt LM, Hickman HD, Jayaraman A, Viswanathan K, Raman R, Sasisekharan R, Bennink JR, Yewdell JW (2009) Hemagglutinin receptor binding avidity drives influenza A virus antigenic drift. Science 326(5953):734–736
Krause JC, Tsibane T, Tumpey TM, Huffman CJ, Basler CF, Crowe JE (2011) A broadly neutralizing human monoclonal antibody that recognizes a conserved, novel epitope on the globular head of the influenza H1N1 virus hemagglutinin. J Virol 85(20):10905–10908
Manicassamy B, Medina RA, Hai R, Tsibane T, Stertz S, Nistal-Villan E, Palese P, Basler CF, Garcia-Sastre A (2010) Protection of mice against lethal challenge with 2009 H1N1 influenza A virus by 1918-like and classical swine H1N1 based vaccines. PLoS Pathog 6(1):e1000745
Rudneva I, Ignatieva A, Timofeeva T, Shilov A, Kushch A, Masalova O, Klimova R, Bovin N, Mochalova L, Kaverin N (2012) Escape mutants of pandemic influenza A/H1N1 2009 virus: variations in antigenic specificity and receptor affinity of the hemagglutinin. Virus Res 166(1–2):61–67
Suguitan AL Jr, Cheng X, Wang W, Wang S, Jin H, Lu S (2011) Influenza H5 hemagglutinin DNA primes the antibody response elicited by the live attenuated influenza A/Vietnam/1203/2004 vaccine in ferrets. PLoS One 6(7):e21942
Wang S, Parker C, Taaffe J, Solorzano A, Garcia-Sastre A, Lu S (2008) Heterologous HA DNA vaccine prime-inactivated influenza vaccine boost is more effective than using DNA or inactivated vaccine alone in eliciting antibody responses against H1 or H3 serotype influenza viruses. Vaccine 26(29–30):3626–3633
Wei CJ, Boyington JC, McTamney PM, Kong WP, Pearce MB, Xu L, Andersen H, Rao S, Tumpey TM, Yang ZY, Nabel GJ (2010) Induction of broadly neutralizing H1N1 influenza antibodies by vaccination. Science 329(5995):1060–1064
Ledgerwood JE, Wei CJ, Hu Z, Gordon IJ, Enama ME, Hendel CS, McTamney PM, Pearce MB, Yassine HM, Boyington JC, Bailer R, Tumpey TM, Koup RA, Mascola JR, Nabel GJ, Graham BS (2011) DNA priming and influenza vaccine immunogenicity: two phase 1 open label randomised clinical trials. Lancet Infect Dis 11(12):916–924
Strengell M, Ikonen N, Ziegler T, Julkunen I (2011) Minor changes in the hemagglutinin of influenza A(H1N1)2009 virus alter its antigenic properties. PLoS One 6(10):e25848
Kuroda M, Katano H, Nakajima N, Tobiume M, Ainai A, Sekizuka T, Hasegawa H, Tashiro M, Sasaki Y, Arakawa Y, Hata S, Watanabe M, Sata T (2010) Characterization of quasispecies of pandemic 2009 influenza A virus (A/H1N1/2009) by de novo sequencing using a next-generation DNA sequencer. PLoS One 5(4):e10256
Fatal H1N1 RBD Change S188N in Hunan China? (2010) Recombinomics Commentary. http://www.recombinomics.com/News/01031002/S188N_Hunan.html
Kaverin NV, Rudneva IA, Ilyushina NA, Lipatov AS, Krauss S, Webster RG (2004) Structural differences among hemagglutinins of influenza A virus subtypes are reflected in their antigenic architecture: analysis of H9 escape mutants. J Virol 78(1):240–249
Kaverin NV, Rudneva IA, Ilyushina NA, Varich NL, Lipatov AS, Smirnov YA, Govorkova EA, Gitelman AK, Lvov DK, Webster RG (2002) Structure of antigenic sites on the haemagglutinin molecule of H5 avian influenza virus and phenotypic variation of escape mutants. J Gen Virol 83(Pt 10):2497–2505
Rudneva IA, Ilyushina NA, Timofeeva TA, Webster RG, Kaverin NV (2005) Restoration of virulence of escape mutants of H5 and H9 influenza viruses by their readaptation to mice. J Gen Virol 86(Pt 10):2831–2838
Theraclone sciences initiates Phase 1 trial of TCN-032 for influenza A. http://wwwbusinesswirecom/news/home/20110921005197/en/Theraclone-Sciences-Announces-Initiation-Phase-1-Clinical
Lanari M, Vandini S, Arcuri S, Galletti S, Faldella G (2013) The use of humanized monoclonal antibodies for the prevention of respiratory syncytial virus infection. Clin Dev Immunol 2013:9
Ison MG (2011) Antivirals and resistance: influenza virus. Curr Opin Virol 1(6):563–573
Koudstaal W, Koldijk MH, Brakenhoff JP, Cornelissen LA, Weverling GJ, Friesen RH, Goudsmit J (2009) Pre- and postexposure use of human monoclonal antibody against H5N1 and H1N1 influenza virus in mice: viable alternative to oseltamivir. J Infect Dis 200(12):1870–1873
Ling LM, Chow AL, Lye DC, Tan AS, Krishnan P, Cui L, Win NN, Chan M, Lim PL, Lee CC, Leo YS (2010) Effects of early oseltamivir therapy on viral shedding in 2009 pandemic influenza A (H1N1) virus infection. Clin Infect Dis 50(7):963–969
Li IW, Hung IF, To KK, Chan KH, Wong SS, Chan JF, Cheng VC, Tsang OT, Lai ST, Lau YL, Yuen KY (2010) The natural viral load profile of patients with pandemic 2009 influenza A(H1N1) and the effect of oseltamivir treatment. Chest 137(4):759–768
Acknowledgments
This study was supported by the following research funds: National 973 Project (2010CB534005, 2010CB530301); National Natural Science Foundation of China (310000088); Foundation for Study Encouragement to Young Scientists, Chinese Academy of Sciences (KSCX2-EW-J-19) and the Ministry of Science and Technology Special Project (2013FY113500).
Author information
Authors and Affiliations
Corresponding authors
Additional information
J. Chen and B. Yan contributed equally.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Chen, J., Yan, B., Chen, Q. et al. Evaluation of neutralizing efficacy of monoclonal antibodies specific for 2009 pandemic H1N1 influenza A virus in vitro and in vivo . Arch Virol 159, 471–483 (2014). https://doi.org/10.1007/s00705-013-1852-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-013-1852-y