Abstract
Purpose
Low back pain (LBP), a widely prevalent and costly disease around the world, is mainly caused by intervertebral disc (IVD) degeneration (IDD). Although numerous factors may trigger this degenerative process, microbiome dysbiosis has recently been implicated as one of the likely causes. However, the exact relationship between the microbiome and IDD is not well understood. This review summarizes the potential mechanisms and discusses microbiome dysbiosis’s possible influence on IDD and LBP.
Methods
Prospective literature review.
Results
Alterations in microbiome composition and host responses to the microbiota causing pathological bone development and involution, led to the concept of gut-bone marrow axis and gut-bone axis. Moreover, the concept of the gut-disc axis was also proposed to explain the microbiome’s role in IDD and LBP. According to the existing evidence, the microbiome could be an important factor for inducing and aggravating IDD through changing or regulating the outside and inside microenvironment of the IVD. Three potential mechanisms by which the gut microbiota can induce IVD and cause LBP are: (1) translocation of the bacteria across the gut epithelial barrier and into the IVD, (2) regulation of the mucosal and systemic immune system, and (3) regulation of nutrient absorption and metabolites formation at the gut epithelium and its diffusion into the IVD. Furthermore, to investigate whether IVD is initiated by pathogenic bacteria and establish the correlation between the presence of certain microbial groups with the disease in question, microbiome diversity analysis based on16S rRNA data can be used to characterise stool/blood microbiota from IVD patients.
Conclusion
Future studies on microbiome, fungi and viruses in IDD is necessary to revolutionize our thinking about their possible role in the development of IVD diseases. Furthermore, we believe that inflammation inhibition and interruption of amplification of cascade reaction in IVD by targeting the gut and IVD microbiome is worthwhile for the treatment of IDD and LBP.
Level of Evidence I
Diagnostic: individual cross-sectional studies with the consistently applied reference standard and blinding.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Low back pain (LBP) is a leading cause of disability worldwide affecting millions of people in their day-to-day activities [1]. There are many potential causes of LBP, including mechanical stress, age, genetic factors, but intervertebral disc (IVD) degeneration (IDD) is regarded as one of the most likely causes [2]. One of the major concerns around IDD is [3], however the exact cause and nature of such inflammation are not clearly understood. One of the putative triggers of inflammation has been hypothesized to be infection of the IVDs by skin bacteria, especially P. acnes [4, 5]. Intriguingly, alterations in the microbiome (microbes in and on the human body) composition in the gastrointestinal system, skin and mouth are associated with the regulation of inflammation and autoimmunity in many other diseases, like rheumatoid arthritis, osteoarthritis, ankylosing spondylitis, septic arthritis and other infectious diseases [6,7,8,9,10,11]. However, it is not yet clear if such microbial changes play any role in IDD. The existing evidence strongly indicates that changes in the microbiome composition and associated metabolites might play a key role in the regulation and management of LBP in patients with IDD.
Human microbiome and IDD
The human microbiome is rapidly emerging as an important player in regulating our health and diseases owing to advances in sequence-based detection methods. Strategies such as metataxonomics are being used as a targeted approach to sequence 16S rRNA genes (encoding bacteria’s ribosomal subunits that include both conserved and variable regions) via probes or primers to detect and identify bacteria, and specify a phylum, a group, a genus or even a species [12]. Further understanding of the microbiome in terms of functional genes can be done using metagenomics that can analyse the composition, function and variations of the microbiome [13]. Compared to16S rRNA gene sequencing, metagenomic sequencing could allow deeper characterization of microorganism complexity and could identify a more accurate definition at the species level. The shotgun metagenomics could help in building a comprehensive description of the microbiome community [14].
Besides maintaining a stable microbial diversity, which in turn affects the characteristics of the host, microbiome can also affect the host phenotypes by secreting several microbial metabolites. Metabolomics can easily identify these markers and strengthen the early detection of diseases [15].
