Abstract
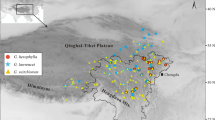
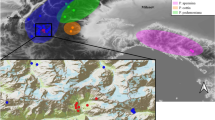
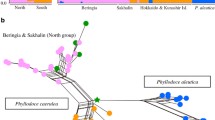
The western Balkans represents an area of significant topographic and environmental heterogeneity, harbouring high species and intraspecific diversity. Similar genetic and distributional splits observed in unrelated species have suggested some common features in their glacial response and biogeographic history. Here, we studied the western Balkan endemic Alyssum austrodalmaticum with the aim to explore and understand its intraspecific structure and processes that shaped the present patterns. We employed data from AFLPs, two low-copy nuclear genes, genome size, morphometrics and species distribution modelling. Four genetic lineages were identified within the species, which were geographically structured and showed cytotype-specific patterns. The observed phylogeographic structure is congruent with the predicted species distribution during the Last Glacial Maximum. Two allopatric diploid lineages (referred to as N2x and S2x) reflect glacial range disruption and survival in distinct refugia inferred in north-western (Istria, Kvarner) and south-eastern Adriatic areas (northern Adriatic palaeo-coastline and southern Dinarides). AFLP analyses with in silico-generated polyploid genotypes and nuclear genes proved that the two tetraploid lineages (C4x, S4x) were genetic allopolyploids and arose independently. The central tetraploids (C4x) originated through secondary contacts of the two diploid lineages. The origin of the southern tetraploids (S4x) is somewhat ambiguous. Apart from the southern diploids (S2x), the involvement, either direct or through later introgression, of the central tetraploids (C4x) or even other Balkan relatives is possible. Our study highlights the role of glacial range dynamics and secondary contacts, triggering introgression and polyploid evolution, in the formation of genetic diversity and intraspecific patterns in the western Balkans.








Similar content being viewed by others
References
Akaike H (1974) A new look at the statistical model identification. IEEE Trans Automatic Control 19:716–723. https://doi.org/10.1109/TAC.1974.1100705
Araújo MB, New M (2007) Ensemble forecasting of species distributions. Trends Ecol Evol 22:42–47. https://doi.org/10.1016/j.tree.2006.09.010
Baduel P, Bray S, Vallejo-Marin M, Kolář F, Yant L (2018) The “polyploid hop”: shifting challenges and opportunities over the evolutionary lifespan of genome duplications. Frontiers Ecol Evol 6:117. https://doi.org/10.3389/fevo.2018.00117
Bardy KE, Albach DC, Schneeweiss GM, Fischer MA, Schönswetter P (2010) Disentangling phylogeography, polyploid evolution and taxonomy of a woodland herb (Veronica chamaedrys group, Plantaginaceae s.l.) in southeastern Europe. Molec Phylogen Evol 57:771–786. https://doi.org/10.1016/j.ympev.2010.06.025
Benac Č, Juračić M (1998) Geomorphological indicators of sea level changes during upper Pleistocene (Würm) and Holocene in the Kvarner region (NE Adriatic sea). Acta Geo Croat 33:27–45
Bivand RS, Pebesma EJ, Gomez-Rubio V (2013) Applied spatial data analysis with R, 2nd edn. Springer, New York. https://doi.org/10.1007/978-1-4614-7618-4
Bogdanović S, Brullo S, Rešetnik I, Satovic Z, Liber Z (2014) Campanula teutana, a new isophyllous Campanula (Campanulaceae) from the Adriatic region. Phytotaxa 162:001–017. https://doi.org/10.11646/phytotaxa.162.1.1
Bogdanović S, Rešetnik I, Jeričević M, Jeričević N, Brullo S (2019) Molecular and morphological survey on Campanula cremnophila (Campanulaceae), a new isophyllous species from Croatia. Pl Syst Evol 305:687–703. https://doi.org/10.1007/s00606-019-01599-x
Brassac J, Jakob SS, Blattner FR (2012) Progenitor-derivative relationships of Hordeum polyploids (Poaceae, Triticeae) inferred from sequences of TOPO6, a nuclear low-copy gene region. PLoS ONE 7:e33808. https://doi.org/10.1371/journal.pone.0033808
Breiner FT, Guisan A, Bergamini A, Nobis MP (2015) Overcoming limitations of modelling rare species by using ensembles of small models. Meth Ecol Evol 6:1210–1218. https://doi.org/10.1111/2041-210X.12403
Brier GW (1950) Verification of forecasts expressed in terms of probability. Monthly Weather Rev 78:1–3. https://doi.org/10.1175/1520-0493(1950)078%3c0001:VOFEIT%3e2.0.CO;2
Caković D, Stešević D, Schönswetter P, Frajman B (2015) How many taxa? Spatiotemporal evolution and taxonomy of Amphoricarpos (Asteraceae, Carduoideae) on the Balkan Peninsula. Organisms Diversity Evol 15:429–445. https://doi.org/10.1007/s13127-015-0218-6
Caković D, Stešević D, Schönswetter P, Frajman B (2018) Long neglected diversity in the Accursed Mountains of northern Albania: Cerastium hekuravense is genetically and morphologically divergent from C. dinaricum. Pl Syst Evol 304:57–69. https://doi.org/10.1007/s00606-017-1448-1
Clement M, Posada D, Crandall KA (2000) TCS: a computer program to estimate gene genealogies. Molec Ecol 9:1657–1660. https://doi.org/10.1046/j.1365-294x.2000.01020.x
Correggiari A, Roveri M, Trincardi F (1996) Late Pleistocene and Holocene evolution of the North Adriatic Sea. Il Quaternario 9:697–704
