Abstract
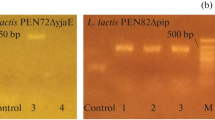
In this study, we isolated and characterized a lytic Lactococcus lactis bacteriophage from the sera of a failed fermentation. The phage was isolated and cultured in L. lactis subsp. cremoris in M17 medium. The isolated bacteriophage was characterized by multiplex PCR, pulsed-field electrophoresis, DNA restriction digestion, analysis of the N-terminal sequence of the phage major structural protein, transmission electron microscopy and sequencing and analysis of a conserved fragment of its genome. Analysis of the viral genome indicates that its genome is composed of a DNA strand of approximately 48 kb in length, and PCR and microscopy confirmed that IL-P1 belongs to the group of 936-type phages in the family Siphoviridae, which is the most abundant type of lactococcal virus in dairy products worldwide. To our knowledge, this is the first report of a virus within this family that has a presumptive genome larger than 40 kb.






Similar content being viewed by others
References
Bogosian G (2006) Control of bacteriophage in commercial microbiology and fermentation facilities. In: Calendar R (ed) The bacteriophages, Oxford University Press, New York, pp 667–673
Labrie SJ, Josephsen J, Neve H, Vogensen FK, Moineau S (2008) Morphology, genome sequence, and structural proteome of type phage P335 from Lactococcus lactis. Appl Environ Microbiol 74(15):4636–4644
McGrath S, Fitzgerald GF, van Sinderen D (2004) The impact of bacteriophage genomics. Curr Opin Biotechnol 15(2):94–99
Raiski A, Belyasova N (2009) Biodiversity of Lactococcus lactis bacteriophages in the Republic of Belarus. Int J Food Microbiol 130(1):1–5
Dinsmore PK, O’Sullivan DJ, Klaenhammer TR (1998) A leucine repeat motif in AbiA is required for resistance of Lactococcus lactis to phages representing three species. Gene 212(1):5–11
Madera C, Monjardin C, Suarez JE (2004) Milk contamination and resistance to processing conditions determine the fate of Lactococcus lactis bacteriophages in dairies. Appl Environ Microbiol 70(12):7365–7371
Josephsen J, Neve H: Bacteriophages and lactic acid bacteria (1998) In: Salminen S, Wright V, Ouwehand A (eds) Lactic Acid bacteria: microbiological and functional aspects, CRC Press, New York, pp 385–436
Oliveira MR, Guimarães WV, Araújo EF, Borges AC (1995) Isolation and characterization of lactococcal bacteriophages from cheese whey. Revista de Microbiologia 26(1):01–05
Binetti AG, Capra ML, Alvarez MA, Reinheimer JA (2008) PCR method for detection and identification of Lactobacillus casei/paracasei bacteriophages in dairy products. Int J Food Microbiol 124(2):147–153
del Rio B, Binetti AG, Martin MC, Fernandez M, Magadan AH, Alvarez MA (2007) Multiplex PCR for the detection and identification of dairy bacteriophages in milk. Food Microbiol 24(1):75–81
Dupuis ME, Moineau S (2010) Genome organization and characterization of the virulent lactococcal phage 1358 and its similarities to Listeria phages. Appl Environ Microbiol 76(5):1623–1632
Hejnowicz MS, Golebiewski M, Bardowski J (2009) Analysis of the complete genome sequence of the lactococcal bacteriophage bIBB29. Int J Food Microbiol 131(1):52–61
Brussow H (2001) Phages of dairy bacteria. Annu Rev Microbiol 55:283–303
Mcgrath S, Van Sinderen D, Fitzgerald GF (2002) Bacteriophage-derived genetic tools for use in lactic acid bacteria. Int Dairy J 12(1):3–15
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor, NY, Cold Spring Harbor Laboratory
Hull RR (1977) Methods for monitoring bacteriophage in cheese factories. Aust J Dairy Technol 6:63–64
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227(5259):680–685
Deveau H, Labrie SJ, Chopin MC, Moineau S (2006) Biodiversity and classification of lactococcal phages. Appl Environ Microbiol 72(6):4338–4346
Jarvis AW, Fitzgerald GF, Mata M, Mercenier A, Neve H, Powell IB, Ronda C, Saxelin M, Teuber M (1991) Species and type phages of lactococcal bacteriophages. Intervirology 32(1):2–9
Braun V, Hertwig D, Neve H, Geis A, Teuber M (1989) Taxonomic differentiation of bacteriophages of Lactococcus lactis by electron microscopy, DNA-DNA hybridization, and protein profiles. J Gen Microbiol 135:2551–2560
Crutz-Le Coq AM, Cantele F, Lanzavecchia S, Marco S (2006) Insights into structural proteins of 936-type virulent lactococcal bacteriophages. Arch Virol 151(6):1039–1053
Lubbers MW, Waterfield NR, Beresford TP, Le Page RW, Jarvis AW (1995) Sequencing and analysis of the prolate-headed lactococcal bacteriophage c2 genome and identification of the structural genes. Appl Environ Microbiol 61(12):4348–4356
Haggaard-Ljungquist E, Jacobsen E, Rishovd S, Six EW, Nilssen O, Sunshine MG, Lindqvist BH, Kim K-J, Barreiro V, Koonin EV, Calendar R (1995) Bacteriophage P2: genes involved in baseplate assembly. Virology 213:109–121
Kageyama Y, Murayama M, Onodera T, Yamada S, Fukada H, Kudou M, Tsumoto K, Toyama Y, Kado S, Kubota K, Takeda S (2009) Observation of the membrane binding activity and domain structure of gpV, which comprises the tail spike of bacteriophage P2. Biochemistry 48:10129–10135
Yamashita E, Nakagawa A, Takahashi J, Tsunoda K, Yamada S, Takeda S (2011) The host-binding domain of the P2 phage tail spike reveals a trimeric iron-binding structure. Acta Crystallogr 67:837–841
Crutz-Le Coq AM, Cesselin B, Commissaire J, Anba J (2002) Sequence analysis of the lactococcal bacteriophage bIL170: insights into structural proteins and HNH endonucleases in dairy phages. Microbiology 148(4):985–1001
Chandry PS, Moore SC, Boyce JD, Davidson BE, Hillier AJ (1977) Analysis of the DNA sequence, gene expression, origin of replication and modular structure of the Lactococcus lactis lytic bacteriophage sk1. Mol Microbiol 26(1):49–64
Loof M, Lembke J, Teuber M (1983) Characterization of the genome of the Streptococcus lactis subsp. diacetylactis bacteriophage P008 wide-spread in German cheese factories. Syst Appl Microbiol 4:413–423
Acknowledgments
We would like to thank the Nucleus of Microscopy and Microanalysis (NMM) and the Nucleus of Biomolecules (NuBio) of the Federal University of Viçosa, for providing materials and technical support. This study was supported by grants from the Fundação de Amparo à Pesquisa do Estado de Minas Gerais (FAPEMIG) and a Master fellowship from Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES) and Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) to MRE. The funders had no role in the study design, data collection, analysis, decision to publish, or preparation of this manuscript.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Eller, M.R., Dias, R.S., De Moraes, C.A. et al. Molecular characterization of a new lytic bacteriophage isolated from cheese whey. Arch Virol 157, 2265–2272 (2012). https://doi.org/10.1007/s00705-012-1432-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-012-1432-6