Recently, Rajasekaran et al. evaluated 24 lumbar IVDs and reported that the composition of the microbiome in healthy IVDs differed from those in the degenerated IVDs and herniated IVDs [16]. The study found Firmicutes, Actinobacteria and Saccharopolyspora to be abundant in normal IVD samples, which were related to the intestinal barrier function and antibacterial protection. On the other hand, few common human pathogens such as Bacillus coagulans and Bacillus clausii were detected in degenerate and herniated IVDs that have been implicated in spondylodiscitis, fracture, joint infections, meningitis and endocarditis. Based on the functional analysis, the microbiome can be distinguished as “good” or “bad” microbiotas. For instance, Saccharopolyspora, defined as “good microbiota” was found in healthy IVDs which produces macrolide antibiotics effectively against most gram-positive bacteria and some gram-negative bacteria. Some “bad” bacteria, which were found in higher abundance in degenerated and herniated IVDs, like Pseudomonas veronii, Pseudomonas stutzeri, Streptococcus anginosus that can cause sub-clinical infection and inflammatory response inducing IVDs degeneration. Microbiome dysbiosis, an imbalance between “good” and “bad” microbiota can be influenced by various environmental factors, including diet, diseases, genetic makeup of the host and medical intervention. This disrupts the diversity of the gut microorganisms by regulating the numbers of bacterial communities [17, 18]. However, the mechanisms by which “good” bacteria protect the IVD tissues, and “bad” bacteria damage the IVD are still unclear and require further investigation.
This study also challenges the dogma associated with IVD sterility. The presence of 58 overlapping bacterial species between the intervertebral IVDs and gut, 29 between IVD and skin suggests that IVD microbiome may talk to the gut microbiome, the gut microbiome may infiltrate into the IVD environment and play a key role in the development of IDD. Findings from this study provided evidence for us to further dwell into the phenomenon of IVD dysbiosis which could be an important cause of inflammation and IVD degeneration. In this regard, P. acnes has been identified as the most frequently isolated anaerobic microbe and has a great association with the vertebral osteomyelitis in bone and joint infection diseases [19]. Moreover, studies in animal models and humans have shown that a persistent imbalance of microbial community could regulate the secretion of inflammatory cytokines TNF-α, IL-1β and PGE-2. Interestingly, these inflammatory molecules have been reported to correlate with IDD. Taken together, all these strongly indicate that there are a variety of environmental factors that could induce dysbiosis by modifying the gut immune system and enhancing inflammation. Moreover, these changes could have far-researching influences as seen in degenerative spinal diseases and bone dysfunctions.
Dysbiosis, IVD degeneration and low back pain
Alterations in microbiome composition and host responses to the microbiota causing pathological bone development and involution [20, 21], led to the concept of gut-bone marrow axis [22, 23] and gut-bone axis [21]. Following the report by Rajasekaran et al., a similar concept of gut-disc axis has emerged that may play a key role in IDD and low back pain [16]. We explain below the cross-talks between gut microbiome and three different tissue types, and how they might be implicated in low back pain.
Gut-bone marrow axis
Gut dysbiosis has been found to enhance barrier permeability and increase intestinal inflammation in animal models of arthritis, diabetes and obesity [24,25,26]. Changes of gut microbiota induce the low-grade inflammation through enhancing the resolution of some specific inflammatory factors like short-chain fatty acids (SCFAs), which are generated by fermentation of complex carbohydrates, or promoting the leakage of bacterial products such as LPS (lipopolysaccharide)[27]. This low-grade inflammation can regulate the gut epithelial barrier by changing the expression of myeloid differentiation primary response gene 88 (MyD88) [28]. Besides, gut bacteria can also influence peripheral immune cells, bone marrow haematopoietic stem and progenitor cells (BM HSPCs). This is clearly demonstrated by the fact that germ-free mice have reduced numbers of myeloid progenitors in BM [29]. Moreover, researchers found the rag1-deficient mice have lower numbers of HSPCs and this effect could be reversed through faecal transplantation from wild-type mice [30]. Gut microbiome has shown the capability of changing HSPC differentiation through destroying BM function in obesity [31]. Furthermore, human spine has 26 vertebral bodies, and each consists of a typical bone marrow niche. They are another rich source of immune cells and gut dysbiosis can cause aberrant immune cell formation from the spinal bone marrows, just like the bone marrows in the hindlimbs. This is another root of systemic inflammation and may indirectly lead to back pain [32]. Therefore, the existence of gut-bone marrow axis can help us to understand the function of microbiome on distant organs.