Darriba D, Taboada GL, Doallo R, Posada D (2012) jModelTest 2: more models, new heuristics and parallel computing. Nature Meth 9:772. https://doi.org/10.1038/nmeth.2109
Degnan JH, Rosenberg NA (2009) Gene tree discordance, phylogenetic inference and the multispecies coalescent. Trends Ecol Evol 24:332–340. https://doi.org/10.1016/j.tree.2009.01.009
Díaz-Pérez A, López-Álvarez D, Sancho R, Catalán P (2018) Reconstructing the origins and the biogeography of species’ genomes in the highly reticulate allopolyploid-rich model grass genus Brachypodium using minimum evolution, coalescence and maximum likelihood approaches. Molec Phylogen Evol 127:256–271. https://doi.org/10.1016/j.ympev.2018.06.003
Doležel J, Sgorbati S, Lucretti S (1992) Comparison of three DNA fluorochromes for flow cytometric estimation of nuclear DNA content in plants. Physiol Pl (Copenhagen) 85:625–631. https://doi.org/10.1111/j.1399-3054.1992.tb04764.x
Dormann CF, Elith J, Bacher S, Buchmann C, Carl G, Carré G, Marquéz JRG, Gruber B, Lafourcade B, Leitão PJ (2013) Collinearity: a review of methods to deal with it and a simulation study evaluating their performance. Ecography 36:27–46. https://doi.org/10.1111/j.1600-0587.2012.07348.x
Drummond A, Rambaut A (2007) BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol Biol 7:214. https://doi.org/10.1186/1471-2148-7-214
Fick SE, Hijmans RJ (2017) WorldClim 2: new 1-km spatial resolution climate surfaces for global land areas. Int J Climatol 37:4302–4315. https://doi.org/10.1002/joc.5086
Frajman B, Pachschwöll C, Schönswetter P (2014) Contributions to the knowledge of the flora of the Dinarides (Balkan Peninsula). Phyton (Horn) 54:27–46. https://doi.org/10.12905/0380.phyton54(1)2014-0027
Frajman B, Rešetnik I, Niketić M, Ehrendorfer F, Schönswetter P (2016) Patterns of rapid diversification in heteroploid Knautia sect. Trichera (Caprifoliaceae, Dipsacoideae), one of the most intricate taxa of the European flora. BMC Evol Biol 16:204. https://doi.org/10.1186/s12862-016-0773-2
Gent PR, Danabasoglu G, Donner LJ, Holland MM, Hunke EC, Jayne SR, Lawrence DM, Neale RB, Rasch PJ, Vertenstein M (2011) The community climate system model version 4. J Climate 24:4973–4991. https://doi.org/10.1175/2011JCLI4083.1
Glasnović P, Temunović M, Lakušić D, Rakić T, Brečko Grubar V, Surina B (2018) Understanding biogeographical patterns in the western Balkan Peninsula using environmental niche modelling and geostatistics in polymorphic Edraianthus tenuifolius. AoB Plants 10:ply064. https://doi.org/10.1093/aobpla/ply064
Gómez A, Lunt DH (2007) Refugia within refugia: patterns of phylogeographic concordance in the Iberian Peninsula. In: Weiss S, Ferrand N (eds) Phylogeography of Southern European refugia. Springer, Berlin, pp 155–188. https://doi.org/10.1007/1-4020-4904-8
Gömöry D, Zhelev P, Brus R (2020) The Balkans: a genetic hotspot but not a universal colonization source for trees. Pl Syst Evol 306:5. https://doi.org/10.1007/s00606-020-01647-x
Gotelli NJ, Stanton-Geddes J (2015) Climate change, genetic markers and species distribution modelling. J Biogeogr 42:1577–1585. https://doi.org/10.1111/jbi.12562
Greilhuber J, Doležel J, Lysak M, Bennett MD (2004) The origin, evolution and proposed stabilization of the terms ‘genome size’ and ‘C-value’ to describe nuclear DNA contents. Ann Bot (Oxford) 95:255–260. https://doi.org/10.1093/aob/mci019
Griffiths HI, Kryštufek B, Reed JM (eds) (2004) Balkan biodiversity: pattern and process in the European hotspot. Kluwer, Dordrecht. https://doi.org/10.1007/978-1-4020-2854-0
Guo Y-P, Wang S-Z, Vogl C, Ehrendorfer F (2012) Nuclear and plastid haplotypes suggest rapid diploid and polyploid speciation in the N Hemisphere Achillea millefolium complex (Asteraceae). BMC Evol Biol 12:2. https://doi.org/10.1186/1471-2148-12-2
Hammer Ø, Harper DAT, Ryan PD (2001) PAST: paleontological statistics software package for education and data analysis. Palaeontol Electronica 4(1):4
Hewitt MG (1999) Post-glacial re-colonization of European biota. Biol J Linn Soc 68:87–112. https://doi.org/10.1006/bijl.1999.0332
Hewitt GM (2011) Mediterranean peninsulas: the evolution of hotspots. In: Zachos FE, Habel JC (eds) Biodiversity hotspots. Springer, Berlin, pp 123–147. https://doi.org/10.1007/978-3-642-20992-5
Hijmans RJ (2019) Raster: geographic data analysis and modeling. R package version 3.0-2
Huelsenbeck JP, Ronquist F (2001) MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 17:754–755. https://doi.org/10.1093/bioinformatics/17.8.754
Hughes PD, Woodward JC, van Calsteren PC, Thomas LE, Adamson KR (2010) Pleistocene ice caps on the coastal mountains of the Adriatic Sea. Quatern Sci Rev 29:3690–3708. https://doi.org/10.1016/j.quascirev.2010.06.032
Jakob SS, Blattner FR (2010) Two extinct diploid progenitors were involved in allopolyploid formation in the Hordeum murinum (Poaceae: Triticeae) taxon complex. Molec Phylogen Evol 55:650–659. https://doi.org/10.1016/j.ympev.2009.10.021
Janković I, Šatović Z, Liber Z, Kuzmanović N, Radosavljević I, Lakušić D (2016) Genetic diversity and morphological variability in the Balkan endemic Campanula secundiflora s.l. (Campanulaceae). Bot J Linn Soc 180:64–88. https://doi.org/10.1111/boj.12359
