Abstract
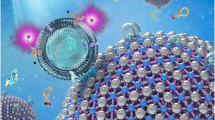
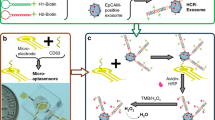
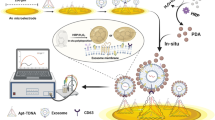
Programed cell death ligand 1 (PD-L1) is a protein biomarker overexpressed on exosomes derived from tumor cells. It plays an important role in tumor diagnosis, screening, evaluation of therapeutic efficacy, and prognosis. In this study, a facile method is presented to detect PD-L1-overexpressing cancer exosomes with high specificity and sensitivity. First, gold nanospheres (GNSs) were attached to the bottom of an eight-well chambered slide by electrostatic adsorption, forming the detection substrate. Then, Cy5-labeled CD63 aptamers (i.e., the capture probes) were modified on the GNSs by Au–S bond. After adding samples containing target exosomes which were stained by membrane dyes DiI in advance, FAM-labeled PD-L1 aptamers (i.e., the immunoprobes) were added to recognize PD-L1 on the target exosomes. By triple-color fluorescence co-localization (TFC) of the Cy5, DiI, and FAM channels, highly sensitive and reliable detection of the PD-L1-overexpressing exosomes was achieved in the concentration range 7.78 × 101 to 7.78 × 104 particles/mL with a detection limit down to 6 particles/mL. The advantages of the proposed detection method include the following; first, the detection substrate is easy to prepare and convenient to clean. Second, the TFC strategy can completely exclude nonspecific reaction sites and thus significantly improves the accuracy. Such a facile and reliable detection method holds a great potential in exosome-based cancer theranostics.
Graphical abstract
In this paper, we proposed a triple-color fluorescence co-localization (TFC) strategy to significantly improve the reliability of exosome detection and the detection substrate is easy to prepare and convenient to clean. In addition, the LOD is down to 6 particles/mL, which is quite low compared with other detection methods







Similar content being viewed by others
References
Gould SJ, Raposo G (2013) As we wait: coping with an imperfect nomenclature for extracellular vesicles. J Extracell Vesicles 2:20389. https://doi.org/10.3402/jev.v2i0.20389
Kim DK, Lee J, Simpson RJ, Lotvall J, Gho YS (2015) EVpedia: a community web resource for prokaryotic and eukaryotic extracellular vesicles research. Semin Cell Dev Biol 40:4–7. https://doi.org/10.1016/j.semcdb.2015.02.005
Raposo G, Stoorvogel W (2013) Extracellular vesicles: exosomes, microvesicles, and friends. J Cell Biol 200(4):373–383. https://doi.org/10.1083/jcb.201211138
Tsatsaronis JA, Franch-Arroyo S, Resch U, Charpentier E (2018) Extracellular vesicle RNA: a universal mediator of microbial communication. Trends Microbiol 26(5):401–410. https://doi.org/10.1083/jcb.201211138
Yanez-Mo M, Siljander PR, Andreu Z, Zavec AB, Borras FE, Buzas EI, Buzas K, Casal E, Cappello F, Carvalho J, Colas E, Cordeiro-da Silva A, Fais S, Falcon-Perez JM, Ghobrial IM, Giebel B, Gimona M, Graner M, Gursel I, Gursel M, Heegaard NH, Hendrix A, Kierulf P, Kokubun K, Kosanovic M, Kralj-Iglic V, Kramer-Albers EM, Laitinen S, Lasser C, Lener T, Ligeti E, Line A, Lipps G, Llorente A, Lotvall J, Mancek-Keber M, Marcilla A, Mittelbrunn M, Nazarenko I, Nolte-’t Hoen EN, Nyman TA, O’Driscoll L, Olivan M, Oliveira C, Pallinger E, Del Portillo HA, Reventos J, Rigau M, Rohde E, Sammar M, Sanchez-Madrid F, Santarem N, Schallmoser K, Ostenfeld MS, Stoorvogel W, Stukelj R, Van der Grein SG, Vasconcelos MH, Wauben MH, De Wever O (2015) Biological properties of extracellular vesicles and their physiological functions. J Extracell Vesicles 4:41–60. https://doi.org/10.3402/jev.v4.27066