Gut-bone axis
Gut hormones secreted from enteroendocrine cells following nutrient ingestion via food intake modulate metabolic processes including glucose homeostasis and food intake, and several of these gut hormones are involved in the regulation of the energy-demanding processes of bone remodelling [33]. Food is a source of nutrients which could be taken up by gut enteroendocrine cell to secrete gut hormones which are involved in the regulation of the energy-demanding process of bone remodelling. Biomarkers of bone turnover reduce slowly after diet consumption with substantially larger influences observed on bone resorption, compared with bone formation [34]. After diet intake, enteroendocrine cells in the gut secrete cytokines and hormones that regulating different metabolic processes [35]. For example, SCFAs can modify the musculoskeletal system by regulating the bone cells [21]. Gastric inhibitory polypeptide (GIP) and glucagon-like peptide 1 (GLP-1) are gut secreted hormones responsible for the amplification of insulin secretion, which In turn can stimulate insulin-mediated increases in bone formation [36]. Moreover, knockout of the GIP receptor in mice resulted in decreased bone mineral density (BMD) (3.7% decrease in total BMD at five months of age compared to wild type) bone strength, and a decrease in markers of bone formation, alkaline phosphatase and osteocalcin [37]. Much research has proved that the changeable level of BMD has a positive relationship with low back pain. [38, 39] Understanding the pathophysiology and physiology of the gut-bone axis may identify new therapeutic candidates for bone disorders and back pain.
Gut-disc axis
IVD is complex fibrocartilaginous joints that are generally referred to as the largest avascular structures in the human body. The blood vessels in the IVD exist only in the longitudinal ligaments and the outer layers of the annulus fibrosis. However, there is neovascularization when disc herniates into the epidural space, physical damage and fracture or a local inflammation on the disc and endplates. Thus, the infiltration of blood vessels into the degenerating disc will also be detected [37]. Gut microbes alter the intestinal microbial environment, fostering the production of various signalling molecules, immune cells and metabolites that may benefit bone. We hypothesize the microbial effects on the IVD mainly through the bloodstream. These metabolites and biological molecules produced by the gut microbiome reach distance tissue and cause a local pathological change.
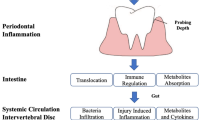
According to the existing evidence, we believe that microbiome could be an important factor for inducing and aggravating IDD through changing or regulating the outside and inside microenvironment of the IVD. Figure 1 illustrates the three potential mechanisms by which the gut microbiota can induce IVD and cause LBP: (1) translocation of the bacteria across the gut epithelial barrier and into the IVD, (2) regulation of the mucosal and systemic immune system and (3) regulation of nutrient absorption and metabolites formation at the gut epithelium and its diffusion into the IVD.
Gut-disc axis. Model of intestinal epithelial disruption signals that can regulate intervertebral disc degeneration. Gut microbes alter the intestinal microbial environment by modulating gut microbiota translocation and composition, increasing intestinal permeability and inflammation and modifying metabolites absorption. These changes can result in local and systemic responses that can cause IDD. A Pathogen displacement, B immune system modifications and C microbial metabolites production and cytokines
Translocation of the bacteria across the gut epithelial barrier and into IVDs
Growing evidence support that low-grade inflammation associated with microbiome dysbiosis is an essential factor for the onset of musculoskeletal diseases, like rheumatoid arthritis [38, 39]. Taking cues from this, it’s likely that the microbiome dysbiosis in the gut and IVD could be related to the IDD and herniation, and possibly LBP.
