Abstract
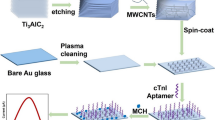
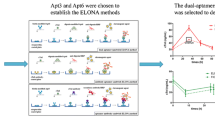
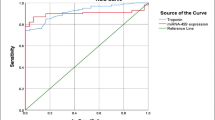
Circulating microRNAs are of diagnostic value for acute myocardial infarction (AMI). This study describes a fluorometric assay for multiplexed detection of the AMI biomarkers microRNA-499, microRNA-133a and microRNA-1. The assay involves the following two steps: (a) duplex-specific nuclease (DSN)-mediated signal amplification using aptamer-based sensor; (b) MoS2 nanosheets-based multiplexed fluorometric signal detection, and fluorometric signals have excitation/emission maxima at 492/518 nm for microRNA-499, 565/580 nm for microRNA-133a and 649/663 nm for microRNA-1. The assay has detection limits of around 100 fM for all three microRNAs. The assay is highly specific and rapid. It demonstrates that the expressions of microRNA-499, microRNA-133a and microRNA-1 are significantly higher in AMI patients. The ROC curves allow a clear distinction to be made between AMI group and non-AMI group.

Schematic representation of a multiplexed fluorometric method based on duplex-specific nuclease (DSN)-mediated signal amplification and MoS2 nanosheets-based fluorometric signal detection (DSN-MoS2) for rapid, sensitive and specific detection of microRNAs in AMI.





Similar content being viewed by others
References
Anderson JL, Morrow DA (2017) Acute myocardial infarction. N Engl J Med 376(21):2053–2064. https://doi.org/10.1056/NEJMra1606915
Roffi M, Patrono C, Collet JP, Mueller C, Valgimigli M, Andreotti F, Bax JJ, Borger MA, Brotons C, Chew DP, Gencer B, Hasenfuss G, Kjeldsen K, Lancellotti P, Landmesser U, Mehilli J, Mukherjee D, Storey RF, Windecker S (2016) 2015 ESC guidelines for the management of acute coronary syndromes in patients presenting without persistent ST-segment elevation: task force for the Management of Acute Coronary Syndromes in patients presenting without persistent ST-segment elevation of the European Society of Cardiology (ESC). Eur Heart J 37(3):267–315. https://doi.org/10.1093/eurheartj/ehv320
Nagele P, Brown F, Gage BF, Gibson DW, Miller JP, Jaffe AS, Apple FS, Scott MG (2013) High-sensitivity cardiac troponin T in prediction and diagnosis of myocardial infarction and long-term mortality after noncardiac surgery. Am Heart J 166(2):325–332.e321. https://doi.org/10.1016/j.ahj.2013.04.018
Shiozaki M, Inoue K, Suwa S, Lee CC, Chikata Y, Ishiura J, Kimura Y, Fukuda K, Tamura H, Fujiwara Y, Sumiyoshi M, Daida H (2017) Utility of the 0-hour/1-hour high-sensitivity cardiac troponin T algorithm in Asian patients with suspected non-ST elevation myocardial infarction. Int J Cardiol 249:32–35. https://doi.org/10.1016/j.ijcard.2017.09.009
Wang J, Chen J, Sen S (2016) MicroRNA as biomarkers and diagnostics. J Cell Physiol 231(1):25–30. https://doi.org/10.1002/jcp.25056
Berindan-Neagoe I, Monroig PC, Pasculli B, Calin GA (2014) MicroRNAome genome: a treasure for cancer diagnosis and therapy. CA Cancer J Clin 64(5):311–336. https://doi.org/10.3322/caac.21244
Kalla R, Ventham NT, Kennedy NA, Quintana JF, Nimmo ER, Buck AH, Satsangi J (2015) MicroRNAs: new players in IBD. Gut 64(3):504–517. https://doi.org/10.1136/gutjnl-2014-307891
Sita-Lumsden A, Dart DA, Waxman J, Bevan CL (2013) Circulating microRNAs as potential new biomarkers for prostate cancer. Br J Cancer 108(10):1925–1930. https://doi.org/10.1038/bjc.2013.192
Hsu A, Chen S-J, Chang Y-S, Chen H-C, Chu P-H (2014) Systemic approach to identify serum microRNAs as potential biomarkers for acute myocardial infarction. Biomed Res Int 2014:418628–418628. https://doi.org/10.1155/2014/418628