Vergauwen G, Dhondt B, Van Deun J, De Smedt E, Berx G, Timmerman E, Gevaert K, Miinalainen I, Cocquyt V, Braems G, Van den Broecke R, Denys H, De Wever O, Hendrix A (2017) Confounding factors of ultrafiltration and protein analysis in extracellular vesicle research. Sci Rep 7(1):2704. https://doi.org/10.1038/s41598-017-02599-y
Wu M, Chen Z, Xie Q, Xiao B, Zhou G, Chen G, Bian Z (2021) One-step quantification of salivary exosomes based on combined aptamer recognition and quantum dot signal amplification. Biosens Bioelectron 171:112733. https://doi.org/10.1016/j.bios.2020.112733
Zhang P, Zhou X, Zeng Y (2019) Multiplexed immunophenotyping of circulating exosomes on nano-engineered ExoProfile chip towards early diagnosis of cancer. Chem Sci 10(21):5495–5504. https://doi.org/10.1039/C9SC00961B
Lee J, Kwon Y, Jung J, Shin H, Park J (2021) Immunostaining extracellular vesicles based on an aqueous two-phase system: for analysis of tetraspanins. ACS Appl Bio Mater 4(4):3294–3303. https://doi.org/10.1021/acsabm.0c01625
Doyle LM, Wang MZ (2019) Overview of extracellular vesicles, their origin, composition, purpose, and methods for exosome isolation and analysis. Cells 8(7):727. https://doi.org/10.3390/cells8070727
Taylor DD, Gercel-Taylor C (2013) The origin, function, and diagnostic potential of RNA within extracellular vesicles present in human biological fluids. Front Genet 4:142. https://doi.org/10.3389/fgene.2013.00142
Suchorska WM, Lach MS (2016) The role of exosomes in tumor progression and metastasis (Review). Oncol Rep 35(3):1237–1244. https://doi.org/10.3892/or.2015.4507
An Y, Jin T, Zhu Y, Zhang F, He P (2019) An ultrasensitive electrochemical aptasensor for the determination of tumor exosomes based on click chemistry. Biosens Bioelectron 142:111503. https://doi.org/10.1016/j.bios.2019.111503
Azmi AS, Bao B, Sarkar FH (2013) Exosomes in cancer development, metastasis, and drug resistance: a comprehensive review. Cancer Metastasis Rev 32(3–4):623–642. https://doi.org/10.1007/s10555-013-9441-9
Shao H, Im H, Castro CM, Breakefield X, Weissleder R, Lee H (2018) New technologies for analysis of extracellular vesicles. Chem Rev 118(4):1917–1950. https://doi.org/10.1021/acs.chemrev.7b00534
Himes BT, Peterson TE, de Mooij T, Garcia LMC, Jung MY, Uhm S, Yan D, Tyson J, Jin-Lee HJ, Parney D, Abukhadra Y, Gustafson MP, Dietz AB, Johnson AJ, Dong H, Maus RL, Markovic S, Lucien F, Parney IF (2020) The role of extracellular vesicles and PD-L1 in glioblastoma-mediated immunosuppressive monocyte induction. Neuro Oncol 22(7):967–978. https://doi.org/10.1093/neuonc/noaa029
Li X, Zhong T, Tang R, Wu C, Xie Y, Liu F, Zhou Z (2020) PD-1 and PD-L1 expression in peripheral CD4/CD8+ T cells is restored in the partial remission phase in type 1 diabetes. J Clin Endocrinol Metab 105(6):1947–1956. https://doi.org/10.1210/clinem/dgaa130
Riazifar M, Mohammadi MR, Pone EJ, Yeri A, Lasser C, Segaliny AI, McIntyre LL, Shelke GV, Hutchins E, Hamamoto A, Calle EN, Crescitelli R, Liao W, Pham V, Yin Y, Jayaraman J, Lakey JRT, Walsh CM, Van Keuren-Jensen K, Lotvall J, Zhao W (2019) Stem cell-derived exosomes as nanotherapeutics for autoimmune and neurodegenerative disorders. ACS Nano 13(6):6670–6688. https://doi.org/10.1021/acsnano.9b01004
Khanova, M.Yu.;Grigoryev, E.V (2019) Roles of PD-1 and PD-L1 receptors in the development of systemic inflammatory response and immunoadjuvant therapy, Patologiya Krovoobrashcheniya i Kardiokhirurgiya 23(3):76–83, https://doi.org/10.21688/1681-3472-2019-3-76-83.