Jarvis A, Reuter HI, Nelson A, Guevara E (2008) Hole-filled seamless SRTM data V4. International Centre for Tropical Agriculture (CIAT). Available at: http://srtm.csi.cgiar.org. Accessed Nov 2019
Joly S, Heenan PB, Lockhart PJ (2009) A Pleistocene inter-tribal allopolyploidization event precedes the species radiation of Pachycladon (Brassicaceae) in New Zealand. Molec Phylogen Evol 51:365–372. https://doi.org/10.1016/j.ympev.2009.02.015
Jones G, Sagitov S, Oxelman B (2013) Statistical inference of allopolyploid species networks in the presence of incomplete lineage sorting. Syst Biol 62:467–478. https://doi.org/10.1093/sysbio/syt012
Katoh K, Misawa K, Kuma K-I, Miyata T (2002) MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucl Acids Res 30:3059–3066. https://doi.org/10.1093/nar/gkf436
Klecka WR (1980) Discriminant analysis. Sage University papers, Series: Quantitative applications in the social sciences, no. 19, Beverly Hills
Koch MA, Haubold B, Mitchell-Olds T (2000) Comparative evolutionary analysis of chalcone synthase and alcohol dehydrogenase loci in Arabidopsis, Arabis, and related genera (Brassicaceae). Molec Biol Evol 17:1483–1498. https://doi.org/10.1093/oxfordjournals.molbev.a026248
Konowalik K, Wagner F, Tomasello S, Vogt R, Oberprieler C (2015) Detecting reticulate relationships among diploid Leucanthemum Mill. (Compositae, Anthemideae) taxa using multilocus species tree reconstruction methods and AFLP fingerprinting. Molec Phylogen Evol 92:308–328. https://doi.org/10.1016/j.ympev.2015.06.003
Krak K, Caklová P, Chrtek J, Fehrer J (2013) Reconstruction of phylogenetic relationships in a highly reticulate group with deep coalescence and recent speciation (Hieracium, Asteraceae). Heredity 110:138–151. https://doi.org/10.1038/hdy.2012.100
Krzanowski WJ (1990) Principles of multivariate analysis. Clarendon Press, Oxford
Kučera J, Tremetsberger K, Vojta J, Marhold K (2008) Molecular study of the Cardamine maritima group (Brassicaceae) from Balkan and Apennine Peninsulas based on amplified fragment length polymorphism (AFLP). Pl Syst Evol 275:193–207. https://doi.org/10.1007/s00606-008-0061-8
Kučera J, Marhold K, Lihová J (2010) Cardamine maritima group (Brassicaceae) in the amphi-Adriatic area: a hotspot of species diversity revealed by DNA sequences and morphological variation. Taxon 59:148–164
Kuhn M, Johnson K (2013) Applied predictive modeling, vol. 26. Springer, New York. https://doi.org/10.1007/978-1-4614-6849-3
Kuittinen H, Aguadé M, Charlesworth D, Haan ADE, Lauga B, Mitchell-Olds T, Oikarinen S, Ramos-Onsins S, Stranger B, van Tienderen P, Savolainen O (2002) Primers for 22 candidate genes for ecological adaptations in Brassicaceae. Molec Ecol Notes 2:258–262. https://doi.org/10.1046/j.1471-8286.2002.00210.x
Kutnjak D, Schönswetter P, Dullinger S, Kuttner M, Niketić M, Frajman B (2014) Escaping to the summits: phylogeography and predicted range dynamics of Cerastium dinaricum, an endangered high mountain plant endemic to the western Balkan Peninsula. Molec Phylogenet Evol 78:365–374. https://doi.org/10.1016/j.ympev.2014.05.015
Kuzmanović N, Comanescu P, Frajman B, Lazarević M, Paun O, Schönswetter P, Lakušić D (2013) Genetic, cytological and morphological differentiation within the Balkan-Carpathian Sesleria rigida sensu Fl. Eur. (Poaceae): a taxonomically intricate tetraploid–octoploid complex. Taxon 62:458–472. https://doi.org/10.12705/623.13
Lakušić D, Liber Z, Nikolić T, Surina B, Kovačić S, Bogdanović S, Stefanović S (2013) Molecular phylogeny of the Campanula pyramidalis species complex (Campanulaceae) inferred from chloroplast and nuclear non-coding sequences and its taxonomic implications. Taxon 62:505–524. https://doi.org/10.12705/623.1
Leitch IJ, Hanson I, Lim KY, Kovarik A, Chase MW, Clarkson JJ, Leitch AR (2008) The ups and downs of genome size evolution in polyploid species of Nicotiana (Solanaceae). Ann Bot (Oxford) 101:805–814. https://doi.org/10.1093/aob/mcm326
Lihová J, Shimizu KK, Marhold K (2006) Allopolyploid origin of Cardamine asarifolia (Brassicaceae): incongruence between plastid and nuclear ribosomal DNA sequences solved by a single-copy nuclear gene. Molec Phylogen Evol 39:759–786. https://doi.org/10.1016/j.ympev.2006.01.027
Ma JX, Li YN, Vogl C, Ehrendorfer F, Guo YP (2010) Allopolyploid speciation and ongoing backcrossing between diploid progenitor and tetraploid progeny lineages in the Achillea millefolium species complex: analyses of single-copy nuclear genes and genomic AFLP. BMC Evol Biol 10:100. https://doi.org/10.1186/1471-2148-10-100
Maddison WP (1997) Gene trees in species trees. Syst Biol 46:523–536. https://doi.org/10.1093/sysbio/46.3.523
Mandák B, Krak K, Vít P, Lomonosova MN, Belyayev B, Habibi F, Wang E, Douda J, Štorchová H (2018) Hybridization and polyploidization within the Chenopodium album aggregate analysed by means of cytological and molecular markers. Molec Phylogen Evol 129:189–201. https://doi.org/10.1016/j.ympev.2018.08.016
Marhold K (2011) Multivariate morphometrics and its application to monography at specific and infraspecific levels. In: Stuessy TF, Lack HW (eds) Monographic plant systematics: fundamental assessment of plant biodiversity. Gantner, Ruggell, pp 73–99
