Abstract
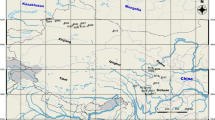
Litsea szemaois (Lauraceae) is an endemic and endangered species from the tropical rain forests of Xishuangbanna, southern Yunnan, SW China, but habitat fragmentation, especially exacerbated by rubber planting, has caused a decline in population size of the species. AFLP and ISSR were used to investigate the genetic diversity and population structure of eight populations from across its known distribution. Three AFLP and ten ISSR primer combinations produced a total of 203 and 77 unambiguous and repeatable bands respectively, of which 164 (80.8%) and 67 (87.0%) were polymorphic for the two markers. These two markers showed that Litsea szemaois exhibits comparatively high genetic diversity at species level (heterozygosity (hs) = 0.2109) relative to some other Lauraceae. Most of the genetic variation was partitioned within populations, but genetic differentiation between populations was significant and relatively high (Φ st = 0.2420, θB = 0.1986) compared with other tropical plants. The genetic characteristics of L. szemaois may be related to its outbreeding system, insect pollination and fragmented distribution. Because L. szemaois is dioecious and slow to mature, ex situ conservation across its genetic diversity is unlikely to succeed, although seedlings grow well under cultivation. Thus, in situ conservation is very important for this endangered species, especially as only 133 adult individuals are known in the wild. In particular, the Nabanhe and Mandian populations should be given a high conservation priority due to their higher genetic diversity, larger population size and better field condition, but wider sampling is required across all populations to determine additional areas with significant genetic conservation value.


Similar content being viewed by others
References
Allen CK (1938) Studies in the Lauraceae. I. Chinese and Indo-Chinese species of Litsea, Neolitsea and Actinodaphne. Ann Missouri Bot Gard 25:361–434
Barrett SCH, Kohn JS (1991) Genetic and evolutionary consequences of small population size in plants: implications for conservation. In: Falk DA, Holsinger KE (eds) Genetics and conservation of rare plants. Oxford University Press, New York, pp 3–30
Bensch S, Åkesson M (2005). Ten years of AFLP in ecology and evolution: why so few animals? Molec Ecol 14:2899–2914
Bussell JD (1999) The distribution of random amplified polymorphic DNA (RAPD) diversity amongst populations of Isotoma petraea (Lobelaceae). Molec Ecol 8:775–789
Cardoso MA, Provan J, Powell W, Ferreira PCG, Oliveira DE (1998) High genetic differentiation among remnant populations of the endangered Caesalpinia echinata Lam. (Leguminosae-Caesalpinioideae). Molec Ecol 7:601–608
Chung MG, Chung MY, Son OHG, Epperson BK (2000) Spatial genetic struture in a Neolitsea sericea population (Lauraceae). Heredity 85:490–497
Chung MY, Nason JD, Epperson BK, Chung MG (2003) Temporal aspects of the fine-scale genetic structure in a population of Cinnamomum insularimontanum (Lauraceae). Heredity 90:98–108
Devy MS, Davidar P (2003) Pollination systems of trees in Kakachi, a mid-elevation wet evergreen forest in Western Ghats, India. Amer J Bot 90:650–657
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull 19:11–15
Ellis JR, Pashiey CH, Burke JM, Mccauley DE (2006) High genetic diversity in a rare and endangered sunflower as compared to a common congener. Molec Ecol 15:2345–2355
Excoffier L, Smouse PE, Quattro JM (1992) Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics 131:479–491
Falk DA, Holsinger KE (1991) Genetics and conservation of rare plants. Oxford University Press, New York
Fan Y-B, Wang R-M, Pan F-J, Yang P-S (2006) Study of the floral and pollination biology of Cinamomum camphora and Litsea cubeba (Lauraceae). J Taiwan Mus 59:75–90
Frankham R, Ballou JD, Briscoe DA (2002) Introduction to conservation genetics. Cambridge University Press, Cambridge
Glover BJ, Abbott RJ (1995) Low genetic diversity in the Scottish endemic Primula scotica Hook. New Phytol 129:147–153
Hamrick JL (1990) Isozymes and the analysis of genetic structure in plant populations. In: Soltis ED, Soltis PS (eds) Isozymes in plant biology. Chapman and Hall, London, pp 87–105
Hamrick JL, Godt MJW (1989) Allozyme diversity in plant species. In: Brown ADH, Clegg MT, Kahler AL, Weir BS (eds) Plant population genetics, breeding and genetic resources. Sinauer, Sunderland, pp 43–63
Hamrick JL, Godt MJW (1996) Effects of life history traits on genetic diversity in plant species. Philos Trans Roy Soc Lond B 351:1291–1298
