Abstract
Locally adapted ecotypes can constitute an important part of the biodiversity, especially in young floras with few endemic species. However, the origins, distinctness and conservation value of many ecotypes remain uncertain because genetic data are lacking or no common-garden study has been carried out. In the present study, we evaluated the distinctness and genetic structure of a phenotypically deviating morph of Rhinanthus angustifolius, growing in calcareous spring fens on the Baltic island of Gotland. Using data from a common-garden experiment and analyses of nuclear microsatellite variation, we compared fen populations on Gotland with conspecific populations from habitats more typical of the study species. We also included the fen specialist R. osiliensis from the Baltic island of Saaremaa in the molecular analyses to make further inferences about the origin of the Gotlandic fen morph. Our data indicate that the Gotlandic fen populations constitute a phenotypically and genetically distinct ecotype that most likely has evolved at least two times on Gotland after the last ice age. In congruence with previous studies, we also infer that fen ecotypes have evolved independently on Gotland and Saaremaa. We propose a varietal status for the Gotlandic fen ecotype and give recommendations for the conservation of this taxon.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
During the last glacial period, which lasted until about 15.000 ybp, Scandinavia was covered by the thick Weichselian ice sheet (Björck 1995). As a consequence, the present-day flora is young with few endemic taxa at the species level (Jonsell 1988, 2004). Many plant species have, however, adapted to local ecological conditions through the formation of phenotypically distinct ecotypes (Turesson 1922). Ecotypes constitute an important part of Scandinavian biodiversity and may be unique, not only to specific habitats but also to the region as a whole (Jonsell 1988). For conservation authorities, it is important to consider ecotypes and other types of habitat-correlated variation (e.g. clines; Gregor 1938) in order to conserve the genetic diversity and long-term adaptive potential of species that occur in Scandinavia (Lundqvist et al. 2007). This work, however, may be hampered by the lack of consensus regarding the taxonomic treatment of ecotypes, which can be variously treated as subspecies or varieties, or remain unnamed, depending on local traditions and other factors (Hamilton and Reichard 1992; Jonsell 2004). To assess the origins, distinctness and conservation value of ecotypes, it is appropriate to use a combination of molecular-genetic analyses (Lowe et al. 2004; Stuessy et al. 2014) and common-garden experiments (Heslop-Harrison 1964), and to apply commonly accepted principles to the taxonomic treatment of the entities recognized.
Species of the annual, hemiparasitic plant genus Rhinanthus (Orobanchaceae), especially the two most widely distributed ones, R. minor L. and R. angustifolius C.C.Gmel., are highly polymorphic and renowned for their capacity to undergo ecotypic differentiation in response to differences in selection pressure (ter Borg 1972; Zopfi 1993a, b, 1995). Several species have independently and recurrently evolved seasonal ecotypes, differing in flowering time and related features, but considerable variation has also been recorded in characters unrelated to phenology such as leaf shape, hairiness and flower morphology (de Soó 1929; de Soó and Webb 1972). Thus, patterns of variation in Rhinanthus species tend to be complex and reticulate, which has resulted in a lack of consensus regarding the number and taxonomic level of described taxa. Ecotypes of R. angustifolius, for example, have variously been recognized as varieties, subspecies or even species (von Sterneck 1901; de Soó 1929; Mizianty 1978). The taxonomy is complicated by extensive hybridization between species (Kwak 1980; Ducarme and Wesselingh 2005) and the existence of intermediate populations that inter-connect taxonomically recognized ecotypes (Karlsson 1974). Besides, several of the characters commonly used in the taxonomic literature are known to be plastic, especially in relation to the identity of the host species (Jonstrup et al. 2016). Hence, there is considerable potential for environmental factors, such as differences in host plant composition, to mask or obscure genetically determined differences. Molecular studies of R. minor, R. angustifolius and R. alectorolophus (Scop.) Polich attest to the complex patterns expressed in previous taxonomic treatments of species in this genus (Houston and Wolff 2012; Pleines et al. 2013; Jonstrup et al. 2018).
The species at focus in the present study, R. angustifolius, is normally found in open dry grasslands such as hay meadows, coastal meadows and road verges, but putative conspecifics also occur in calcareous spring fens on the Baltic island of Gotland (SE Sweden) (Marklund 1982; Martinsson 1997; Johansson et al. 2016). The fen plants are similar to other plants of R. angustifolius in floral characters and leaf morphology but have a later flowering time, produce more nodes and fewer and shorter branches and possess a distinctive red colouration due to anthocyanin pigmentation, unlike the green colour normally found in R. angustifolius (Lindell 2006). On the island of Saaremaa (W Estonia), just 200 km away from Gotland, yet another form of Rhinanthus is found in calcareous spring fens. This form has been named R. osiliensis (Ronniger & Saarsoo) Vassilcz. and has long been considered as a local endemic for Saaremaa by Estonian botanists (Ronniger 1934). Fen plants on Gotland share many features with R. osiliensis, but possess fewer and shorter glandular hairs than those of the Estonian taxon (Lindell 2006). At present, the few fen populations on Gotland are provisionally treated as R. osiliensis and listed as near threatened (NT) in the Swedish Redlist (ArtDatabanken 2015).
A recent molecular study has revealed that R. osiliensis and the Gotland fen populations constitute distinct evolutionary units with putatively independent origins (Talve et al. 2014a, b). However, the distinctness and conservation value of the Gotland fen plants remain uncertain, as does their genetic relationship to the more typical R. angustifolius phenotype. In the study presented here, we analysed nuclear microsatellite variation as well as phenotypic data from a common-garden experiment to answer the following questions:
-
1.
Is there a phenotypically and genetically distinct entity of R. angustifolius related to calcareous spring fens on Gotland?
-
2.
Are the fen populations on Gotland related to R. osiliensis on Saaremaa and how do they relate to the widespread R. angustifolius?
-
3.
If there is a distinct entity on Gotland, should it be treated as a taxon of its own and does it constitute one or several conservation units at the genetic level?
Materials and methods
Study species
Rhinanthus angustifolius is a summer annual hemiparasitic herb that grows in various types of open grassland in western Eurasia. Plants of R. angustifolius are 20–50 cm tall with a glabrous to moderately hairy stem, opposite simple leaves and a green to reddish colour. Bracts and calyces are glabrous, but the margins of the calyces may be scabrid or puberulent. The yellow, zygomorphic flowers are placed in a terminal raceme on the central stem but can also be produced on primary and secondary branches (ter Borg 1972; de Soó and Webb 1972). The flowering period spans from May to September, with great overlap between populations (Jonstrup et al. 2016). Individual flowering date is partly determined by the number of nodes below the first flower; as a result, early flowering individuals generally have the fewest nodes (Wesselingh 2016). The flowers are mainly pollinated by bumblebees and produce large, winged seeds in flat capsules (Kwak 1978; Natalis and Wesselingh 2012). The heavy seeds fall down close to the maternal plant when shaken by wind or other agents, but long-distance dispersal by animals or hay transfer is also possible (ter Borg 1972). The outcrossing rate within populations has been estimated to vary between 0.32 and 1.00 (mean 0.76; Ducarme and Wesselingh 2013).