A healthy IVD microenvironment is very similar to the central nervous system (CNS). CNS is regarded as an immune-privileged, where blood–brain barrier (BBB) and blood–spinal cord barrier separate the CNS from any systemic inflammatory states to maintain homeostasis within this specialized, vulnerable organ [40]. The blood–disc barrier keeps the IVD immune-privileged and protects it from systemic infection. The blood–disc barrier keeps the IVD separated from other potential inflammation, but it also blocks the immune surveillance (playing the role of sentry) started by our body's immune system on the inner of disc. Immune cell cannot get timely feedback from the inner microenvironment of disc, which causes the worse inflammatory reaction. So it is the lack of IVD immune surveillance and hypoxic conditions that foster ideal conditions for the reproduction of anaerobic bacteria within the IVD upon invasion [41]. Furthermore, intestinal inflammation can promote intestinal permeability allowing more bacteria to cross the epithelial barrier [42]. Though most of the translocated bacteria are killed quickly by the immune system, a few bacteria can enter and survive while evading the immune response [43]. These bacteria arrive near the IVD and recruit more inflammatory cells (e.g. T cells, B cells, dendritic cells and macrophages) via release of pro-inflammatory factors (e.g. IL-6 and TNFα) [44]. Inflammatory cellular infiltration causes blood vessels to grow into the IVDs, which destroys the IVD’s appropriate anaerobic environment. Additionally, abnormal mechanical loads and continuous microscopic injuries may also induce irreversible IVD damage and microfractures formation. A broken IVD provides an ideal place for the reproduction and growth of bacteria that evade humoral and cellular immunity, as well as diffusion of harmful microbiome metabolites. Microfractures also allow the immune cells to enter the IVD, which further aggravates IVD damage.
Bacterial invasion into the broken IVDs stimulates the IVD cells to secret inflammatory cytokines, e.g. IL-1α/β, IL-17, TNF-α and IL-6. At the same time, these cytokines destroy the IVD extracellular matrix (ECM) by promoting aggrecan and collagen degradation. A series of chain reactions started by these cytokines results in chemokine production, which further damages the ECM [45]. Such inflammation inside the IVD also induces an imbalance in anabolic and catabolic responses by IVD cells, resulting in IDD, herniation and discogenic back pain. Moreover, the release of inflammatory molecules secreted by damaged IVDs and activation of macrophages, T and B cells, mast cells and neutrophils further amplify the inflammatory cascade and thereby IDD. Bacterial translocation into the IVDs can also lead to activation of these immune cells by the release of lipopolysaccharide (LPS) and cause persistent pain. Such inflammation also results in increased innervation into degenerate IVDs that amplifies pain and transmit the pain signal to peripheral afferent nerve fibres located in dorsal root ganglia (DRG) and brain [46].
Regulation of the mucosal and systemic immune system
IDD has been proven to be correlated with abnormal production of pro-inflammatory factors by IVD cells (nucleus pulposus cells and annulus fibrosus cells) themselves, as well as immune cells, e.g. neutrophils, T cells and macrophages [45]. These molecules trigger a series of pathogenic and inflammatory reactions in IVD, which can activate senescence, apoptosis and autophagy and cause IDD [47, 48].
Gut microbiota communicates with the human immune system in a reciprocal manner [49, 50]. It has been reported that in a germ-free mouse model, the absence of gut microbiota produces an immature mucosal immune system and reduces proper immune signalling [51]. Immune system also regulates the localization and composition of microbiome [52]. Besides, communication with different niche-specific microbiomes is very important to develop and enhance the function of the immune system. When there is a gut dysbiosis, weakening of the epithelial barrier promotes increased permeability resulting in increased contact between the intestinal microbiota and the mucosal immune system [53, 54]. This excessive contact causes a massive influx of activated immune cells. These cells release vast quantities of pro-inflammatory cytokines (e.g. IL-6 and TNFα) into the bloodstream and regulate bone metabolism. These inflammatory cytokines and activated immune cells can migrate and gather near IVDs and induce IDD by modulating bone resorption and remodelling [45, 55].
Notably, the migration of immune cells into the IVD stimulates the IVD cells to generate neurogenic factors, e.g. brain-derive neurotrophic factor (BDNF) and nerve growth factor (NGF) [45, 56, 57]. This induces the appearance of nociceptive nerve fibres in the IVD as well as dorsal root ganglion (DRG). In addition, immune cells cause an elevated expression of pain-associated cation channels in the DRG due to the creation of an inflammatory milieu. In fact, DRG stimulation is used as a treatment strategy for chronic LBP [58].