Marjanac L, Marjanac T (2004) Glacial history of the Croatian Adriatic and coastal Dinarides. In: Ehlers J, Gibbard PL (eds) Quaternary glaciations—extent and chronology. Elsevier, Amsterdam, pp 19–26
McCullagh P, Nelder JA (1989) Generalized linear models, 2nd edn. Chapman and Hall/CRC, Boca Raton
Médail F, Diadema K (2009) Glacial refugia infl uence plant diversity patterns in the Mediterranean Basin. J Biogeogr 36:1333–1345. https://doi.org/10.1111/j.1365-2699.2008.02051.x
Melichárková A, Španiel S, Brišková D, Marhold K, Zozomová-Lihová J (2017) Unravelling allopolyploid origins in the Alyssum montanum–A. repens (Brassicaceae) species complex: low-copy nuclear gene data complement cpDNA sequences and AFLPs. Bot J Linn Soc 184:485–502. https://doi.org/10.1093/botlinnean/box039
Melichárková A, Španiel S, Marhold K, Hurdu B-I, Drescher A, Zozomová-Lihová J (2019) Diversification and independent polyploid origins in the disjunct species Alyssum repens from the SE Alps and the Carpathians. Amer J Bot 106:1499–1518. https://doi.org/10.1002/ajb2.1370
Mereďa P, Hodálová I, Kučera J, Zozomová-Lihová J, Letz DR, Slovák M (2011) Genetic and morphological variation in Viola suavis s.l. (Violaceae) in the western Balkan Peninsula: two endemic subspecies revealed. Syst Biodivers 9:211–231. https://doi.org/10.1080/14772000.2011.603903
Miller MA, Pfeiffer W, Schwartz T (2010) Creating the CIPRES science gateway for inference of large phylogenetic trees. In: Anonymous (ed) 2010 Gateway computing environments workshop (GCE). Institute of Electrical and Electronics Engineers, New Orleans, pp 1–8. https://doi.org/10.1109/GCE.2010.5676129
Naciri Y, Linder HP (2015) Species delimitation and relationships: the dance of the seven veils. Taxon 64:3–16. https://doi.org/10.12705/641.24
Naimi B, Hamm NAS, Groen TA, Skidmore AK, Toxopeus AG (2014) Where is positional uncertainty a problem for species distribution modelling. Ecography 37:191–203. https://doi.org/10.1111/j.1600-0587.2013.00205.x
Nieto Feliner G (2011) Southern European glacial refugia: a tale of tales. Taxon 60:365–372. https://doi.org/10.1002/tax.602007
Nieto Feliner G (2014) Patterns and processes in plant phylogeography in the Mediterranean Basin: a review. Perspect Pl Ecol Evol Syst 16:265–278. https://doi.org/10.1016/j.ppees.2014.07.002
Nikolić T (ed) (2019) Flora Croatica Database. Sveučilište u Zagrebu, Prirodoslovnomatematički fakultet, Botanički zavod s botaničkim vrtom, Zagreb. Available at: http://hirc.botanic.hr/fcd/. Accessed 1 Sep 2019
Nikolić T, Antonić O, Alegro AL, Dobrovič I, Bogdanović S, Liber Z, Rešetnik I (2008) Plant species diversity of Adriatic islands: an introductory survey. Pl Biosystems 142:435–445. https://doi.org/10.1080/11263500802410769
Nocedal J, Wright S (2006) Numerical optimization, 2nd edn. Springer, Berlin
Oxelman B, Brysting AK, Jones GR, Marcussen T, Oberprieler Ch, Pfeil BE (2017) Phylogenetics of allopolyploids. Annual Rev Ecol Evol Syst 48:543–557. https://doi.org/10.1146/annurev-ecolsys-110316-022729
Phillips SJ, Anderson RP, Schapire RE (2006) Maximum entropy modeling of species geographic distributions. Ecol Modelling 190:231–259. https://doi.org/10.1016/j.ecolmodel.2005.03.026
Podani J (2001) SYN-TAX 2000: computer programs for data analysis in ecology and systematics. User’s manual. Scientia, Budapest
QGIS Development Team (2019) QGIS geographic information system, ver. 3.6.0. Open Source Geospatial Foundation project. Available at: http://qgis.osgeo.org. Accessed 1 Nov 2019
R Core Team (2019) R: a language and environment for statistical computing. R Foundation for statistical computing, Vienna. http://www.r-project.org
Rambaut A, Drummond AJ, Xie D, Baele G, Suchard MA (2018) Posterior summarisation in Bayesian phylogenetics using Tracer 1.7. Syst Biol 67:901–904. https://doi.org/10.1093/sysbio/syy032
Rešetnik I, Španiel S (2018) The new circumscription of the genus Alyssum L. (Brassicaceae) in the flora of Croatia. Glas Hrvatsk Bot Društva 6(2):4–16
Rešetnik I, Satovic Z, Schneeweiss GM, Liber Z (2013) Phylogenetic relationships in Brassicaceae tribe Alysseae inferred from nuclear ribosomal and chloroplast DNA sequence data. Molec Phylogen Evol 69:772–786. https://doi.org/10.1016/j.ympev.2013.06.026
Rešetnik I, Frajman B, Bogdanović S, Ehrendorfer F, Schönswetter P (2014) Disentangling relationships among the diploid members of the intricate genus Knautia (Caprifoliaceae, Dipsacoideae). Molec Phylogen Evol 74:97–110. https://doi.org/10.1016/j.ympev.2014.01.028
Rešetnik I, Baričevič D, Batîr Rusu D, Carović-Stanko K, Chatzopoulou P, Dajić-Stevanović Z, Gonceariuc M, Grdiša M, Greguraš D, Ibraliu A, Jug-Dujaković M, Krasniqi E, Liber Z, Murtić S, Pećanac D, Radosavljević I, Stefkov G, Stešević D, Šoštarić I, Šatović Z (2016a) Genetic diversity and demographic history of wild and cultivated/naturalised plant populations: evidence from Dalmatian sage (Salvia officinalis L., Lamiaceae). PLoS ONE 11:e0159545. https://doi.org/10.1371/journal.pone.0159545
Rešetnik I, Frajman B, Schönswetter P (2016b) Heteroploid Knautia drymeia includes K. gussonei and cannot be separated into diagnosable subspecies. Amer J Bot 103:1300–1313. https://doi.org/10.3732/ajb.1500506