Plants of R. angustifolius have reduced root systems and attach to roots of other plants to support their requirement of water and essential nutrients (ter Borg 1972). The life cycle can be completed without attaching to a host, but then resulting in small plants with few flowers and seeds (A. Jonstrup, pers. obs.).
Seed collection, cultivation and phenotypic scoring
The Gotland fen phenotype has been reported from approximately six sites in two main areas on Gotland (Johansson et al. 2016). In August 2011, we collected seeds from three of these populations as well as from three geographically adjacent populations that conform to the more common and typical R. angustifolius phenotype (hereafter referred to as the fen morph and the common morph, respectively; Fig. 1, Table 1). In this study, we have chosen not to distinguish between two previously described taxa of R. angustifolius (ssp. angustifolius and ssp. grandiflorus (Wallr.) D.A. Webb) since results from common-garden experiments and molecular analyses contradict this division (Jonstrup et al. 2016, 2018). The two southern populations of the fen morph (SO2, SO3) inhabit fens grazed by Gotland ponies, while the northern one (SO1) inhabits a previously drained fen with no current management. Two of the common morph populations represent road verges (A11, A13) and the third an unmanaged coastal grassland (A12, Table 1). Capsules with ripe seeds were collected from 20 plants in each of the six study populations. To avoid collecting from closely related genotypes, we chose plants that grew > 2 m apart. We stored the seeds in separate paper bags at room temperature until they were sown in late November–early December the same year.
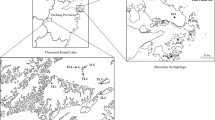
Map of southern Sweden and western Estonia (with the Baltic islands of Gotland and Saaremaa enlarged) showing the location of the 22 Rhinanthus populations used in the present study (see Table 1 for full population names)
Seeds from each maternal plant were sown in three separate pots, filled with standard soil, consisting of peat (93%), clay (4%) and sand (3%), and the following contents of essential macronutrients (nitrogen: 182 g/m3; phosphorus: 91 g/m3 and potassium: 194 g/m3; pH 5.5–6.5). The pots were planted with one of three host species: Festuca rubra (Poaceae), Lolium perenne (Poaceae) and Trifolium repens (Fabaceae) (hereafter referred to as Festuca, Lolium and Trifolium, respectively). Seeds of the host plants were obtained from a commercial seed supplier (Weibulls Horto) and had been sown earlier during the autumn of 2011. Because many Lolium and Trifolium plants died during the winter, additional seeds of these species were sown in the spring of 2012. Pots were placed randomly within and between four outdoor cold frames (blocks) in a sunny part of an experimental garden at Lund University. The pots were watered, but no extra fertilizer was added.
The Rhinanthus seeds started to germinate in late March 2012. Excess seedlings were continuously removed until only two Rhinanthus plants remained in each pot. In the summer, we scored the Rhinanthus plants for phenotypic characters deemed taxonomically important according to the previous literature (Ronniger 1934; de Soó and Webb 1972; Oja and Talve 2012) (Table 2). For scoring of hairiness, a single flower with its adjoining bract was collected and stored in 70% ethanol until measurements were taken under a stereomicroscope. The density of glandular hairs on the calyx margin was determined by counting all glandular hairs within a single ocular field at 4× magnification, centred along the upper margin at the widest part of the calyx. The mean length of the three longest glandular hairs (expressed in units of gland widths) in the ocular field was also determined, as was the corresponding measure for the bract (Table 2). During the course of the experiment, we had problems with fungal infections and aphids, which resulted in some deformation of flowers and inflorescences. To account for these factors, we estimated the rate of deformation for each plant (scored visually on a scale of 0–2) to serve as a covariate in some analyses (see below).
Voucher specimens, preserved from plants cultivated in the common garden (see Table 1), have been deposited in the Botanical Museum, Lund University (LD).
Statistical analyses of phenotypic data
We performed a combination of univariate and multivariate analyses in SPSS version 23 (IBM) to evaluate the phenotypic distinctness of the fen morph and the common morph on Gotland. Because of the unbalanced experimental design resulting from missing data, it was necessary to pool data across maternal seed families from the same population. Based on visual inspections of residuals, the error variation for quantitative traits was approximately normally distributed; thus, no transformation was deemed necessary.
The overall pattern of variation between populations and environments was assessed in a multivariate analysis of variance (MANOVA) with flowering start and measures of overall plant morphology as dependent variables; morph, population (nested within morph), host species, block and the morph × host species and population × host species interactions as independent variables; and rate of infection as covariate. Population was treated as fixed (as all the other factors) since a large part of the existing fen populations were included in the analysis, i.e. the studied populations could not be considered as a strict sample. Hair characters were analysed separately because of large gaps in the data set.
To visualize patterns of variation between morphs and populations, we performed a discriminant function analysis (DFA; also known as canonical variates analysis, Dunn and Everitt 1982), based on data pooled across blocks, with each population grown on a particular host species being a group and individual plants serving as within-group replicates. A DFA creates a reduced set of discriminant functions (DFs) that maximize the separation between pre-determined groups relative to the variation within groups (Dunn and Everitt 1982).
As a complementary approach, we assessed the phenotypic distinctness of the fen morph and the common morph by pooling quantitative data over populations, blocks and host environments, and subjecting the data to a DFA with morphs as groups. This approach mimics a natural situation in which the measured Rhinanthus plants represent a mix of sites and potentially co-occurring microhabitats (e.g. different host species) (Jonstrup et al. 2016). The distinctness of morphs was quantified as the percentage of plants classified into the original morph based on all DFs. An individual was reassigned to the other morph if its multivariate phenotype was closer to the centroid of that morph than to the centroid of its original morph.
Because of gaps in the data set, the density and length of glandular hairs on calyces and bracts were analysed separately with analysis of variance (ANOVA), based on the same model as was used in the MANOVA, with all factors treated as fixed.
Categorical data (plant colour, position of glandular hairs on bracts) were subjected to ordinal linear regression (position of glandular hairs on bracts), based on the same model as described above and a statistical testing procedure based on log-likelihood ratios. Analysis of plant colour was only performed for the common morph, since almost all fen plants fell into the same colour category (see Results).
Leaf collection and DNA extraction
Samples for molecular analyses were collected from the same fen morph populations on Gotland as those sampled for the common-garden study (SO1-SO3), whereas the common morph was represented by four populations on Gotland (A8, G2, G10, SA1), all from the same range as the fen populations, two populations on the Swedish mainland (A4, A7) and three populations on Saaremaa (EA1–EA3). Microsatellite data from seven previously studied populations of R. osiliensis (Rosi2–10) obtained from Talve et al. (2014a, b), were also included for reference purposes, giving a total of 19 sampled populations and molecular data from 509 individual plants (Fig. 1, Table 1). Leaves for the present study were collected from 40 plants per site, growing > 2 m apart. Leaves were dried and stored in plastic bags with silica gel until DNA extraction. Total genomic DNA was extracted using a modified CTAB-protocol (Doyle and Doyle 1990).
