Abstract
A “signal-off” photoelectrochemical (PEC) sensing platform has been designed for the ultrasensitive detection of DNA methylation levels and multiple methylated sites. The platform employs tungsten trioxide and TpPa-1-COF loaded by gold nanoparticle (AuNPs@WO3@TpPa-1-COF) composite material as the photoactive component and p-type reduced graphene (rGO) as an efficient quencher. The PEC signal of AuNPs@WO3@TpPa-1-COF composite is effectively quenched in the presence of p-type rGO, because p-type rGO can compete with AuNPs@WO3@TpPa-1-COF to deplete light energy and electron donors. In addition, a hybrid strand reaction (HCR) amplification strategy fixes more target DNA and then combines with rGO-modified anti-5-methylcytosine antibody to facilitate ultrasensitive DNA methylation detection. Under optimal conditions, DNA methylation can be measured within a linear concentration range of 10–14 to 10–8 M, with an exceptionally low detection limit of 0.19 fM (S/N = 3). At the same time, the platform can conduct quantitative determination of multi-site methylation, with the linear equation △I = 44.19LogA + 61.43, and the maximum number of methylation sites is 5. The sensor demonstrates high sensitivity, excellent selectivity, and satisfactory stability. Furthermore, the proposed signal-off PEC strategy was successfully employed to detect DNA methylation in spiked human serum samples, with recoveries ranging from 93.17 to 107.28% and relative standard deviation (RSD) ranging from 1.15 to 5.49%.
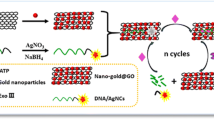
Graphical Abstract








Similar content being viewed by others
Data availability
The data sets supporting the results of this article are included within the article and its Supplementary Information.
References
Schubeler D (2015) Function and information content of DNA methylation. Nature 517:321–326. https://doi.org/10.1038/nature14192
Li SZ, Tollefsbol TO (2020) DNA methylation methods: global DNA methylation and methylomic analyses. Methods 187:28–43. https://doi.org/10.1016/j.ymeth.2020.10.002
Veland N, Hardikar S, Zhong Y, Gayatri S, Dan J, Strahl BD, Rothbart SB, Bedford MT, Chen TP (2017) The arginine methyltransferase PRMT6 regulates DNA methylation and contributes to global DNA hypomethylation in cancer. Cell Rep 21(12):3390–3397. https://doi.org/10.1016/j.celrep.2017.11.082
Wei W, Gao CY, Xiong YX, Zhang YJ, Liu SQ, Pu YP (2015) A fluorescence method for detection of DNA and DNA methylation based on graphene oxide and restriction endonuclease HpaII. Talanta 131:342–347. https://doi.org/10.1016/j.talanta.2014.07.094
Wang J, Yang J, Li DD (1875) Li JN (2021) Technologies for targeting DNA methylation modifications: basic mechanism and potential application in cancer. Biochim Biophys Acta Rev Cancer 1:188454. https://doi.org/10.1016/j.bbcan.2020.188454
Yan Q, Tang Y, He F, Xue J, Zhou RS, Zhang XY, Luo HY, Zhou DH, Wang XW (2021) Global analysis of DNA methylation in hepatocellular carcinoma via a whole-genome bisulfite sequencing approach 113(6):3618–3634. https://doi.org/10.1016/j.ygeno.2021.08.024
Dong NN, Wang WJ, Qin YR, Wang Y, Shan HB (2022) Sensitive lateral flow assay for bisulfite-free DNA methylation detection based on the restriction endonuclease GlaI and rolling circle amplification 1227:340307. https://doi.org/10.1016/j.aca.2022.340307