Kondkar AA, Abu-Amero KK (2015) Utility of circulating microRNAs as clinical biomarkers for cardiovascular diseases. Biomed Res Int 2015:821823–821823. https://doi.org/10.1155/2015/821823
Zhang L, Chen X, Su T, Li H, Huang Q, Wu D, Yang C, Han Z (2015) Circulating miR-499 are novel and sensitive biomarker of acute myocardial infarction. J Thorac Dis 7(3):303–308. https://doi.org/10.3978/j.issn.2072-1439.2015.02.05
Wang F, Long G, Zhao C, Li H, Chaugai S, Wang Y, Chen C, Wang DW (2013) Plasma microRNA-133a is a new marker for both acute myocardial infarction and underlying coronary artery stenosis. J Transl Med 11:222–222. https://doi.org/10.1186/1479-5876-11-222
Li J, Dong X, Wang Z, Wu J (2014) MicroRNA-1 in cardiac diseases and cancers. Korean J Physiol Pharmacol 18(5):359–363. https://doi.org/10.4196/kjpp.2014.18.5.359
Fiedler SD, Carletti MZ, Christenson LK (2010) Quantitative RT-PCR methods for mature microRNA expression analysis. Methods Mol Biol 630:49–64. https://doi.org/10.1007/978-1-60761-629-0_4
Aw SS, Tang MX, Teo YN, Cohen SM (2016) A conformation-induced fluorescence method for microRNA detection. Nucleic Acids Res 44(10):e92–e92. https://doi.org/10.1093/nar/gkw108
Sempere LF (2014) Fully automated fluorescence-based four-color multiplex assay for co-detection of microRNA and protein biomarkers in clinical tissue specimens. Methods Mol Biol 1211:151–170. https://doi.org/10.1007/978-1-4939-1459-3_13
Lan L, Chen D, Yao Y, Peng X, Wu J, Li Y, Ping J, Ying Y (2018) Phase-dependent fluorescence quenching efficiency of MoS2 Nanosheets and their applications in multiplex target biosensing. ACS Appl Mater Interfaces 10:42009–42017. https://doi.org/10.1021/acsami.8b15677
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116(2):281–297
Hoshida Y, Villanueva A, Kobayashi M, Peix J, Chiang DY, Camargo A, Gupta S, Moore J, Wrobel MJ, Lerner J, Reich M, Chan JA, Glickman JN, Ikeda K, Hashimoto M, Watanabe G, Daidone MG, Roayaie S, Schwartz M, Thung S, Salvesen HB, Gabriel S, Mazzaferro V, Bruix J, Friedman SL, Kumada H, Llovet JM, Golub TR (2008) Gene expression in fixed tissues and outcome in hepatocellular carcinoma. N Engl J Med 359(19):1995–2004. https://doi.org/10.1056/NEJMoa0804525
Chen X, Ba Y, Ma L, Cai X, Yin Y, Wang K, Guo J, Zhang Y, Chen J, Guo X, Li Q, Li X, Wang W, Zhang Y, Wang J, Jiang X, Xiang Y, Xu C, Zheng P, Zhang J, Li R, Zhang H, Shang X, Gong T, Ning G, Wang J, Zen K, Zhang J, Zhang CY (2008) Characterization of microRNAs in serum: a novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res 18(10):997–1006. https://doi.org/10.1038/cr.2008.282
Lawrie CH, Soneji S, Marafioti T, Cooper CD, Palazzo S, Paterson JC, Cattan H, Enver T, Mager R, Boultwood J, Wainscoat JS, Hatton CS (2007) MicroRNA expression distinguishes between germinal center B cell-like and activated B cell-like subtypes of diffuse large B cell lymphoma. Int J Cancer 121(5):1156–1161. https://doi.org/10.1002/ijc.22800
Gustafson D, Tyryshkin K, Renwick N (2016) microRNA-guided diagnostics in clinical samples. Best Pract Res Clin Endocrinol Metab 30(5):563–575. https://doi.org/10.1016/j.beem.2016.07.002
Wang Q, Ma J, Jiang Z, Wu F, Ping J, Ming L (2017) Identification of microRNAs as diagnostic biomarkers for acute myocardial infarction in Asian populations: a systematic review and meta-analysis. Medicine 96(24):e7173–e7173. https://doi.org/10.1097/MD.0000000000007173
Goretti E, Devaux Y (2016) Which future for circulating microRNAs as biomarkers of acute myocardial infarction? Ann Transl Med 4(21):440–440. https://doi.org/10.21037/atm.2016.11.21