Molecular methods
Samples were prepared for microsatellite typing at 11 loci described by Ducarme et al. (2008), Houston and Wolff (2009) and Talve et al. (2014a) (Table 3). PCR reactions were done in a reaction volume of approx. 11 μL containing: 7.5 μL sterilized deionized water, 1.1 μL 10 × reaction buffer (100 mM Tris–HCl pH 8.3, 500 mM KCl, 15 mM MgCl2; Applied Biosystems), 0.93 μL dNTPs (2.5 mM of each nucleotide; Applied Biosystems), 0.17 μL M13-tailed dye (25 pmol/μL; either NED, PET, VIC or 6-FAM; DNA-Technology), 0.17 μL of reverse primer (25 pmol/μL; DNA-Technology), 0.042 μl of M13-tailed forward primer (25 pmol/μL; DNA-Technology), 0.05 μl of Taq-polymerase (5U/μL, Applied Biosystems) and 1 μl of template DNA (< 14 ng/μL) (Jonstrup et al. 2018).
PCR amplifications were carried out in a PTC-100 thermal cycler using the following program: initial denaturation at 94 °C for 5 min followed by 30 cycles of: 94 °C for 30 s, the annealing temperature (Ta, Table 2) for 45 s and 72 °C for 45 s, followed by 9 cycles of: 94 °C for 30 s, 53 °C for 45 s and 72 °C for 45 s, and a final extension of 72 °C for 10 min (Jonstrup et al. 2018). PCR products were sent to Uppsala Genome Centre, where an internal size standard (Genescan, Liz500) was added and fragments were separated using an ABI3730XL DNA Analyser. Fragments were sized using the software GENIOUS 8.0 (Kearse et al. 2012) and quality-checked by eye. To enable exact comparison of our microsatellite data with those previously published for R. osiliensis (Talve et al. 2014a, b), eight of the R. osiliensis samples already analysed in Talve et al. were rerun together with the samples from Gotland. As it was still difficult to match the allele sizes at three loci (Ra58, Ra77 and Ra84), these loci were excluded from analyses including both Swedish and Estonian material.
Analyses of molecular data
Genetic structure and differentiation
To test whether the fen morph and the common morph on Gotland constitute distinct units and whether a genetic structure exists within the fen morph, we analysed the microsatellite data with the software STRUCTURE 2.3.4, which identifies clusters that minimize Hardy–Weinberg and linkage disequilibria based on multilocus genotypes (Pritchard et al. 2000). A second analysis was performed to assess the relationship between the Gotland fen morph, the common morph of R. angustifolius from both Sweden and Estonia, and R. osiliensis from Saaremaa. We used settings according to recommendations given in the STRUCTURE manual (Pritchard et al. 2000). Simulations were run for 1–10 clusters (K = 1 to K = 10) in the first analysis and for K = 1 to K = 24 in the second analysis. For each value of K, the analysis was repeated 20 times with a burn-in period of 100 000 iterations followed by additional 100 000 iterations to obtain parameter estimates. We ran the analyses with prior information on population identity and applied a model that assumes admixture between populations and correlated allele frequencies (Falush et al. 2003). The optimal K value, i.e. the number of clusters that best fit the data, for each analysis was determined by the Evanno method (Evanno et al. 2005), implemented in the web platform STRUCTURE HARVESTER 0.6.94 (Earl Dent and von Holdt 2012). Separate runs for each K were aligned using CLUMPP 1.1.2 (Jakobsson and Rosenberg 2007), and the results were visualized with DISTRUCT 1.1 (Rosenberg 2004).
Patterns of differentiation between and within morphs were also evaluated using analysis of molecular variance (AMOVA), as implemented in ARLEQUIN 3.5.2.2 (Excoffier and Lischer 2010). We grouped populations by morph (the two different morphs of R. angustifolius with or without R. osiliensis as the third group) and ran two sets of analyses, based on pairwise FST and RST-values as measures of differentiation, respectively. FST, also called the fixation index, is based on the infinite alleles model (IAM; Kimura and Crow 1964) and treats all distinct alleles as equally different, while RST assumes a stepwise mutation model (SMM; Slatkin 1995; Rousset 1996) by taking into account the length of each microsatellite allele. FST-based analyses suffer less from sampling variance than analyses based on RST (Balloux and Goudet 2002). We visualized the genetic structure by means of principal coordinates analyses (PCOs), performed in the program NTSYS-pc 2.20 (Rohlf 1994) using pairwise FST-values obtained from ARLEQUIN. To test for geographical structure at the regional scale, we performed a Mantel permutation test on geographical distances and FST-values for pairs of populations on Gotland, using ARLEQUIN with data obtained in SPAGeDi 1.5 (Hardy and Vekemans 2002).
Genetic diversity within populations
The genetic diversity of individual populations was estimated as the number of alleles (A), the number of effective alleles (Ae), the number of private alleles (Ap), the observed and expected heterozygosity (Ho, He) and the coefficient of inbreeding (FIS), all calculated using GenAlEx 6.502 (Peakall and Smouse 2012). Since sample sizes differed between populations (Table 1), we also estimated allelic richness adjusted for sample size (Aadj) by means of rarefaction, as implemented in SPAGeDi 1.5 (Hardy and Vekemans 2002).
Input files for analyses in STRUCTURE, ARLEQUIN and SPAGeDi were created in CREATE (Coombs et al. 2008).
Results
Phenotypic variation
Flowering time and overall plant morphology differed significantly between the fen morph and the common morph, and between the three populations representing each morph in the common-garden experiment (morph: F4,414 = 81.52, P < 0.001; population: F16,1265 = 39.36, P < 0.001; MANOVA). Judging from the DF plot, individuals of the fen morph generally had larger values along the first axis of separation (DF1, Fig. 2a, b) owing to the presence of more nodes, more branches and a later flowering start (Table 4). Environmental factors exerted significant effects on the mean phenotype (host: F8,828 = 2.79, P < 0.01; block: F12,1096 = 3.50, P < 0.001); however, there was no clearly visible pattern of response across the three host species along the two major axes of separation (Fig. 2b). Our analysis revealed a weak, though statistically significant, population-by-host interaction (F32,1528 = 1.63, P < 0.05) but no morph-by-host interaction (F8,828 = 0.89, P > 0.05).
In the DFA of data pooled over blocks, host environments and replicate populations, 41% of the variation in flowering time and plant morphology could be attributed to differences between morphs (1-Wilks lambda = 0.41, P < 0.001). 81% of the individuals were correctly assigned to their original morph when they were re-classified according to the DFs (data not shown).
None of the individuals of either the fen morph or the common morph had normal or glandular hairs on the main surface of the calyx or corolla, but a thin zone of glandules and non-glandular hairs was always present along the margin of the calyx (A. Jonstrup, personal observation). Most of the glandules were sessile, but a few stalked glandular hairs were often present. Plants of the fen morph and the common morph differed in both the density and length of glandular hairs on the calyx (P < 0.001–0.01, ANOVA; Table 5) with fen morph plants having a denser cover (mean ± SD 10.8 ± 8.7 vs. 5.7 ± 5.1 for the common morph) and longer hairs (3.6 ± 3.1 vs. 3.0 ± 1.2 for the common morph); note, however, that there was considerable overlap between sd’s in both cases. Populations within morphs differed significantly (P < 0.001) and both characters were significantly influenced by host species (P < 0.05 in both cases, ANOVA; Table 5). Effect sizes for block and interaction terms were too weak to be declared significant (P > 0.05, ANOVA; Table 5).