Rešetnik I, Temunović M, Liber Z, Satovic Z, Bogdanović S (2020) Phylogeography of Campanula fenestrellata s.l. (Campanulaceae) in the Northern Adriatic. Pl Syst Evol 306:42. https://doi.org/10.1007/s00606-020-01668-6
Rojas-Andrés BM, Albach DC, Martínez-Ortega MM (2015) Exploring the intricate evolutionary history of the diploid–polyploid complex Veronica subsection Pentasepalae (Plantaginaceae). Bot J Linn Soc 179:670–692. https://doi.org/10.1111/boj.12345
Royle JA, Chandler RB, Yackulic C, Nichols JD (2012) Likelihood analysis of species occurrence probability from presence-only data for modelling species distributions. Meth Ecol Evol 3:545–554. https://doi.org/10.1111/j.2041-210X.2011.00182.x
SAS Institute Inc. (2009) SAS/STAT® 9.2 user’s guide, 2nd edn. SAS Institute, Cary
Schlüter PM, Harris SA (2006) Analysis of multilocus fingerprinting data sets containing missing data. Molec Ecol Notes 6:569–572. https://doi.org/10.1111/j.1471-8286.2006.01225.x
Schmitt T (2007) Molecular biogeography of Europe: pleistocene cycles and postglacial trends. Frontiers Zool 4:11. https://doi.org/10.1186/1742-9994-4-11
Schönswetter P, Schneeweiss GM (2009) Androsace komovensis sp. nov., a long mistaken local endemic from the southern Balkan Peninsula with biogeographic links to the Eastern Alps. Taxon 58:544–549. https://doi.org/10.1002/tax.582018
Shimizu-Inatsugi R, Lihová J, Iwanaga H, Kudoh H, Marhold K, Savolainen O, Watanabe K, Yakubov VV, Shimizu KK (2009) The allopolyploid Arabidopsis kamchatica originated from multiple individuals of Arabidopsis lyrata and Arabidopsis halleri. Molec Ecol 18:4024–4048. https://doi.org/10.1111/j.1365-294X.2009.04329.x
Sikora M, Mihanović H, Vilibić I (2014) Paleo-coastline of the Central Eastern Adriatic Sea, and Paleo-Channels of the Cetina and Neretva rivers during the last glacial maximum. Acta Adriat 55(1):3–18
Simmons MP, Ochoterena H (2000) Gaps as characters in sequence-based phylogenetic analyses. Syst Biol 49:369–381. https://doi.org/10.1093/sysbio/49.2.369
Skokanová K, Hodálová I, Mereďa P, Slovák M, Kučera J (2019) The Cyanus tuberosus group (Asteraceae) in the Balkans: biological entities require correct names. Pl Syst Evol 305:569–596. https://doi.org/10.1007/s00606-019-01576-4
Small RL, Cronn RC, Wendel JF (2004) Use of nuclear genes for phylogeny reconstruction in plants. Austral Syst Bot 17:145–170
Španiel S, Marhold K, Passalacqua NG, Zozomová-Lihová J (2011) Intricate variation patterns in the diploid-polyploid complex Alyssum montanum–A. repens in the Apennine Peninsula: an evidence for long-term persistence and diversification. Amer J Bot 98:1887–1904. https://doi.org/10.3732/ajb.1100147
Španiel S, Marhold K, Zozomová-Lihová J (2017a) The polyploid Alyssum montanum–A. repens complex in the Balkans: a hotspot of species and genetic diversity. Pl Syst Evol 303:1443–1465. https://doi.org/10.1007/s00606-017-1470-3
Španiel S, Zozomová-Lihová J, Marhold K (2017b) Revised taxonomic treatment of the Alyssum montanum–A. repens complex in the Balkans: a multivariate morphometric analysis. Pl Syst Evol 303:1413–1442. https://doi.org/10.1007/s00606-017-1468-x
Španiel S, Rešetnik I, Buzjak S (2018) The lost and rediscovered holotype of Alyssum austrodalmaticum (Brassicaceae) is in line with the latest taxonomic treatment. Phytotaxa 379:57. https://doi.org/10.11646/phytotaxa.379.1.5
Španiel S, Marhold K, Zozomová-Lihová J (2019) Polyphyletic Alyssum cuneifolium (Brassicaceae) revisited: morphological and genome size differentiation of recently recognized allopatric taxa. J Syst Evol 57:287–301. https://doi.org/10.1111/jse.12464
Stevanović V, Kit T, Petrova A (2007) Mapping the endemic flora of the Balkans—a progress report. Bocconea 21:131–137
Stevanovič V, Vukojičić S, Šinžar-Sekulić J, Lazarević M, Tomović G, Tan K (2009) Distribution and diversity of arctic-alpine species in the Balkans. Pl Syst Evol 283:219–235. https://doi.org/10.1007/s00606-009-0230-4
Surina B, Schönswetter P, Schneeweiss GM (2011) Quarternary range dynamics of ecologically divergent species (Edraianthus serpyllifolius and E. tenuifolius, Campanulaceae) within the Balkan refugium. J Biogeogr 38:1381–1393. https://doi.org/10.1111/j.1365-2699.2011.02493
Surina B, Schneeweiss GM, Glasnović P, Schönswetter P (2014) Testing the efficiency of nested barriers to dispersal in the Mediterranean high mountain plant Edraianthus graminifolius (Campanulaceae). Molec Ecol 23:2861–2875. https://doi.org/10.1111/mec.12779
Voges A (ed) (1995) International quaternary map of Europe 1:2500000. B 10 Bern. Bundesanstalt für Geowissenschaften und Rohstoffe/UNESCO, Hannover
Winkler M, Escobar García P, Gattringer A, Sonnleitner M, Hülber K, Schönswetter P, Schneeweiss GM (2017) A novel method to infer the origin of polyploids from Amplified Fragment Length Polymorphism data reveals that the alpine polyploid complex of Senecio carniolicus (Asteraceae) evolved mainly via autopolyploidy. Molec Ecol Resources 17:877–892. https://doi.org/10.1111/1755-0998.12641
Young ND, Healy J (2003) GapCoder automates the use of indel characters in phylogenetic analysis. BMC Bioinf 4:6. https://doi.org/10.1186/1471-2105-4-6