Hamrick JL, Loveless MD (1989) The genetic structure of tropical tree populations: Associations with reproductive biology. In: Bock JH, Linhart YB (eds) Plant evolutionary ecology. Westview Press, Boulder Colo, pp 131–146
Hamrick JL, Godt MJW, Murawski DA, Loveless MD (1991) Correlations between species traits and allozyme diversity: implications for conservation biology. In: Falk DA, Holsinger KE (eds) Genetics and conservation of rare plants. Oxford University Press, New York, p 5–86
Hogbin PM, Peakall R (1999) Evaluation of the contribution of genetic research to the management of the endangered plant Zieria prostrata. Conserv Biol 13:514–522
Holsinger KE (1999) Analysis of genetic diversity in geographically structured population: a Bayesian perspective. Hereditas 130:245–255
Holsinger KE, Gottlieb LD (1991) Conservation of rare and endangered plants: principles and prospects. In: Falk DA, Holsinger KE (eds) Genetics and conservation of rare plants. Oxford University Press, New York, pp 195–208
Holsinger KE, Lewis PO (2006) HICKORY version 1.0.4. Department of Ecology and Evolutionary Biology, University of Connecticut, USA (Program available from http://darwin.eeb.uconn.edu/hickory/software.html)
Holsinger KE, Wallace LE (2004) Bayesian approaches for the analysis of population genetic structure: an example from Platanthera leucophaea (Orchidaceae). Molec Ecol 13:887–897
Holsinger KE, Lewis PO, Dey DK (2002) A Bayesian approach to inferring population structure from dominant markers. Molec Ecol 11:1157–1164
Jones CJ, Dwards KJ, Castaglione S, Sala F, van de Wiel C, Bredemeijer G, Vosman B, Matthes M, Daly A, Brettschneider R, Bettini P, Buiatti M, Maestri E, Malcevschi A, Marmiroli N, Aert R, Volckaert G, Rueda J, Linacero R, Vazquez A, Karp A (1997) Reproducibility testing of RAPD, AFLP and SSR markers in plants by a network of European laboratories. Molec Breed 3:381–390
Kam TL, Rob M, Tien YC, Edward T (2004) A comparison of AFLP and ERIC-PCR analyses for discriminating Escherichia coli from cattle, pig and human sources. FEMS Microbiol Ecol 47:111–119
Kang M, Ye Q-G, Huang H-W (2005) Genetic consequence of restricted habitat and population decline in endangered Isoetes sinensis (Isoetaceae). Ann Bot 96:1265–1274
Karron JD (1991) Patterns of genetic variation and breeding system in rare plant species. In: Falk DA, Holsinger KE (eds) Genetics and conservation of rare plants. Oxford University Press, New York, pp 87–98
Li J-M, Jin Z-X (2006) High genetic differentiation revealed by RAPD analysis of narrowly endemic Sinocalycanthus chinensis Cheng et S.Y. Chang, an endangered species of China. Biochem Syst Ecol 34:725–735
Li J, Li H-W (2006) Notes on the plants of genus Litsea (Lauraceae) from China. Act Bot Yun 28:103–107
Li H-W, Pai P-Y, Lee S-K, Wei F-N, Wei Y-T, Yang Y-C, Huang P-H, Tsui H-P, Shia Z-D, Li J-L (1984) Lauraceae. In: Li H-W (eds) Flora reipublicae popularis sinicae, vol 31. Science Press, Beijing, pp 1–463
Li J, Christophel DC, Conran JG, Li H-W (2004) Phylogenetic relationships within the Litsea complex (Lauraceae) inferred from sequences of the chloroplast gene matK and nuclear ribosomal DNA ITS regions. Pl Syst Evol 246:19–34
Li H-M, Aide TM, Ma Y-X, Liu W-J, Cao M (2007) Demand for rubber is causing the loss of high diversity rain forest in SW China. Biodivers Conserv 16:1731–1745
Liou H (1934) Lauracées de Chine et D’Indochine. Hermann & Gie, Éditeurs, Paris
Liu W-J, Ma Y-X, Hu H-B, Cao M, Wang W (2005) Land use and land cover change in the tropical rainforest region of southern Yunnan - a case study of Menglun, Xishuangbanna. J Mt Sci 23:71–79
Loveless MD (1992) Isozyme variation in tropical trees: patterns of genetic organization. New For 6:67–94
Loveless MD, Hamrick JL (1984) Ecological determinants of genetic structure in plant populations. Annual Rev Ecol Syst 15:65–95
Masayuki M (2003) Population genetics of threatened wild plants in Japan. J Pl Res 116:169–174
Miller MP (1997) Tools for population genetic analysis (TFPGA), version 1.3: a Windows program for the analysis of allozyme and molecular population genetic data. Department of Biological Sciences Northern Arizona University, USA (Program available from: http://www.marksgeneticsoftware.net/tfpga.htm)
Moyle LC (2006) Correlates of genetic differentiation and isolation by distance in 17 congeneric Silene species. Molec Ecol 15:1067–1081
Myers N, Mittermeier RA, Mittermeier CG, da Fonseca GAB, Kent J (2000) Biodiversity hotspots and conservation priorities. Nature 403:853–858