Plants of the fen morph had glandular hairs over a larger proportion of the bracts (with 32% having more than half of the bract covered by hairs) than plants of the common morph (no plant with the entire bract covered by hairs; P < 0.001, ordinal logistic regression), whereas the effect of population within morph was nonsignificant (P > 0.05, ordinal logistic regression). Growth on different host species influenced the area covered by glandular hairs (P < 0.01, ordinal logistic regression), with individuals grown on Trifolium having the highest scores (data not shown). The length of the glandular hairs on the bracts was not significantly affected by any of the factors considered (P > 0.05, ANOVA; Table 5).
The fen morph was distinctively uniform in colouration, with 99% of the plants being red and 1% being green. Plants of the common morph were more variable (75% green, 25% red), with no consistent differences between blocks, populations and host environments (P > 0.10, ordinal logistic regression).
Molecular-genetic differentiation
We performed separate analyses in computer program STRUCTURE (Pritchard et al. 2000) to explore the genetic structuring of populations from Gotland only, and to explore the genetic structuring of the combined material from Gotland, mainland Sweden and Estonia. The resulting patterns are illustrated graphically in Figs. 3 and 4, respectively, and we have chosen to present the results given by K = 2 to K = 4, since in each case these are the most relevant partitions relating to the hypothesis formulated in the introduction.
The probability of each individual of Rhinanthus angustifolius from Gotland being member of a particular cluster identified by STRUCTURE, based on 11 microsatellite loci. Individuals are arranged according to their population and region. Colours denote estimated membership fraction. K = 4 gave the best fit, but results are also shown for K = 2–3
The probability of each individual of Rhinanthus from Sweden and Estonia being member of a particular cluster identified by STRUCTURE, based on a subset of 8 microsatellite loci. Individuals are arranged according to their population and region. Colours denote estimated membership fraction. K = 2 gave the best fit, but results are also shown for K = 3–4
In the STRUCTURE-analysis restricted to the fen morph and the common morph on Gotland (Fig. 3), the optimal partitioning was obtained at K = 4. Individuals from fen morph populations split into two distinct clusters, one comprising individuals from two southern populations (SO2, SO3) and another comprising individuals from a northern population (SO1), together with those from a nearby roadside population (SA1). Individuals from two southern common morph populations (A8, G2) formed a third cluster, and individuals from a northern common morph population (G10) were joined in a fourth cluster. For K = 3, individuals from fen morph populations remained separated into two clusters. Population SO1 was still grouped with the nearby common morph population SA1, while individuals from the remaining common morph populations formed their own cluster (Fig. 3). For K = 2, individuals from the northern fen morph population (SO1) continued to form a distinct cluster together with individuals from the SA1 population, while individuals from the northern common morph population (G10) constituted another distinct cluster. The remaining fen morph and common morph plants were about equally likely to belong to either of the two clusters (Fig. 3).
In the STRUCTURE-analysis based on the whole material from Sweden and Estonia (Fig. 4), the optimal partitioning was obtained at K = 2. The two clusters corresponded to individuals from Sweden and Estonia, with exception of individuals from the Estonian population EA3, which clustered together with the Swedish individuals. Swedish and Estonian individuals of the common morph had mixed probability of belonging to any of the two clusters, while individuals of the Gotland fen morph and R. osiliensis were assigned to different regional clusters. For K = 3, the Swedish individuals separated into two clusters, with individuals from the two southern fen morph populations (SO2, SO3) forming one cluster and the remaining ones, including the northern fen morph population SO1, forming an admixture of the two Swedish clusters. Most of Estonian individuals were kept as a single cluster, whereas those of R. angustifolius, especially population EA3, continued to show some similarity to those from Sweden.
The PCO on FST-values for population pairs on Gotland (Fig. 5) confirmed the pattern found in the STRUCTURE-analysis. Populations were spread out in the PCO plot (Fig. 5), more or less forming three clusters, one with the two southern fen morph populations (SO2, SO3), one with the northern fen morph population (SO1) and the nearby roadside population (SA1), and one cluster with the remaining common morph populations (A8, G2, G10). No significant correlation was found between geographical and genetic distance between populations on Gotland (r = 0.23, P > 0.05, Mantel test).
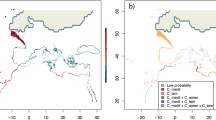
Similarly, PCO on data from all Swedish and Estonian populations showed the same large-scale geographical pattern (Fig. 6) as that seen in the STRUCTURE-analysis. Swedish and Estonian populations tended to fall into separate clusters (except for EA3), the most distinct being the Estonian R. osiliensis populations. Gotlandic fen morph populations continued to form two separate clusters, in this case with the northern cluster including not only populations SO1 and SA1, but also the Estonian R. angustifolius population EA3.
In the AMOVA based on fen morph and common morph populations from Gotland, a large portion of the diversity was due to differentiation between populations within morphs (FST: 37.7%; RST: 44.7%; Table 6), while only a minor portion was due to differentiation between morphs (FST: 7.2%; RST: 0.5%; Table 6). In the AMOVA based on all Swedish and Estonian populations, the between-morph component of differentiation was higher (FST: 18.2%; RST: 13.3%; Table 6), but much of the variation was still explained by differentiation between populations within morphs (FST: 24.7%; RST: 29.1%; Table 6).
Genetic diversity within populations
Analysis of diversity at all 11 microsatellite loci studied in the Gotland material showed fen morph populations (SO1–3) to have lower values than common morph populations (Table 7). In particular, measures of numbers of alleles per locus, richness and heterozygosity were usually the lowest for either of the two southern fen morph populations (SO2, SO3) and the highest for one of the southern common morph populations (G2, A8). Judging from a closer examination of allele frequencies (Supplementary data), the fen morph populations contained different subsets of alleles found in the common morph, especially at locus Rhos8 (fixed in SO1) and at loci Ra73, Ra75, Ra87 and Rhos8 (fixed in both SO2 and SO3).
Analyses based on the eight loci whose alleles could be matched between Swedish and Estonian populations confirmed the trend seen in the analysis of all 11 loci—relatively low diversity in Gotlandic fen populations—but also a tendency towards reduced genetic diversity in the Estonian R. osiliensis populations (Rosi2–10, Table 8).
The number of private alleles (Ap) was low for all populations, regardless of whether the analysis was based on all 11 loci (Ap = 0–0.7, Table 7) or the restricted set of eight ‘matching’ loci (Ap = 0–0.4, Table 8). Coefficients of inbreeding (FIS) were always positive, with a trend towards higher values for common morph populations of R. angustifolius (Tables 7, 8).
Discussion
Although adaptation ultimately occurs at the level of individual populations, it is often useful to identify more broadly defined ecotypes, i.e. groups of populations connected to particular habitat types (Turesson 1922), especially if the aim is to conserve the genetic diversity and long-term adaptive potential of species at the intraspecific level (Jonsell 2004; Lundqvist et al. 2007). Decisions on taxonomic status and conservation value are, however, only meaningful if ecotypes correspond to biologically meaningful entities at both the genetic and phenotypic level (Lowe et al. 2004; Mace 2004; Stuessy et al. 2014).