Zozomová-Lihová J, Marhold K, Španiel S (2014) Taxonomy and evolutionary history of Alyssum montanum (Brassicaceae) and related taxa in southwestern Europe and Morocco: diversification driven by polyploidy, geographic and ecological isolation. Taxon 63:562–591. https://doi.org/10.12705/633.18
Zozomová-Lihová J, Malánová-Krásná I, Vít P, Urfus T, Senko D, Svitok M, Kempa M, Marhold K (2015) Cytotype distribution patterns, ecological differentiation and genetic structure in a diploid–tetraploid contact zone of Cardamine amara. Amer J Bot 102:1380–1395. https://doi.org/10.3732/ajb.1500052
Zwickl DJ (2006) Genetic algorithm approaches for the phylogenetic analysis of large biological sequence datasets under the maximum likelihood criterion. PhD Thesis, The University of Texas, Austin
Acknowledgements
This study was financially supported by the Slovak Research and Development Agency (APVV; Grant No. APVV-17-0616) and by the GAČR—Czech Science Foundation (Grant No. 19-06632S). We are grateful to M. Perný and J. Šibík (Plant Science and Biodiversity Centre, SAS, Bratislava, Slovakia), who accompanied us in the field; D. Brišková (Berlin-Chemie AG, Bratislava, Slovakia), who generated DET1 sequence data for her diploma thesis and kindly provided them for the present study; I. Rešetnik (University of Zagreb, Croatia) for extracting and providing distribution data from the Flora Croatica Database; and J. Eriksson (University of Gothenburg, Sweden) for her advice on assembling the input file for AlloppNET computations. We also wish to thank two reviewers, who provided valuable comments and helped to improve the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Handling Editor: Ivana Rešetnik.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Contribution to “Plants of the Balkan Peninsula in Space and Time”.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Appendices
Appendix
List of population samples of Alyssum austrodalmaticum and related species under study. Populations of A. austrodalmaticum were subjected to morphometric and relative genome size measurements, and to AFLP and nuclear gene sequence analyses. The other listed species were included only in analyses of nuclear genes. The locality description and collection data are followed by the number of individuals included in AFLP, morphological (morph) and genome size (FCM-gs) analyses (if applicable), and by GenBank accession numbers for the DET1 and CHS genes (if applicable). GenBank accession numbers with superscripts refer to sequences generated in earlier studies: Melichárková et al. (2017a, 2019b); otherwise, the sequences were generated in the present study. Population codes (in bold) are identical to the collection numbers. All voucher specimens were deposited in the herbarium of the Institute of Botany, SAS, Bratislava, Slovakia (SAV).
Alyssum austrodalmaticum Trinajstić: northern diploids (lineage N2x).
53KAS, Slovenia, Primorska, Submediteransko območje, NW of the village of Kastelec, karst grassland near the cemetery, 45° 35.017′ N, 13° 51.699′ E, 357 m a. s. l., 27 May 2007, S. Španiel and M. Perný; AFLP—7 ind., morph—13 ind., FCM-gs—9 ind., DET1: MN685397–MN685399; CHS: MN685498–MN685503. 97SKR, Croatia, Primorsko-goranska županija, Krk Island, Skrbčići, along the road S of the village, 45° 02.45′ N, 14° 29.6′ E, 78 m a. s. l., 17 Apr 2008, S. Španiel and M. Perný; AFLP—7 ind., morph—20 ind., FCM-gs—10 ind., DET1: MN685439–MN685442; CHS: MN685556–MN685561. 98OSO, Croatia, Primorsko-goranska županija, Cres Island, Osor, along the road NE of the village, 44° 41.9′ N, 14° 23.883′ E, 28 m a. s. l., 17 Apr 2008, S. Španiel and M. Perný; AFLP—7 ind., morph—20 ind., FCM-gs—10 ind., DET1: MN685419–MN685422; CHS: MN685530–MN685534. 99CRE, Croatia, Primorsko-goranska županija, Cres Island, along the road N of the town of Cres, 44° 58.95′ N, 14° 24.667′ E, 157 m a. s. l., 17 Apr 2008, S. Španiel and M. Perný; AFLP—0 ind., morph—20 ind., FCM-gs—10 ind. 100BKAa,b, Croatia, Primorsko-goranska županija, Krk Island, Baška valley, two microlocalities—45° 01.683′ N, 14° 40.633′ E, 273 m a. s. l. and 44° 58.133′ N, 14° 46.017′ E, 41 m a. s. l., 17 Apr 2008, S. Španiel and M. Perný; AFLP—7 ind., morph—33 ind., FCM-gs—17 ind. 101GRO, Croatia, Primorsko-goranska županija, town of Grobnik, Grobničko polje, grassland near the motor-racing circuit, 45° 23.000′ N, 14° 31.046′ E, 318 m a. s. l., 18 Apr 2008, S. Španiel and M. Perný; AFLP—7 ind., morph—23 ind., FCM-gs—9 ind., DET1: MN685381–MN685386; CHS: MN685483–MN685485. 104RAB, Croatia, Primorsko-goranska županija, Rab Island, rocky area near the port of Mišnjak, 44° 42.417′ N, 14° 51.583′ E, 10 m a. s. l., 18 Apr 2008, S. Španiel and M. Perný; AFLP—7 ind., morph—29 ind., FCM-gs—10 ind., DET1: MN685433–MN685436; CHS: MN685551–MN685554. 105KAG, Croatia, Ličko-senjska županija, S of the town of Karlobag, rocks near the road along the coast, 44° 31.05′ N, 15° 05.333′ E, 25 m a. s. l., 18 Apr 2008, S. Španiel and M. Perný; AFLP—7 ind., morph—28 ind., FCM-gs—9 ind. 106PAG, Croatia, Zadarska županija, Pag Island, rocks near the road above the town of Pag, 44° 26.333′ N, 15° 02.233′ E, 162 m a. s. l., 19 Apr 2008, S. Španiel and M. Perný; AFLP—7 ind., morph—30 ind., FCM-gs—10 ind., DET1: MN685423-MN685427; CHS: MN685535-MN685540. 