Regulation of nutrient absorption and metabolites at the gut epithelium
A layer of mucus cover on the gastrointestinal epithelium forms a physical and firm barrier to prevent the invasion of pathogens in our gut [54]. Goblet cells produce mucins (MUC) continuously in normal physiological circumstances; however, gut toxins, inflammatory cytokines, microbiome and microbial metabolites modify this process positively or negatively [59]. Some inflammatory molecules like TNFα, IL-1β and IL-6 are the main regulators of mucin exocytosis and synthesis [60]. It has been known that IL-6 promotes the expression of the secreted gel-forming mucins (MUC5AC, MUC2, MUC6 and MUC5B) [61,62,63]. The prominent secreted gel-forming mucin in the small and large intestine is Mucin2 (MUC2). MUC2 is an important protective barrier against external pathogens, and it has diverse functions in intestinal homeostasis [64]. The mucus layer serves as an energy provider for gut microbiome. Moreover, it also serves as a matrix for commensal microbiome colonization and attachment, which prevents pathogenic bacteria from growing/binding within the mucus [65]. Currently, the capability of microbiota to degrade MUC2 has been thought of as a pathogenicity factor. Pathogens, including enterotoxigenic E. coli, could degrade MUC2 mucin through different mechanisms [66, 67]. Besides, acute intestinal infection induces rapid mucous secretion, which may aid in eliminating pathogens. Cross talk between the intestinal microbiota and mucus layer contributes to the regulated production of mucin by goblet cells [68]. Moreover, goblet cells are heavily influenced by interactions with the immune system. The epithelial barrier could be influenced or impaired by the damaged goblet cell, synthesis dysregulation and altered post-translational modifications when immune system changes in our gut [69]. The impaired epithelial barrier results in translocation of bacteria and their toxic metabolites. Cell wall components like endotoxin/LPS, microbiome metabolites, such as SCFAs/D-lactate and inflammatory factors produced by immune cells infiltrate into bloodstream and induce long-distance inflammation in the IVD [70]. For instance, SCFAs are fermented by the bacteria in the gut where they provide energy to epithelial cells and promote the activity of immune cells [71, 72]. SCFAs have emerged as key regulatory metabolites produced by the gut microbiota.
Although many studies have evaluated the effects of prebiotic consumption on bone resorption or osteoclast formation, our understanding of how gut microbes communicate with IVD remains ill-defined [73]. The degenerative disc is accompanied by vertebral bone remodelling, including bony endplate thickening and osteophyte formation [74]. Calcification of the IVD has been correlated with osteoblast formation [75]. Animal research showed reduced osteoclast numbers in C57BL/6 mice and osteoporotic mice following propionate and butyrate treatment. SCFAs promote the differentiation of naive CD4 + cells into Tregs resided preferentially on the endosteal surfaces of bone [76]; Tregs could promote osteoblast differentiation and suppress osteoclastogenesis [77, 78], moreover, it is also required for parathyroid hormone-stimulated (PTH-stimulated) bone formation [79]. Besides, SCFAs have direct effects on inhibition of bone resorption or osteoclast formation either via activation of G Protein-Coupled Receptors (GPCR) or through histone deacetylase (HDAC) inhibition. Moreover, the appearance of calcium deposits in the disc and expression of the extracellular calcium-sensing receptor (CaSR) are closely related to the GPCR in the degenerated discs [80], which means that the diffusion of gut-derived SCFAs into the IVDs can therefore lead to calcification and IDD. SCFAs can also induce pro-inflammatory phenotypes of immune cells in the IVD and lead to the pathogenesis of neuropathic pain.
Taxonomic characteristics of gut microbiota in patients with IVD
The toxaemic factor hypothesis has been proposed for the relationship between microbiome dysbiosis and degenerative disease. This hypothesis assumed that the overgrowth of gram-negative bacteria in the intestines leads to an increase in toxic metabolites that enter the circulation and ultimately promote inflammation [81]. To investigate whether IVD is initiated by pathogenic bacteria and establish the correlation between the presence of certain microbial groups with the disease in question [75], taxonomic characteristics of stood/blood microbiota from IVD patients can be performed using microbiome diversity analysis based on16S rRNA data. 16S rRNA gene sequencing is the most commonly used method of microbiome diversity analysis [74] that aims to identify the composition of the microbial (bacterial) group in the biological samples. Microbiome diversity analysis includes alpha diversity and beta diversity analysis. Alpha diversity is used to measure how many microbial species exist in a single sample and the proportion of each species. Beta diversity aims to measure the similarity of the composition of the microbiome across different samples [76]. LEfSe (linear discriminant analysis effect size) analysis is frequently used to analyse differences in the microbiome and find the biomarkers [77]. PICRUSt (Phylogenetic Investigation of Communities by Reconstruction of Unobserved States) [78] helps us to predict the functional composition of a metagenome using marker gene data and a database of reference genomes.