In this study, we performed a common-garden experiment and obtained microsatellite data to evaluate the distinctness and taxonomic status of populations of R. angustifolius found in calcareous spring fens on the Baltic island of Gotland. We compared them with other conspecific populations (collectively called the common morph), and also with R. osiliensis, an endemic taxon known from calcareous spring fens on the island of Saaremaa. Our experimental results are consistent with the presence of a phenotypically distinct fen ecotype (sensu Turesson 1922), characterized by red colouration as well as a later flowering start, a larger node and branch number, and a denser cover of glandular hairs on bracts and calyces than normally found in R. angustifolius. The Gotland fen ecotype is genetically different from R. osiliensis but appears to be closely related to conspecific populations of the common morph from Gotland. Based upon these and other observations, we propose a varietal status for the Gotlandic fen ecotype and give some recommendations for the conservation of this taxon.
Phenotypic differentiation
In accordance with previous observations (Lindell 2006), we found significant differences between the two morphs in flowering time and general plant morphology, similar to what has been observed in other rhinanthoid taxa (e.g. ter Borg 1972; Karlsson 1982; Zopfi 1993a, b, 1995). In particular, fen morph plants started flowering later and had a larger node number than individuals of the common morph, a direct consequence of the close developmental link between flowering start and node number in annual rhinanthoids (Wesselingh 2016). After pooling of data across environments, 41% of the total variation in phenology and gross morphology could be attributed to differences between the fen morph and the common morph; further, a majority of the plants (81%) were correctly re-classified to their original morph in the multivariate analysis. Thus, there was relatively little overlap between morphs across the host environments used in the common-garden experiment.
The percentage of correctly classified individuals was higher than the 61–73% correctly classified individuals in similar comparisons of two putative seasonal ecotypes within the common morph (ssp. angustifolius vs. ssp. grandiflorus; Jonstrup et al. 2016). These results indicate a greater separation between the Gotlandic fen morph and the common morph than between ecologically differentiated populations within the latter group. However, Jonstrup et al. (2016) used a broader range of characters and host environments, two factors that may have resulted in a larger phenotypic overlap between populations in their experiment. In particular, in the present study, we did not use the strong-growing legume Trifolium pratense as host species, a species that exerted strong and highly specific phenotypic effects on the plants used in the previous study (Jonstrup et al. 2016).
Consistent with work on other Rhinanthus species (de Soó 1929; de Soó and Webb 1972), we also identified differences in characters unrelated to flowering time and node number. First, almost all fen plants were distinctly red because of anthocyanin pigmentation, while a majority of the common morph plants had normal green colouration, irrespective of population and environment. Second, plants of the fen morph had a denser and more extensive cover of glandular hairs on the calyx and the bracts subtending the flowers, and the glandular hairs on the calyx margin were longer. There was, however, considerable overlap between morphs and the glandular hairs on the fen plants were never as long and dense as observed in R. osiliensis (A. Jonstrup, personal observation).
Calcareous spring fens on Gotland are characterized by cold upwelling water all year around and by a soil poor in available nutrients (especially phosphorous) due to high calcium content (Sjörs 1956; Pettersson 1958; Marklund 1982), two factors that could enhance the selective advantage of features characterizing the fen morph phenotype. For example, the high content of anthocyanin may act as a defence against oxidative stress caused by mineral imbalance (Landi et al. 2015). A similar hypothesis applies to the relatively dense cover of glandular hairs observed in the Gotland fen morph and, to an even greater extent, the densely pubescent R. osiliensis on Saaremaa. Glandular hairs of Rhinanthus have recently been shown to function as water excreting hydathode trichomes, which help to create a low water potential and thus to facilitate the transport of water and nutrients from the roots of the host plant (Světlíková et al. 2015). However, as the observed differences only concern structures directly associated with the flowers (calyces, bracts), more detailed studies are required to determine the function and adaptive significance of glandular hairs in the Gotlandic fen morph.
Differentiation at molecular marker loci
In contrast to the phenotypic differences found between the two morphs of R. angustifolius, there was no corresponding differentiation at the microsatellite loci investigated: the fen morph populations on Gotland were more strongly differentiated from one another than from geographically adjacent populations of the common morph, a pattern seen in both the PCO and STRUCTURE analyses. Thus, only a modest part of the microsatellite variation (< 20%) could be attributed to differences between morphs, irrespective of whether the AMOVA was based on all 11 microsatellite loci studied in the Gotland material or the subset of eight loci studied in both Swedish and Estonian populations. These results are in agreement with other studies that found no, or only weak, molecular support for the delimitation of infraspecific taxa within Rhinanthus species (Houston and Wolff 2012; Pleines et al. 2013; Jonstrup et al. 2018).
Incongruence between phenotypic and molecular-genetic variation has several possible explanations. Houston and Wolff (2012) suggested that such incongruence found in R. minor could be a result of phenotypic plasticity. However, plasticity is not likely to be a major reason for the incongruence observed in the present study: environmental effects and their interactions with morph and population effects had minor influence on the mean phenotype in the common-garden experiment. Instead, it appears that the Gotlandic fen morph evolved in parallel from different populations of the common morph as an adaptive response to the same selection pressures, similar to what has been inferred for other plant species (Levin 2001) including rhinanthoids in other habitats (e.g. Euphrasia: Karlsson 1974, 1976). Consistent with this scenario, the northern and southern fen populations contained different subsets of alleles found in the common morph, and more specifically, there was a close genetic similarity between the fen population SO1 and the common morph population SA1.
Because we performed a common-garden experiment, we infer that the differentiation between the fen morph and the common morph of Rhinanthus has a heritable basis, and that they accordingly constitute true ecotypes. However, our data provide no evidence as to whether the observed difference has a pure genetic background or whether epigenetic modifications of the genome also contribute. Epigenetic modifications have been shown to influence phenotypic traits in many plant systems, and these modifications can be stable over multiple generations (Pikaard and Scheid 2014). For instance, epigenetic factors seem to explain much of the phenotypic variation expressed within genetically homogeneous accessions of the inbreeding annual Arabidopsis thaliana (Zhang et al. 2018). Epigenetic modifications can also contribute to morphological differentiation between geographical populations and, in some cases, taxonomically recognized entities (e.g. Paun et al. 2010). Nevertheless, studies linking ecotypes in the original sense of Turesson (1922) to epigenetic mechanisms seem to be rare.
The genetic divergence between populations was generally large (mean FST ≈ 0.3; Online Resources 1–2), as commonly observed for annual plants with gravity-dispersed seeds (mean GST = 0.38; Hamrick and Godt 1996), indicating limited gene flow between populations. However, a history of gene flow is still possible between populations in close proximity, such as the two southern fen populations on Gotland (SO2, SO3) and the aforementioned SO1 and SA1 populations. Regarding SO2 and SO3, which shared the same subset of alleles found in the common morph, we cannot exclude a common origin, perhaps from a more ancient lineage of R. angustifolius, as the cause of this and other similarities. Furthermore, we note that common morph populations from both Sweden and Estonia were separated by relatively short genetic distances. Most of these populations represent managed grassland sites (pastures, road verges and hay fields) and they may accordingly be connected through seed dispersal by machinery, animals or hay transport, both within and between countries (ter Borg 1972). The most striking case concerns the Saaremaa population EA3, which clustered with the Swedish populations (especially those from Gotland) rather than with the other Estonian populations (Fig. 6). This connection could reflect fairly recent transport by humans, for example in hay with cattle, as previously invoked to explain the genetic proximity between Icelandic and Danish populations of R. minor (Vrancken et al. 2012).