107OBR, Croatia, Zadarska županija, S of the town of Obrovać, near the crossroad Knin-Zadar-Obrovać, 44° 11.783′ N, 15° 40.85′ E, 98 m a. s. l., 19 Apr 2008, S. Španiel and M. Perný; AFLP—7 ind., morph—24 ind., FCM-gs—9 ind., DET1: MN685413–MN685418; CHS: MN685524–MN685529. 108PJS, Croatia, Šibensko-kninska županija, N of Knin, near Prevjes, between villages of Padene and Palanka, 44° 06.733′ N, 16° 04.65′ E, 338 m a. s. l., 19 Apr 2008, S. Španiel and M. Perný; AFLP—0 ind., morph—22 ind., FCM-gs—10 ind. 126KNI, Croatia, Šibensko-kninska županija, town of Knin, grazing area near the road, 44° 03.05′ N, 16° 11.15′ E, 327 m a. s. l., 26 Apr 2008, S. Španiel and M. Perný; AFLP—7 ind., morph—24 ind., FCM-gs—10 ind., DET1: MN685400–MN685403; CHS: MN685506–MN685511. 127LJU-diploids, Croatia, Ličko-senjska županija, the village of Ljubovo, grazing land and rocks (minefield), 44° 39.667′ N, 15° 33.433′ E, 935 m a. s. l., 28 Apr 2008, S. Španiel and M. Perný; AFLP—0 ind., morph—4 ind., FCM-gs—5 ind. 128BRS, Croatia, Primorsko-goranska županija, W of the village of Brseč (ca 13 km S of Lovran), rocky slope and the summit area of Mt. Sisol, 45° 10.6′ N, 14° 12.2′ E, 764 m a. s. l., 27 Apr 2008, S. Španiel and M. Perný; AFLP—7 ind., morph—0 ind., FCM-gs—10 ind. 129OCI, Slovenia, Primorska, between villages of Ocizla and Petrinje, ruderal site near the road, 45° 34′30″ N, 13° 54′39″ E, 411 m a. s. l., 28 Apr 2008, S. Španiel and M. Perný; AFLP—0 ind., morph—0 ind., FCM-gs—10 ind. 130LOR, Italy, Friuli-Venezia Giulia, SE of Basovizza near Trieste, ca 3 km from village of San Lorenzo, rocks and hayfield near the road, 45° 37′46″ N 13° 53′34″ E, 482 m a. s. l., 28 Apr 2008, S. Španiel and M. Perný; AFLP—0 ind., morph—0 ind., FCM-gs—10 ind. 131OPI, Italy, Friuli-Venezia Giulia, the village of Opicina near the town of Trieste, grassland near the road, 45° 41.633′ N, 13° 48.85′ E, 318 m a. s. l., 28 Apr 2008, S. Španiel and M. Perný; AFLP—7 ind., morph—22 ind., FCM-gs—10 ind.
Alyssum austrodalmaticum: central tetraploids (lineage C4x).
127LJU-tetraploids, Croatia, Ličko-senjska županija, the village of Ljubovo, grazing land and rocks (minefield), 44° 39.667′ N, 15° 33.433′ E, 935 m a. s. l., 28 Apr 2008, S. Španiel and M. Perný; AFLP—0 ind., morph—6 ind., FCM-gs—23 ind. 124DRR, Bosnia and Herzegovina, Kanton 10 (Livanjski kanton), rocky face near the road between the village of Drvar and the settlement of Resanovci, 44° 20.85′ N, 16° 21.117′ E, 750 m a. s. l., 26 Apr 2008, S. Španiel and M. Perný; AFLP—7 ind., morph—0 ind., FCM-gs—10 ind., DET1: MN685369–MN685380; CHS: MN685472–MN685482. 240HUM, Croatia, Splitsko-dalmatinska županija, Brač Island, along the road NW of the settlement of Gornji Humac, 43° 16.497′ N, 16° 42.623′ E, 435 m a. s. l., 16 May 2009, S. Španiel and J. Šibík; AFLP—7 ind., morph—21 ind., FCM-gs—10 ind., DET1: MN685387–MN685396; CHS: MN685486–MN685497. 241CAA, Croatia, Dubrovačko-neretvanska županija, Korčula Island, along the road between the villages of Čara and Pupnat, 42° 56.395′ N, 16° 57.132′ E, 284 m a. s. l., 17 May 2009, S. Španiel and J. Šibík; AFLP—6 ind., morph—18 ind., FCM-gs—10 ind., DET1: MN685360–MN685368; CHS: MN685460–MN685471.
Alyssum austrodalmaticum: southern diploids (lineage S2x).
111NEU, Croatia, Dubrovačko-neretvanska županija, SE of the town of Neum, rock face along the road near the shore, 1.5 km S of the border Bosnia and Herzegovina—Croatia, 42° 52.917′ N, 17° 39.75′ E, 60 m a. s. l., 20 Apr 2008, S. Španiel and M. Perný; AFLP—7 ind., morph—30 ind., FCM-gs—10 ind., DET1: MN685407–MN685412; CHS: MN685518–MN685523. 117PON, Croatia, Dubrovačko-neretvanska županija, Pelješac Peninsula, rocks near the road in the village of Ponikve (Boljenovići), 42° 50.933′ N, 17° 37.317′ E, 223 m a. s. l., 23 Apr 2008, S. Španiel and M. Perný; AFLP—7 ind., morph—30 ind., FCM-gs—10 ind., DET1: MN685429–MN685432; CHS: MN685544–MN685548. 119KOR, Croatia, Dubrovačko-neretvanska županija, Mljet Island, between the villages of Korita and Maranovići, 42° 42.6′ N, 17° 41.267′ E, 250 m a. s. l., 23 Apr 2008, S. Španiel and M. Perný; AFLP—7 ind., morph—30 ind., FCM-gs—10 ind., DET1: MN685404–MN685406; CHS: MN685512–MN685515.
Alyssum austrodalmaticum: southern tetraploids (lineage S4x).
116SRD, Croatia, Dubrovačko-neretvanska županija, Mt. Srd above the town of Dubrovnik, 42° 38.85′ N, 18° 06.95′ E, 348 m a. s. l., 23 Apr 2008, S. Španiel and M. Perný; AFLP—7 ind., morph—29 ind., FCM-gs—11 ind., DET1: MN685443–MN685454; CHS: MN685562–MN685571.
Alyssum bosniacum Beck.
199BOR, Bosnia and Herzegovina, Hercegovačko-neretvanski kanton, Prenj Mts, Mt. Borašnica, W of the village of Borci (15 km S of the town of Konjic), 43° 33.902′ N, 17° 57.695′ E, 1590 m a. s. l., 22 Jun 2008, S. Španiel and M. Perný; DET1: MK164705b; CHS: MN685458, MN685459. 200PRO, Bosnia and Herzegovina, Srednjobosanski kanton, Vranica Mts, W of the town of Fojnica, pasture above Prokoško jezero lake, 43° 56.912′ N, 17° 45.370′ E, 1808 m a. s. l., 23 Jun 2008, S. Španiel and M. Perný; DET1: MK164805b; CHS: MN685549, MN685550.