Future directions
Findings by Rajasekaran et al. will lead to a paradigm shift and help us gain greater insight into the role of human microbiome in the pathophysiological changes of IDD. But it still has limitations with respect to sample size and challenges to avoid the cross-contamination of air pollutants and blood-borne bacteria while collecting IVD tissues. Moreover, this study only detected the existence of bacteria in IVDs. It has been shown that there is a large diversity of both eukaryotic viruses and prokaryotic phages in healthy humans [82]. Besides, human blood, which like IVD tissues was thought to be sterile in healthy conditions, has been detected to contain fungi and viruses [83]. We strongly believe that recognition of the effects of microbiome on IDD will broaden our horizons about microbial ecology in the gut and IVD in LBP patients. Future studies on fungi and viruses in IDD are necessary to revolutionize our thinking about their possible role in the development of IVD diseases. Furthermore, we believe that inflammation inhibition and interruption of amplification of cascade reaction in IVD by targeting gut and IVD microbiome are worthwhile for the treatment of IDD and LBP.
References
Hoy D, March L, Brooks P, Blyth F, Woolf A, Bain C, Williams G, Smith E, Vos T, Barendregt J, Murray C, Burstein R, Buchbinder R (2014) The global burden of low back pain: estimates from the Global Burden of Disease 2010 study. Ann Rheum Dis 73:968–974. https://doi.org/10.1136/annrheumdis-2013-204428
Khan AN, Jacobsen HE, Khan J, Filippi CG, Levine M, Lehman RA Jr, Riew KD, Lenke LG, Chahine NO (2017) Inflammatory biomarkers of low back pain and disc degeneration: a review. Ann N Y Acad Sci 1410:68–84. https://doi.org/10.1111/nyas.13551
Weisman MH (2012) Inflammatory back pain: the United States perspective. Rheum Dis Clin North Am 38:501–512. https://doi.org/10.1016/j.rdc.2012.09.002
Coscia MF, Denys GA, Wack MF (2016) Propionibacterium acnes, coagulase-negative staphylococcus, and the “Biofilm-like” intervertebral disc. Spine (Phila Pa 1976) 41:1860–1865. https://doi.org/10.1097/brs.0000000000001909
Capoor MN, Birkenmaier C, Wang JC, McDowell A, Ahmed FS, Brüggemann H, Coscia E, Davies DG, Ohrt-Nissen S, Raz A, Ruzicka F, Schmitz JE, Fischetti VA, Slaby O (2019) A review of microscopy-based evidence for the association of Propionibacterium acnes biofilms in degenerative disc disease and other diseased human tissue. Eur Spine J 28:2951–2971. https://doi.org/10.1007/s00586-019-06086-y
Boer CG, Radjabzadeh D, Medina-Gomez C, Garmaeva S, Schiphof D, Arp P, Koet T, Kurilshikov A, Fu J, Ikram MA, Bierma-Zeinstra S, Uitterlinden AG, Kraaij R, Zhernakova A, van Meurs JBJ (2019) Intestinal microbiome composition and its relation to joint pain and inflammation. Nat Commun 10:4881. https://doi.org/10.1038/s41467-019-12873-4