Genetic diversity and conservation biology
Calcareous spring fens on Gotland are unique environments with a characteristic flora that includes several rare and redlisted species, including cold-adapted taxa with their only Swedish occurrences in this habitat, e.g. Tofieldia calyculata and the local endemic Euphrasia salisburgensis var. schoenicola (Sjörs 1956; Pettersson 1958). Many fens on Gotland have been lost or have become increasingly fragmented as results of drainage and exploitation during the nineteenth and twentieth centuries (Marklund 1982; Martinsson 1997). General legislations protect wetlands in southern Sweden from further exploitation, but increasing nutrient supply due to atmospheric nitrogen deposition and overgrowth still pose a threat to spring fen vegetation (Martinsson 1997). These changes call for further actions to prevent habitat degradation and raise the question of how to best preserve the genetic diversity and evolutionary potential of taxa in this habitat (Moritz 1994; Frankham 2010).
The present study suggests that the Gotland fen morph of R. angustifolius constitutes a distinct spring fen ecotype that evolved locally on Gotland and that the fen morph plants fall into two genetic clusters, one containing two southern populations and another containing a northern population (together with a nearby common morph population). Thus, to conserve the genetic diversity of R. angustifolius in general, and the fen ecotype in particular, it seems reasonable to treat the southern and northern fen populations as separate management units, each with a high conservation value (Moritz 1994; Frankham 2010). In this regard, we also call for further studies of the SA1 population to assess the taxonomic status and conservation value of this population.
Gotlandic fen populations were not particularly inbred but had lower genetic diversity and fewer private alleles than other populations, especially those of the common morph. Low genetic diversity could reduce fitness and the long-term evolutionary potential of populations by increasing the expression of recessive deleterious alleles and by decreasing the heritable variation available to selection (Young et al. 1996; Storfer 1996). Microsatellite markers are, however, assumed to be selectively neutral with little or no phenotypic effect, making it difficult to draw firm conclusions about population viability (Frankham 2010). Further studies of the fen ecotype are needed to determine whether the low genetic diversity is associated with any reduction in population viability (Leimu et al. 2006) and whether present-day populations of the fen morph have been purged of alleles with negative effects on survival and fecundity (Crnokrak and Barrett 2002).
Taxonomic treatment
Although fen morph populations on Gotland are provisionally treated as R. osiliensis by Swedish conservation authorities (ArtDatabanken 2015), there is a clear separation between fen populations on Gotland and Saaremaa (Talve et al. 2014a, b). Our current results, based on microsatellite data from both Swedish and Estonian populations, support the hypothesis of Talve et al. (2014a, b) that fen-adapted populations on Gotland and Saaremaa have separate origins—extending the parallel-adaptation scenario proposed above for individual populations of the fen morph on Gotland. These considerations, together with the patterns observed at the phenotypic level, suggest that the Gotlandic fen morph warrants recognition as a separate taxon. Since the fen morph contains a subset of the variation found within the common morph on Gotland, not only at putatively neutral marker loci but also in phenotypic characters such as plant colour, there should be great potential for rapid and recurrent shifts towards the fen morph, as a result of selection operating on pre-existing genetic variation within ancestral populations (Barrett and Schluter 2008; Pleines et al. 2013; Jonstrup et al. 2016). Thus, it seems reasonable to distinguish the Gotlandic fen morph as an infraspecific taxon within R. angustifolius rather than as a separate species. On the basis of guidelines given in the ongoing Flora Nordica project Jonsell (2004), varietal status is deemed most appropriate on the ground that it originated independently on at least two occasions on Gotland and that the different populations are most closely related to geographically adjacent populations of the common grassland morph rather than to each other (Hamilton and Reichard 1992; Jonsell 2004). Other floras often restrict the usage of infraspecific categories to the level of subspecies (e.g. de Soó and Webb 1972), but in Flora Nordica this level is used for geographically or ecogeographically subdivisions of a species in which the component populations are more closely related to each other than to other subsets of populations (Jonsell 2004).
As R. osiliensis was included as reference material only, a final evaluation of its taxonomic status must await careful comparison with other sympatrically occurring Rhinanthus taxa on Saaremaa.
Rhinanthus angustifolius C.C.Gmel. var.gotlandicus Jonstrup, H.S.Andersson & Hedrén, var. nov.—HOLOTYPE: Sweden. Gotland: Levide parish, Mallgårds källmyr, 57°19′ N, 18°17′ E, Aug 2011, Jonstrup MA12T R1 (LD). Only seeds were collected at the original locality. The type material consists of a plant cultivated and harvested 2012 in the Experimental garden, Department of Biology, Lund University, using Trifolium repens as host.
Vernacular name: källmyrsskallra.
Diagnosis: Most closely related to Rhinanthus angustifolius var. angustifolius from which it differs in having a consistently red-coloured calyx. It is phenotypically similar to R. osiliensis from which it is separated by a shorter and more lax indumentum.
Phenology: Flowering in late July and August.
Distribution area and habitat: Only known from two distribution areas on Gotland, Sweden. Calcareous spring fens.
Additional specimens examined: Sweden. Gotland: Levide parish, Mallgårds källmyr, 57°19′ N, 18°17′ E, Aug 2011, Jonstrup MA1L R1 (LD); cultivated and harvested 2012 in the Experimental garden, Department of Biology, Lund University, using Lolium perenne as host. Gerum parish, Botes källmyr, 57°20′ N, 18°19′ E, Aug 2011, Jonstrup BO17FR1 (LD); cultivated and harvested 2012 in the Experimental garden, Department of Biology, Lund University, using Festuca rubra as host.
Proposed conservation status: Near Threatened (NT) according to The Swedish Species Information Centre (ArtDatabanken).