Liu G, Niu X, Meng X, Zhang Z (2018) Sensitive miRNA markers for the detection and management of NSTEMI acute myocardial infarction patients. J Thorac Dis 10(6):3206–3215. https://doi.org/10.21037/jtd.2018.05.141
Varallyay E, Burgyan J, Havelda Z (2007) Detection of microRNAs by northern blot analyses using LNA probes. Methods (San Diego Calif) 43(2):140–145. https://doi.org/10.1016/j.ymeth.2007.04.004
Clancy E, Burke M, Arabkari V, Barry T, Kelly H, Dwyer RM, Kerin MJ, Smith TJ (2017) Amplification-free detection of microRNAs via a rapid microarray-based sandwich assay. Anal Bioanal Chem 409(14):3497–3505. https://doi.org/10.1007/s00216-017-0298-6
Kim KJ, Kwak J, Lee JH, Lee SS (2017) Real-time qRT-PCR assay for the detection of miRNAs using bi-directional extension sequences. Anal Biochem 536:32–35. https://doi.org/10.1016/j.ab.2017.08.006
Shagin DA, Rebrikov DV, Kozhemyako VB, Altshuler IM, Shcheglov AS, Zhulidov PA, Bogdanova EA, Staroverov DB, Rasskazov VA, Lukyanov S (2002) A novel method for SNP detection using a new duplex-specific nuclease from crab hepatopancreas. Genome Res 12(12):1935–1942. https://doi.org/10.1101/gr.547002
Zhulidov PA, Bogdanova EA, Shcheglov AS, Vagner LL, Khaspekov GL, Kozhemyako VB, Matz MV, Meleshkevitch E, Moroz LL, Lukyanov SA, Shagin DA (2004) Simple cDNA normalization using Kamchatka crab duplex-specific nuclease. Nucleic Acids Res 32(3):e37–e37. https://doi.org/10.1093/nar/gnh031
Wang Y, Howes PD, Kim E, Spicer CD, Thomas MR, Lin Y, Crowder SW, Pence IJ, Stevens MM (2018) Duplex-specific nuclease-amplified detection of MicroRNA using compact quantum dot-DNA conjugates. ACS Appl Mater Interfaces 10(34):28290–28300. https://doi.org/10.1021/acsami.8b07250
Xi Q, Zhou DM, Kan YY, Ge J, Wu ZK, Yu RQ, Jiang JH (2014) Highly sensitive and selective strategy for microRNA detection based on WS2 nanosheet mediated fluorescence quenching and duplex-specific nuclease signal amplification. Anal Chem 86(3):1361–1365. https://doi.org/10.1021/ac403944c
Xiao M, Man T, Zhu C, Pei H, Shi J, Li L, Qu X, Shen X, Li J (2018) MoS2 Nanoprobe for MicroRNA quantification based on duplex-specific nuclease signal amplification. ACS Appl Mater Interfaces 10(9):7852–7858. https://doi.org/10.1021/acsami.7b18984
Author contribution statement
Chengjian Yang and Zhijun Han design experiments; Yan Jin, Shuya Wang, Xiaoxiao Liu and Haohao Liu carry out experiments; Xue Zhu, Ke Wang and Peiling Zhou analyze experimental results; Xue Zhu and Ke Wang write the manuscript.
Funding
This work was supported by grant from Natural Science Foundation of Jiangsu Province (No. BK20171147), Jiangsu Young Medical Talents (QNRC2016167), Six talent peaks project in Jiangsu Province (WSW-047), The Science and Technology Projects of Wuxi City (WX18IVJN016) and The Major Science and Technology Projects of Wuxi City (Z201804, MS201653).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Declarations of interest
The authors declare that there is no conflict of interest regarding the publication of this paper.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 72 kb)
Rights and permissions
About this article
Cite this article
Zhu, X., Wang, K., Jin, Y. et al. Multiplexed fluorometric determination for three microRNAs in acute myocardial infarction by using duplex-specific nuclease and MoS2 nanosheets. Microchim Acta 187, 15 (2020). https://doi.org/10.1007/s00604-019-3896-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00604-019-3896-5