Carney RP, Hazari S, Rojalin T, Knudson A, Gao T, Tang Y, Liu R, Viitala T, Yliperttula M, Lam KS (2017) Targeting tumor-associated exosomes with integrin-binding peptides. Adv Biosyst 1(5):160038. https://doi.org/10.1002/adbi.201600038
Lachal R (2019) Proceedings of Reanimation 2019, the French Intensive Care Society International Congress. Ann Intensive Care 9(Suppl 1):1–153. https://doi.org/10.1186/s13613-018-0474-7
Liu C, Zeng X, An Z, Yang Y, Eisenbaum M, Gu X, Jornet JM, Dy GK, Reid ME, Gan Q, Wu Y (2018) Sensitive detection of exosomal proteins via a compact surface plasmon resonance biosensor for cancer diagnosis. ACS Sens 3(8):1471–1479. https://doi.org/10.1021/acssensors.8b00230
Lin B, Tian T, Lu Y, Liu D, Huang M, Zhu L, Zhu Z, Song Y, Yang C (2021) Tracing tumor-derived exosomal PD-L1 by dual-aptamer activated proximity-induced droplet digital PCR. Angew Chem Int Ed Engl 60(14):7582–7586. https://doi.org/10.1002/anie.202015628
Pang Y, Shi J, Yang X, Wang C, Sun Z, Xiao R (2020) Personalized detection of circling exosomal PD-L1 based on Fe3O4@TiO2 isolation and SERS immunoassay. Biosens Bioelectron 148:111800. https://doi.org/10.1016/j.bios.2019.111800
Yu J, Lin Y, Xiong X, Li K, Yao Z, Dong H, Jiang Z, Yu D, Yeung SJ, Zhang H (2019) Detection of Exosomal PD-L1 RNA in Saliva of Patients With Periodontitis. Front Genet 10:202. https://doi.org/10.3389/fgene.2019.00202
Li C, Li C, Zhi C, Liang W, Wang X, Chen X, Lv T, Shen Q, Song Y, Lin D, Liu H (2019) Clinical significance of PD-L1 expression in serum-derived exosomes in NSCLC patients. J Transl Med 17(1):355. https://doi.org/10.1186/s12967-019-2101-2
Zong S, Liu Y, Yang K, Yang Z, Wang Z, Cui Y (2021) Eliminating nonspecific binding sites for highly reliable immunoassay via super-resolution multicolor fluorescence colocalization. Nanoscale 13(13):6624–6634. https://doi.org/10.1039/d0nr08103e
Ruan Q, Shao L, Shu Y, Wang J, Wu H (2014) Growth of monodisperse gold nanospheres with diameters from 20 nm to 220 nm and their core/satellite nanostructures. Adv Opt Mater 2(1):65–73. https://doi.org/10.1002/adom.201300359
Ivanusic D, Madela K, Laue M, Denner J (2015) Three-dimensional imaging of CD63 recruitment at the virological synapse: HIV-1. AIDS Res Hum Retroviruses 31(6):579–580. https://doi.org/10.1089/aid.2014.0252
Jung S, Kim MJ, Sellaththurai S, Kim S, Lee S, Lee J (2021) Generation of cd63-deficient zebrafish to analyze the role of cd63 in viral infection. Fish Shellfish Immunol 111:152–159. https://doi.org/10.1016/j.fsi.2021.01.016
Green GN, Fang H, Lin RJ, Newton G, Mather M, Georgiou CD, Gennis RB (1988) The nucleotide sequence of the cyd locus encoding the two subunits of the cytochrome d terminal oxidase complex of Escherichia coli. J Biol Chem 263(26):13138–13143. https://doi.org/10.1016/S0021-9258(18)37682-8
Liddell S, Smith JR, Zhang X, Hall H, Swan A, Bacardit J, Mobasheri A (2012) Proteomic and bioinformatic analysis of hydrophobic membrane protein extracts reveals the presence of several novel CD antigens, glucose transporters and voltage gated anion channels in articular chondrocytes. Osteoarthritis Cartilage 20:S261. https://doi.org/10.1016/j.joca.2012.02.438
Sina AA, Vaidyanathan R, Dey S, Carrascosa LG, Shiddiky MJ, Trau M (2016) Real time and label free profiling of clinically relevant exosomes. Sci Rep 6:30460. https://doi.org/10.1038/srep30460
Moura SL, Martin CG, Marti M, Pividori MI (2020) Multiplex detection and characterization of breast cancer exosomes by magneto-actuated immunoassay. Talanta 211:120657. https://doi.org/10.1016/j.talanta.2019.120657