Li X, Franke AA (2011) High-throughput and cost-effective global DNA methylation assay by liquid chromatography-mass spectrometry. Anal Chim Acta 703(1):58–63. https://doi.org/10.1016/j.aca.2011.07.014
Feng QM, Wang MY, Qin L, Wang P (2019) Dual-signal readout of DNA methylation status based on the assembly of a supersandwich electrochemical biosensor without enzymatic reaction. ACS Sens 4:2615–2622. https://doi.org/10.1021/acssensors.9b00720
Chen DF, Wu YF, Tilley RD, Gooding JJ (2022) Rapid and ultrasensitive electrochemical detection of DNA methylation for ovarian cancer diagnosis. Biosens Bioelectron 206:114126. https://doi.org/10.1016/j.bios.2022.114126
Li MJ, Zheng YN, Liang WB, Yuan R, Chai YQ (2017) Using p-type PbS quantum dots to quench photocurrent of fullerene-Au NP@MoS2 composite structure for ultrasensitive photoelectrochemical detection of ATP. ACS Appl Mater Interfaces 9(48):42111–42120. https://doi.org/10.1021/acsami.7b13894
Hu T, Zheng YN, Li MJ, Liang WB, Chai YQ, Yuan R (2018) A highly sensitive photoelectrochemical assay with donor-acceptor-type material as photoactive material and polyaniline as signal enhancer. Anal Chem 90(10):6096–6101. https://doi.org/10.1021/acs.analchem.8b00093
Yang H, Shen HR, Qileng A, Cui GH, Liang ZQ, Liu YJ, Liu WP (2023) Well-aligned track-accelerated tripedal DNA walker for photoelectrochemical recognition of dual-miRNAs based on molecular logic gates. Anal Chem 95:5764–5772. https://doi.org/10.1021/acs.analchem.3c00027
Liu XP, Chen JS, Mao CJ, Niu HL, Song JM, Bao-Kang Jin BK (2018) A label-free photoelectrochemical biosensor for urokinase-type plasminogen activator detection based on a g-C3N4/CdS nanocomposite. Anal Chim Acta 1025:99–107. https://doi.org/10.1016/j.aca.2018.04.051
Yan PC, Jiang DH, Tian YH, Xu L, Qian JC, Li HN, Xia JX, Li HM (2018) A sensitive signal-on photoelectrochemical sensor for tetracycline determination using visible-light-driven flower-like CN/BiOBr composites. Biosens Bioelectron 111:74–81. https://doi.org/10.1016/j.bios.2018.03.054
Qian YR, Feng JH, Fan DW, Zhang Y, Kuang X, Wang H, Qin Wei Q, Ju HX (2019) A sandwich-type photoelectrochemical immunosensor for NT-pro BNP detection based on F-Bi2WO6/Ag2S and GO/PDA for signal amplification. Biosens Bioelectron 131:299–309. https://doi.org/10.1016/j.bios.2019.02.029
Zeng RJ, Gong HX, Li YL, Li YX, Lin W, Tang DP, Knopp D (2022) CRISPR-cas12a-derived photoelectrochemical biosensor for point of-care diagnosis of nucleic acid. Anal Chem 94:7442–7448. https://doi.org/10.1021/acs.analchem.2c01373
Lu JY, Wang MH, Han YW, Deng Y, Zeng YJ, Li C, Yang J, Li GX (2022) Functionalization of covalent organic frameworks with DNA via covalent modification and the application to exosomes detection. Anal Chem 94:5055–5061. https://doi.org/10.1021/acs.analchem.1c05222
Wang L, Xie HJ, Lin YX, Wang MH, Sha LJ, Yu XM, Yang J, Zhao J, Li GX (2022) Covalent organic frameworks (COFs)-based biosensors for the assay of disease biomarkers with clinical applications. Biosens Bioelectron 217:114668. https://doi.org/10.1016/j.bios.2022.114668
Pérez-Carvajal J, Boix G, Imaz I, Maspoch D (2019) The imine-based COF TpPa-1 as an efficient cooling adsorbent that can be regenerated by heat or light. Adv Energy Mater 9:1901535. https://doi.org/10.1002/aenm.201901535