References
ArtDatabanken (2015) Rödlistade arter i Sverige 2015. ArtDatabanken SLU, Uppsala
Balloux F, Goudet J (2002) Statistical properties of population differentiation estimators under stepwise mutation in a finite island model. Molec Ecol 11:771–783. https://doi.org/10.1046/j.1365-294X.2002.01474.x
Barrett RHD, Schluter D (2008) Adaptation from standing genetic variation. Trends Ecol Evol 23:38–44. https://doi.org/10.1016/j.tree.2007.09.008
Björck S (1995) A review of the history of the Baltic sea, 13.0–8.0 ka BP. Quatern Int 27:19–40. https://doi.org/10.1016/1040-6182(94)00057-C
Coombs JA, Letcher BH, Nislow KH (2008) CREATE: a software to create input files from diploid genotypic data for 52 genetic software programs. Molec Ecol Resources 8:578–580. https://doi.org/10.1111/j.1471-8286.2007.02036.x
Crnokrak P, Barrett SCH (2002) Purging the genetic load: a review of the experimental evidence. Evolution 56:2347–2358. https://doi.org/10.1111/j.0014-3820.2002.tb00160.x
de Soó R (1929) Die mittel- und südosteuropäischen Arten und Formen der Gattung Rhinanthus und ihre Verbreitung in Südosteuropa. Repert Spec Nov Regni Veg 26:179–219
de Soó R, Webb DA (1972) Rhinanthus L. In: Tutin TG, Heywood VH, Burges NA, Moore DH, Valentine DH, Walters SM, Webb DA (eds) Flora Europaea. Diapensiaceae to Myoporaceae, vol 3. Cambridge University Press, Cambridge, pp 276–280
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus 12:13–15
Ducarme V, Wesselingh RA (2005) Detecting hybridization in mixed populations of Rhinanthus minor and Rhinanthus angustifolius. Folia Geobot 40:151–161. https://doi.org/10.1007/BF02803231
Ducarme V, Wesselingh RA (2013) Outcrossing rates in two self-compatible, hybridising Rhinanthus species: implications for hybrid formation. Pl Biol 15:541–547. https://doi.org/10.1111/j.1438-8677.2012.00668.x
Ducarme V, Risterucci MA, Wesselingh RA (2008) Development of microsatellite markers in Rhinanthus angustifolius and cross-species amplification. Molec Ecol Resources 8:384–386. https://doi.org/10.1111/j.1471-8286.2007.01966.x
Dunn G, Everitt BS (1982) Introduction to mathematical taxonomy. Cambridge University Press, Cambridge
Earl Dent A, von Holdt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conservation Genet Resources 4:359–361. https://doi.org/10.1007/s12686-011-9548-7
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Molec Ecol 14:2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x
Excoffier L, Lischer HEL (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Molec Ecol Resources 10:564–567. https://doi.org/10.1111/j.1755-0998.2010.02847.x
Falush D, Stephens M, Pritchard JK (2003) Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164:1567–1587
Frankham R (2010) Challenges and opportunities of genetic approaches to biological conservation. Biol Conservation 143:1919–1927. https://doi.org/10.1016/j.biocon.2010.05.011
Gregor JW (1938) Experimental taxonomy II. Initial population differentiation in Plantago maritima L. of Britain. New Phytol 37:15–49. https://doi.org/10.1111/j.1469-8137.1938.tb06925.x
Hamilton CW, Reichard SH (1992) Current practice in the use of subspecies, variety, and forma in the classification of wild plants. Taxon 41:485–498. https://doi.org/10.2307/1222819
Hamrick JL, Godt MJW (1996) Effects of life history traits on genetic diversity in plant species. Philos Trans, ser B 351:1291–1298. https://doi.org/10.1098/rstb.1996.0112
Hardy OJ, Vekemans X (2002) SPAGeDi: a versatile computer program to analyse spatial genetic structure at the individual or population levels. Molec Ecol Notes 2:618–620. https://doi.org/10.1046/j.1471-8286.2002.00305.x
Heslop-Harrison J (1964) Forty years of genecology. In: Cragg JB (ed) Advances in ecological research, vol. 2. Academic Press, New York, pp 159–247
Houston K, Wolff K (2009) Eight polymorphic microsatellite markers for Rhinanthus minor. Molec Ecol Resources 9:174–176. https://doi.org/10.1111/j.1755-0998.2008.02415.x
Houston K, Wolff K (2012) Rhinanthus minor population genetic structure and subspecies: potential seed sources of a keystone species in grassland restoration projects. Perspect Pl Ecol Evol Syst 14:423–433. https://doi.org/10.1016/j.ppees.2012.08.001
Jakobsson M, Rosenberg NA (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23:1801–1806. https://doi.org/10.1093/bioinformatics/btm233
Johansson BG, Petersson J, Ingmansson G (2016) Gotlands flora, vol. 2. SBF-förlag, Uppsala
Jonsell B (1988) Mikroendemism i det baltiska landhöjningsområdet. Blyttia 46:65–73
Jonsell B (ed) (2004) Flora Nordica, general volume. The Bergius Foundation, Stockholm
Jonstrup A, Hedrén M, Andersson S (2016) Host environment and local genetic adaptation determine phenotype in parasitic Rhinanthus angustifolius. Bot J Linn Soc 180:89–103. https://doi.org/10.1111/boj.12352
Jonstrup A, Andersson S, Hedrén M (2018) Genetic structure in parasitic Rhinanthus angustifolius is determined by geographical scale rather than habitat—implications for taxonomy and conservation. Nordic J Bot 2018:e01941. https://doi.org/10.1111/njb.01941
Karlsson T (1974) Recurrent ecotypic variation in Rhinantheae and Gentianaceae in relation to hemiparasitism and mycotrophy. Bot Not 127:527–539
Karlsson T (1976) Euphrasia in Sweden: hybridization, parallelism, and species concept. Bot Not 129:49–60
Karlsson T (1982) Euphrasia rostkoviana i Sverige. Växtekol Stud 15. Uppsala, Sweden
Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, Buxton S, Cooper A, Markowitz S, Duran C, Thierer T, Ashton B, Mentjies P, Drummond A (2012) Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28:1647–1649. https://doi.org/10.1093/bioinformatics/bts199
Kimura M, Crow JF (1964) The number of alleles that can be maintained in a finite population. Genetics 49:725–738
Kwak M (1978) Pollination, hybridization and ethological isolation of Rhinanthus minor and R. serotinus (Rhinanthoideae: Scrophulariaceae) by bumblebees (Bombus Latr.). Taxon 27:145–158. https://doi.org/10.2307/1220235
Kwak M (1980) Artificial and natural hybridization and introgression in Rhinanthus (Scrophulariaceae) in relation to bumblebee pollination. Taxon 29:613–628. https://doi.org/10.2307/1220333
Landi M, Tattini M, Gould KS (2015) Multiple functional roles of anthocyanins in plant-environment interactions. Environm Exp Bot 119:4–17. https://doi.org/10.1016/j.envexpbot.2015.05.012
Leimu R, Mutikainen P, Koricheva J, Fischer M (2006) How general are positive relationships between plant population size, fitness and genetic variation? J Ecol 94:942–952. https://doi.org/10.1111/j.1365-2745.2006.01150.x
Levin DA (2001) The recurrent origin of plant races and species. Syst Bot 26:197–204
Lindell T (2006) Finns öselskallra på Gotland? Svensk Bot Tidskr 100:261–262
Lowe A, Harris S, Ashton P (2004) Ecological genetics. Design, analysis and application. Blackwell Publishing, Oxford
Lundqvist A-C, Andersson S, Lönn M (2007) Genetic variation in wild plants and animals in Sweden. A review of case studies from the perspective of conservation genetics. Swedish Environmental Protection Agency Report, Solna, Sweden
Mace G (2004) The role of taxonomy in species conservation. Philos Trans, ser B 359:711–719. https://doi.org/10.1098/rstb.2003.1454
Marklund S (1982) Källmyrar. Botanisk inventering. Länsstyrelsen i Gotlands län, Visby