Neigel JE (1996) Estimation of effective population size and migration parameters from genetic data. In: Smith TB, Wayne RK (eds) Molecular genetic approaches in conservation. Oxford University Press, New York, pp 329–346
Nybom H (2004) Comparison of different nuclear DNA markers for estimating intraspecific genetic diversity in plants. Molec Ecol 13:1143–1155
Ranker TA (1994) Evolution of high genetic variability in the rare Hawaiian fern Adenophorus periens and implications for conservation management. Biol Conserv 70:19–24
Rohwer JG (1993) Lauraceae. In: Kubitzki K, Rohwer JG, Bittrich V (eds) The families and genera of vascular plants, vol 2. Springer-Verlag, Berlin, pp 366–391
Rolf JF (2000) NTSYS-pc. Numerical taxonomy and multi-variate analysis system, version 2.1. Exeter Software, Setauket
Rousset F (1997) Genetic differentiation and estimation of gene flow from F-Statistics under Isolation by distance. Genetics 145:1219–1228
Schaal BA, Leverich WJ, Rogstad SH (1991) Comparison of methods for assessing genetic variation in plant conservation biology. In: Falk DA, Holsinger KE (eds) Genetics and conservation of rare plants. Oxford University Press, New York, pp 123–134
Schmidt K, Jensen K (2000) Genetic structure and AFLP variation of remnant populations in the rare plant Pedicularis palustris (Scrophulariacae) and its relation to population size and reproductive components. Amer J Bot 87:678–689
SPSS (2005) Statistical software package, Version 13.0. SPSS Inc, Chicago
Suyama Y, Obayashi K, Hayashi I (2000) Clonal structure in a dwarf bamboo (Sasa senanensis) population inferred from amplified fragment length polymorphism (AFLP) fingerprints. Molec Ecol 9:901–906
Tero N, Aspi J, Siikamäki P, Jäkäläniemi A, Tuomi J (2003) Genetic structure and gene flow in a metapopulation of an endangered plant species, Silene tatarica. Molec Ecol 12:2073–2085
Vos P, Hogers R, Bleeker M, Reijand M, van de Lee T, Hornes M, Frijers A, Pot J, Peleman J, Kuiper M, Zabeau M (1995) AFLP: a new technique for DNA fingerprinting. Nucleic Acids Res 23:4407–4414
Wang S, Xie Y (eds) (2003) China species red list. High Education Press, Beijing, pp 330–334
Wang Z-F, Gao S-H, Tian S-N, Fu S-L, Ren H, Peng S-L (2005a) Genetic structure of Cryptocarya chinensis in fragmented lower subtropical forests in China based on ISSR markers. Biodivers Sci 13:324–331
Wang Z-S, An S-Q, Liu H, Leng X, Zheng J-W, Liu Y-H (2005b) Genetic structure of the endangered plant Neolitsea sericea (Lauraceae) from the Zhoushan Archipelago using RAPD markers. Ann Bot 95:305–313
Wright S (1978) Evolution and the genetics of populations, variability within and among natural populations, vol 4. University of Chicago Press, Chicago
Zabeau M, Vos P (1993) European Patent Application Number: 92402629 7 (Publication Number: 0534858 A)
Zhang F-M (2002) DCFA 1.1, a program companied with AMOVA to compute the matrix of distance. Laboratory of Systematics and Evolutionary Botany, Institute of Botany, the Chinese Academy of Sciences, Beijing
Zietkiewicz E, Rafalski A, Labuda D (1994) Genome fingerprinting by simple sequence repeats (SSR)-anchored PCR amplification. Genomics 20:176–183
Zhu H (2004) A tropical seasonal rain forest at its altitudinal and latitudinal limits in southern Yunnan, SW China. Gard Bull Singap 56:55–72
Acknowledgments
Guo-Da Tao and Hsi-Wen Li are thanked for their assistance with field collection, and we are indebted to Fu-Min Zhang for access to the software DCFA 1.1. We are also grateful to Song Ge, Yun-Qian Hu, Long-Qian Xiao, Hong-Mei Li, Shu-Li Wang and Zhi-Ming Li for their helpful suggestions, and Barbara Rudolph and Kristina Groth from University of Hamburg for comments and changes to an earlier manuscript. John Conran from The University of Adelaide is thanked for assistance with the current manuscript. The project “Conservation Genetics of Litsea szemaois” from the program “the Light of Western China” of CAS, and The National Natural Science Foundation of China (grant number 30470123) jointly funded this work.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ci, Xq., Chen, Jq., Li, Qm. et al. AFLP and ISSR analysis reveals high genetic variation and inter-population differentiation in fragmented populations of the endangered Litsea szemaois (Lauraceae) from Southwest China. Plant Syst Evol 273, 237–246 (2008). https://doi.org/10.1007/s00606-008-0012-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00606-008-0012-4