Alyssum gmelinii Jord. & Fourr.
52KRA, Austria, Steiermark, Kraubath an der Mur, serpentine rocks in the pine forest, 47° 16.93′ N, 14° 55.65′ E, 635 m, 26 May 2007, S. Španiel and M. Perný; DET1: KX697250a; CHS: KX697101a. 133KEL, Serbia, Severna Bačka, Kelebija near the town of Subotica, 46° 09.154′ N, 19° 38.627′ E, 129 m, 13 May 2008, S. Španiel and J. Šibík; DET1: KX697247a; CHS: KX697098a.
Alyssum moellendorfianum Asch. ex Beck.
120KIC, Bosnia and Herzegovina, Hercegovačko-neretvanski kanton, town of Konjic, limestone rocks and screes, 43° 39.85′ N, 17° 58.267′ E, 370 m a. s. l., 25 Apr 2008, S. Španiel and M. Perný; DET1: MK164739–MK164740b; CHS: MN685504, MN685505. 121POD, Bosnia and Herzegovina, Hercegovačko-neretvanski kanton, village of Podorašac, limestone rocks and screes, 43° 41.583′ N, 17° 59.1′ E, 360 m a. s. l., 25 Apr 2008, S. Španiel and J. Šibík; DET1: MK164795b; CHS: MN685543.
Alyssum montenegrinum (Bald.) Španiel, Lihová & Marhold.
115PET, Montenegro, Budva, rock face along the road near the town of Petrovac (towards the town of Virpazar), 42° 13.028′ N, 18° 57.334′ E, 392 m a. s. l., 22 Apr 2008, S. Španiel and M. Perný; DET1: MN685428; CHS: MN685541–MN685542. 192BOA, Montenegro, Šavnik, between the settlements of Šavnik and Boan near a crossroad to Timar, along the road above the river Bukovica, limestone screes, 42° 59.202′ N, 19° 11.302′ E, 1193 m a. s. l., 19 Jun 2008, S. Španiel and M. Perný; DET1: MN685358, MN685359; CHS: MN685456, MN685457.
Alyssum repens Baumg.
419JHN, Austria, Kärnten, SE of Sankt Paul im Lavanttal, Johannesberg, grasslands near the church in the village, 46° 41.633′ N, 14° 52.733′ E, 606 m a. s. l., 14 May 2013, W. Franz, A. Drescher, S. Španiel, J. Zozomová-Lihová and K. Marhold; DET1: KX697246a; CHS: KX697097a. 416PEG, Austria, Steiermark, Steirisches Randgebirge, Murtal N Peggau, northern part of the Peggauer Wand, rock face, 47° 12.550′ N, 15° 20.816′ E, 522 m a. s. l., 13 May 2013, A. Drescher, S. Španiel, J. Zozomová-Lihová and K. Marhold; DET1: KX697264–KX697265a; CHS: KX697115–KX697116a.
Alyssum spruneri Jord. & Fourr. subgroup ‘serbicum’ (for its circumscription, see Španiel et al. 2017a, b).
140TOP, Serbia, Pirot, the village of Topli do (NE of the town of Pirot) ruderal sites and rocks along the road, 43° 17.407′ N, 22° 35.283′ E, 446 m a. s. l., 17 May 2008, S. Španiel and J. Šibík; DET1: MK164830–MK164831b; CHS: MN685573. 143MAG, Serbia, Raška, Maglič fortress above the river Ibar, serpentine slopes, 43° 36.782′ N, 20° 33.065′ E, 374 m a. s. l., 18 May 2008, S. Španiel and J. Šibík; DET1: MK164759b; CHS: MN685516, MN685517.
Alyssum vernale Kit. ex Hornem.
27SKE, Romania, Mehedinţi, Schela Cladovei (part of the town of Drobeta-Turnu Severin, 3 km W of the town), 44° 38.065′ N, 22° 35.610′ E, 70 m a. s. l., 5 May 2007, S. Španiel, J. Zozomová-Lihová, K. Marhold and V. Kolarčik [The locality was recently destroyed due to the construction of a road.]; DET1: MN685437, MN685438; CHS: MN685555. 134SUM, Serbia, Južni Banat, Deliblatska peščara, the village of Šumarak, between the villages of Gaj and Dubovac, 44° 49.106′ N, 21° 08.581′ E, 105 m a. s. l., 13 May 2008, S. Španiel and J. Šibík; DET1: MN685455; CHS: MN685572.
Information on electronic supplementary material
Online Resource 1. Map showing distribution records of Alyssum austrodalmaticum used for species distribution modelling, gathered from the here sampled populations and from the data stored in the Flora Croatica Database (Nikolić 2019, available at: https://hirc.botanic.hr/fcd).
Online Resource 2. Morphological characters used in the present study, and the results of morphometric analyses (eigenvectors of PCA, total canonical structure of CDA).
Online Resource 3. Cohen’s d and its 95% confidence interval of Nei-Li genetic distances between in silico tetraploids inferred under different scenarios and natural sampled tetraploids of Alyssum austrodalmaticum (lineages C4x and S4x) based on AFLP data.
Online Resource 4. Bayesian majority-rule consensus tree based on CHS sequence data obtained from Alyssum austrodalmaticum and its close relatives.
Online Resource 5. Summary data of flow cytometric measurements of relative genome size in Alyssum austrodalmaticum and its genetic lineages.
Online Resource 6. Principal component analysis of Alyssum austrodalmaticum at the level of individual plants based on 20 morphological characters.
Rights and permissions
About this article
Cite this article
Zozomová-Lihová, J., Melichárková, A., Svitok, M. et al. Pleistocene range disruption and postglacial expansion with secondary contacts explain the genetic and cytotype structure in the western Balkan endemic Alyssum austrodalmaticum (Brassicaceae). Plant Syst Evol 306, 47 (2020). https://doi.org/10.1007/s00606-020-01677-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00606-020-01677-5