Zheng JT, Huang SY, Tong YJ, Wei SB, Chen GS, Huang SM, Ouyang JF (2020) In-situ layer-by-layer synthesized TpPa-1 COF solid-phase microextraction fifiber for detecting sex hormones in serum. Anal Chim Acta 1137:28–36. https://doi.org/10.1016/j.aca.2020.08.047
Dang XM, Zhao HM (2021) Signal amplifified sandwich-type photoelectrochemical sensing assay based on rGO-Znln2S4 functionalized Au-WO3 IOPCs Z-scheme heterojunction. Electrochim Acta 365:137382. https://doi.org/10.1016/j.electacta.2020.137382
Wang HY, Wang M, Wang H, Ren X, Wang H, Wei Q, Wu D (2022) Detection of NSE by a photoelectrochemical self-powered immunosensor integrating RGO photocathode and WO3/Mn:CdS nanomaterial photoanode. Biosens Bioelectron 207:114196. https://doi.org/10.1016/j.bios.2022.114196
Pei FB, Feng SS, Wu Y, Lv XC, Wang HL, Chen SM, Hao QL, Cao Y, Wu Lei W, Tong ZY (2021) Label-free photoelectrochemical immunosensor for aflatoxin B1 detection based on the Z-scheme heterojunction of g-C3N4/Au/WO3. Biosens Bioelectron 189:113373. https://doi.org/10.1016/j.bios.2021.113373
Yan H, Liu YH, Yang Y, Zhang HY, Liu XR, Wei JZ, Bai LL, Wang YW, Zhang FM (2022) Covalent organic framework based WO3@COF/rGO for efficient visible-light-driven H2 evolution by two-step separation mode. Chem Eng J 431:133404. https://doi.org/10.1016/j.cej.2021.133404
Sun L, Li LL, Fan JJ, Xu QL, Ma DK (2022) Construction of highly active WO3/TpPa-1-COF S-scheme heterojunction toward photocatalytic H2 generation. J Mater Sci Technol 123:41–48. https://doi.org/10.1016/j.jmst.2021.12.065
Deng XD, Chen MS, Fu Q, Smeets NMB, Xu F, Zhang ZY, Filipe CDM, Hoare T (2016) An highly-sensitive immunosorbent assay based on biotinylated graphene oxide and the quartz crystal microbalance. ACS Appl Mater Interfaces 8(3):1893–1902. https://doi.org/10.1021/acsami.5b10026
Saleem W, Salinas C, Watkins B, Garvey G, Sharma AC, Ghosh R (2016) Antibody functionalized graphene biosensor for label-free electrochemical immunosensing of fifibrinogen, an indicator of trauma induced coagulopathy. Biosens Bioelectron 86:522–529. https://doi.org/10.1016/j.bios.2016.07.009
Xu C, Shi X, Ji A, Shi L, Zhou C, Cui Y (2015) Fabrication and characteristics of reduced graphene oxide produced with different green reductants. PLoS ONE 10(12):e0144842. https://doi.org/10.1371/journal.pone.0144842
Sahoo DP, Patnaik S, Parida K (2019) Construction of a z-scheme dictated WO3-X/Ag/ZnCr LDH synergistically visible light-induced photocatalyst towards tetracycline degradation and H2 evolution. ACS Omega 4(12):14721–14741. https://doi.org/10.1021/acsomega.9b01146
Funding
Financial support for this work was provided by the National Natural Science Foundation of China (No. 82272430, No. 81873982, and No. 82003109), Special Project of Scientific and Technological Innovation Capability Improvement of the Army Military Medical University (No. 2022XQN04 and No. 2021XQN03), and Special Support from the Chongqing Postdoctoral Research Project (No. 2021XM1018).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, H., Li, C., Wang, L. et al. Photoelectrochemical sensor based on AuNPs@WO3@TpPa-1-COF for quantification of DNA methylation levels. Microchim Acta 191, 167 (2024). https://doi.org/10.1007/s00604-024-06235-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00604-024-06235-1