Lima Moura S, Marti M, Pividori MI (2020) Matrix effect in the isolation of breast cancer-derived nanovesicles by immunomagnetic separation and electrochemical immunosensing-a comparative study. Sensors (Basel) 20(4):965. https://doi.org/10.3390/s20040965
Wang Y, Luo D, Fang Y, Wu W, Wang Y, Xia Y, Wu F, Li C, Lan J, Chen J (2019) An aptasensor based on upconversion nanoparticles as LRET donors for the detection of exosomes. Sensors Actuators B: Chem 298:126900. https://doi.org/10.1016/j.snb.2019.126900
Wang L, Zeng L, Wang Y, Chen T, Chen W, Chen G, Li C, Chen J (2021) Electrochemical aptasensor based on multidirectional hybridization chain reaction for detection of tumorous exosomes. Sens Actuators B Chem 332:129471. https://doi.org/10.1016/j.snb.2021.129471
Sun Y, Jin H, Jiang X, Gui R (2020) Assembly of black phosphorus nanosheets and MOF to form functional hybrid thin-film for precise protein capture dual-signal and intrinsic self-calibration sensing of specific cancer-derived exosomes. Anal Chem 92(3):2866–2875. https://doi.org/10.1021/acs.analchem.9b05583
Zhu X, Liu Z, Li J, Li Z, Si F, Yang H, Kong J (2021) Dual signal amplification based on polysaccharide-initiated ring-opening polymerization and click polymerization for exosomes detection. Talanta 233:122531. https://doi.org/10.1016/j.talanta.2021.122531
Wang Y, Zhang K, Huang X, Qiao L, Liu B (2021) Mass spectrometry imaging of mass tag immunoassay enables the quantitative profiling of biomarkers from dozens of exosomes. Anal Chem 93(2):709–714. https://doi.org/10.1021/acs.analchem.0c03904
Xu L, Chopdat R, Li D, Al-Jamal KT (2020) Development of a simple, sensitive and selective colorimetric aptasensor for the detection of cancer-derived exosomes. Biosens Bioelectron 169:112576. https://doi.org/10.1016/j.bios.2020.112576
Liu H, Liu W, Jin G (2021) Detection of exosomes using total internal reflected imaging ellipsometry. Biosensors (Basel) 11(5):164. https://doi.org/10.3390/bios11050164
Li R, An Y, Jin T, Zhang F, He P (2021) Detection of MUC1 protein on tumor cells and their derived exosomes for breast cancer surveillance with an electrochemiluminescence aptasensor. J Electroanal Chem 882:115011. https://doi.org/10.1016/j.jelechem.2021.115011
Liu L, Shen Y, Zhu X, Lv R, Li S, Zhang Z, Shi YG, Tan L (2018) ERa is a negative regulator of PD-L1 gene transcription in breast cancer. Biochem Biophys Res Commun 505(1):157–161. https://doi.org/10.1016/j.bbrc.2018.09.005
Deng H, Kan A, Lyu N, Mu L, Han Y, Liu L, Zhang Y, Duan Y, Liao S, Li S, Xie Q, Gao T, Li Y, Zhang Z, Zhao M (2020) Dual vascular endothelial growth factor receptor and fibroblast growth factor receptor inhibition elicits antitumor immunity and enhances programmed cell death-1 checkpoint blockade in hepatocellular carcinoma. Liver Cancer 9(3):338–357. https://doi.org/10.1159/000505695
Funding
This work was supported by the Natural Science Foundation of China (NSFC) (62175027, 62175030), the Natural Science Foundation of Jiangsu Province (BK20201261), the Zhishan Scholars Program of Southeast University (2242021R41151) and the National Key Basic Research Program of China (2015CB352002).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wei, J., Zhu, K., Chen, Z. et al. Triple-color fluorescence co-localization of PD-L1-overexpressing cancer exosomes. Microchim Acta 189, 182 (2022). https://doi.org/10.1007/s00604-022-05278-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00604-022-05278-6