Martinsson M (1997) Våtmarker på Gotland Del 1. Länsstyrelsen i Gotlands län, Visby
Mizianty M (1978) Variability of Rhinanthus serotinus (Schönh.) Oborny in Poland. Fragm Florist Geobot 24:387–425
Moritz C (1994) Defining ‘Evolutionarily Significant Units’ for conservation. Trends Ecol Evol 9:373–375. https://doi.org/10.1016/0169-5347(94)90057-4
Natalis LC, Wesselingh RA (2012) Shared pollinators and pollen transfer dynamics in two hybridizing species, Rhinanthus minor and R. angustifolius. Oecologia 170:709–721. https://doi.org/10.1007/s00442-012-2346-4
Oja T, Talve T (2012) Genetic diversity and differentiation in six species of the genus Rhinanthus (Orobanchaceae). Pl Syst Evol 298:901–911. https://doi.org/10.1007/s00606-012-0599-3
Paun O, Bateman RM, Fay MF, Hedrén M, Civeyrel L, Chase MW (2010) Stable epigenetic effects and adaptation in allopolyploid orchids. Molec Biol Evol 27:2465–2473. https://doi.org/10.1093/molbev/msq150
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research—an update. Bioinformatics 28:2537–2539. https://doi.org/10.1093/bioinformatics/bts460
Pettersson B (1958) Dynamik och konstans i Gotlands flora och vegetation. Acta Phytogeographica Suecica, vol. 40. Almquist & Wiksell, Uppsala
Pikaard CS, Scheid OM (2014) Epigenetic regulation in plants. Cold Spring Harb Perspect Biol 6:a019315. https://doi.org/10.1101/cshperspect.a019315
Pleines T, Esfeld K, Blattner FR, Thiv M (2013) Ecotypes and genetic structure of Rhinanthus alectorolophus (Orobanchaceae) in southwestern Germany. Pl Syst Evol 299:1523–1535. https://doi.org/10.1007/s00606-013-0816-8
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Rohlf FJ (1994) NTSYS-pc, Numerical taxonomy and multivariate analysis system, version 1.80. Exeter Software, Setauket
Ronniger K (1934) Auffindung einer neuen Rasse des Rhinanthus rumelicus Vel. auf der Insel Ösel (Estland). Feddes Repert 35:97–99
Rosenberg NA (2004) Distruct: a program for the graphical display of population structure. Molec Ecol Notes 4:137–138. https://doi.org/10.1046/j.1471-8286.2003.00566.x
Rousset F (1996) Equilibrium values of measures of population subdivision for stepwise mutation processes. Genetics 142:1357–1362
Sjörs H (1956) Nordisk växtgeografi. Bonnier, Stockholm
Slatkin M (1995) A measure of population subdivision based on microsatellite allele frequencies. Genetics 139:457–462
Storfer A (1996) Quantitative genetics: a promising approach for the assessment of genetic variation in endangered species. Trends Ecol Evol 11:343–347. https://doi.org/10.1016/0169-5347(96)20051-5
Stuessy TF, Crawford DJ, Soltis DE, Soltis PS (2014) Plant systematics: the origin, interpretation, and ordering of plant biodiversity. Koeltz Scientific Books, Königstein
Světlíková P, Hájek T, Těšitel J (2015) Hydathode trichomes actively secreting water from leaves play a key role in the physiology and evolution of root-parasitic rhinanthoid Orobanchaceae. Ann Bot (Oxford) 116:61–68. https://doi.org/10.1093/aob/mcv065
Talve T, McGlaughlin ME, Helenurm K, Wallace LE, Oja T (2014a) Population genetic diversity and species relationships in the genus Rhinanthus L. based on microsatellite markers. Pl Biol (Stuttgart) 16:495–502. https://doi.org/10.1111/plb.12057
Talve T, Mürk M, Lindell T, Oja T (2014b) Rhinanthus plants found on Gotland (Sweden): are they related to Rhinanthus osiliensis from Saaremaa (Estonia)? Biochem Syst Ecol 54:113–122. https://doi.org/10.1016/j.bse.2014.01.006
ter Borg SJ (1972) Variability of Rhinanthus serotinus (Schönh.) Oborny in relation to the environment. PhD Thesis, University of Groningen, Groningene
Turesson G (1922) The genotypical response of the plant species to the habitat. Hereditas 3:211–350. https://doi.org/10.1111/j.1601-5223.1922.tb02734.x
von Sterneck J (1901) Monographie der Gattung Alectorolophus. Abh K K Zool-Bot Ges Wien 1:1–150
Vrancken J, Brochmann C, Wesselingh RA (2012) A European phylogeography of Rhinanthus minor compared to Rhinanthus angustifolius: unexpected splits and signs of hybridization. Ecol Evol 2:1531–1548. https://doi.org/10.1002/ece3.276
Wesselingh RA (2016) Within-population variation in the relation between node number and flowering time in Rhinanthus angustifolius (Orobanchaceae). Pl Ecol Evol 149:21–30
Young A, Boyle T, Brown T (1996) The population genetic consequences of habitat fragmentation for plants. Trends Ecol Evol 11:413–418. https://doi.org/10.1016/0169-5347(96)10045-8
Zhang Y-Y, Latzel V, Fisher M, Bossdorf O (2018) Understanding the evolutionary potential of epigenetic variation: a comparison of heritable phenotypic variation in rpiRILs, RILs, and natural ecotypes of Arabidopsis thaliana. Heredity 121:257–265. https://doi.org/10.1038/s41437-018-0095-9
Zopfi H-J (1993a) Ecotypic variation in Rhinanthus alectorolophus (Scopoli) Pollich (Scrophulariaceae) in relation to grassland management. I. Morphological delimitations and habitats of seasonal ecotypes. Flora 188:15–39. https://doi.org/10.1016/S0367-2530(17)32244-2
Zopfi H-J (1993b) Ecotypic variation in Rhinanthus alectorolophus (Scopoli) Pollich (Scrophulariaceae) in relation to grassland management. II. The genotypic basis of seasonal ecotypes. Flora 188:153–173. https://doi.org/10.1016/S0367-2530(17)32261-2
Zopfi H-J (1995) Life history variation and intraspecific heterochrony in Rhinanthus glacialis (Scrophulariaceae). Pl Syst Evol 198:209–233. https://doi.org/10.1007/BF00984738
Acknowledgements
Open access funding provided by Lund University. We are grateful to Margarita Mürk who helped with the fieldwork, to Joel Widéen who assisted with the morphometric measurements, to Anupong Saenruen and Nina Strömberg who assisted in the laboratory, and to Renate Wesselingh and two anonymous reviewers, who provided valuable comments on previous versions of the manuscript. We also thank Jörgen Petersson for information about Rhinanthus populations on Gotland. The work was supported by Elly Olssons minnesfond (to AJ and SA), Anna och Svante Murbecks minnesfond and Hallströms minnesfond (to AJ) and the European Regional Development Fund (Centre of Excellence EcolChange; to TO). Seeds of the host species used in common-garden experiments were donated by Weibulls Horto.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Handling Editor: Pablo Vargas.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
606_2020_1662_MOESM1_ESM.pdf
Supplementary material 1 (see Table 1) (PDF 91 kb)
Information on Electronic Supplementary Material
Information on Electronic Supplementary Material
Online Resource 1. Genetic distances (pairwise FST-values) based on 11 microsatellite loci and geographical distances in km between Rhinanthus populations from Gotland (see Table 1).
Online Resource 2. Genetic distances (pairwise FST-values) based on 8 microsatellite loci and geographical distances in km between Rhinanthus populations from Sweden and Estonia.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Jonstrup, A., Hedrén, M., Oja, T. et al. The evolution of spring fen ecotypes in Rhinanthus: genetic evidence for parallel origins in Scandinavia after the last ice age. Plant Syst Evol 306, 35 (2020). https://doi.org/10.1007/s00606-020-01662-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00606-020-01662-y