Biver E, Berenbaum F, Valdes AM, Araujo de Carvalho I, Bindels LB, Brandi ML, Calder PC, Castronovo V, Cavalier E, Cherubini A, Cooper C, Dennison E, Franceschi C, Fuggle N, Laslop A, Miossec P, Thomas T, Tuzun S, Veronese N, Vlaskovska M, Reginster JY, Rizzoli R (2019) Gut microbiota and osteoarthritis management: An expert consensus of the European society for clinical and economic aspects of osteoporosis, osteoarthritis and musculoskeletal diseases (ESCEO). Ageing Res Rev 55:100946. https://doi.org/10.1016/j.arr.2019.100946
De Luca F, Shoenfeld Y (2019) The microbiome in autoimmune diseases. Clin Exp Immunol 195:74–85. https://doi.org/10.1111/cei.13158
Zhang X, Zhang D, Jia H, Feng Q, Wang D, Liang D, Wu X, Li J, Tang L, Li Y, Lan Z, Chen B, Li Y, Zhong H, Xie H, Jie Z, Chen W, Tang S, Xu X, Wang X, Cai X, Liu S, Xia Y, Li J, Qiao X, Al-Aama JY, Chen H, Wang L, Wu QJ, Zhang F, Zheng W, Li Y, Zhang M, Luo G, Xue W, Xiao L, Li J, Chen W, Xu X, Yin Y, Yang H, Wang J, Kristiansen K, Liu L, Li T, Huang Q, Li Y, Wang J (2015) The oral and gut microbiomes are perturbed in rheumatoid arthritis and partly normalized after treatment. Nat Med 21:895–905. https://doi.org/10.1038/nm.3914
Asquith M, Sternes PR, Costello ME, Karstens L, Diamond S, Martin TM, Li Z, Marshall MS, Spector TD, le Cao KA, Rosenbaum JT, Brown MA (2019) HLA alleles associated with risk of ankylosing spondylitis and rheumatoid arthritis influence the gut microbiome. Arthritis Rheumatol 71:1642–1650. https://doi.org/10.1002/art.40917
Tajik N, Frech M, Schulz O, Schälter F, Lucas S, Azizov V, Dürholz K, Steffen F, Omata Y, Rings A, Bertog M, Rizzo A, Iljazovic A, Basic M, Kleyer A, Culemann S, Krönke G, Luo Y, Überla K, Gaipl US, Frey B, Strowig T, Sarter K, Bischoff SC, Wirtz S, Cañete JD, Ciccia F, Schett G, Zaiss MM (2020) Targeting zonulin and intestinal epithelial barrier function to prevent onset of arthritis. Nat Commun 11:1995. https://doi.org/10.1038/s41467-020-15831-7
Sokol H, Seksik P, Furet JP, Firmesse O, Nion-Larmurier I, Beaugerie L, Cosnes J, Corthier G, Marteau P, Doré J (2009) Low counts of Faecalibacterium prausnitzii in colitis microbiota. Inflamm Bowel Dis 15:1183–1189. https://doi.org/10.1002/ibd.20903
Durazzi F, Sala C, Castellani G, Manfreda G, Remondini D, De Cesare A (2021) Comparison between 16S rRNA and shotgun sequencing data for the taxonomic characterization of the gut microbiota. Sci Rep 11:3030. https://doi.org/10.1038/s41598-021-82726-y
Xiao L, Feng Q, Liang S, Sonne SB, Xia Z, Qiu X, Li X, Long H, Zhang J, Zhang D, Liu C, Fang Z, Chou J, Glanville J, Hao Q, Kotowska D, Colding C, Licht TR, Wu D, Yu J, Sung JJ, Liang Q, Li J, Jia H, Lan Z, Tremaroli V, Dworzynski P, Nielsen HB, Bäckhed F, Doré J, Le Chatelier E, Ehrlich SD, Lin JC, Arumugam M, Wang J, Madsen L, Kristiansen K (2015) A catalog of the mouse gut metagenome. Nat Biotechnol 33:1103–1108. https://doi.org/10.1038/nbt.3353
Daliri EB, Wei S, Oh DH, Lee BH (2017) The human microbiome and metabolomics: Current concepts and applications. Crit Rev Food Sci Nutr 57:3565–3576. https://doi.org/10.1080/10408398.2016.1220913
Rajasekaran S, Soundararajan DCR, Tangavel C, Muthurajan R, Sri Vijay Anand KS, Matchado MS, Nayagam SM, Shetty AP, Kanna RM, Dharmalingam K (2020) Human intervertebral discs harbour a unique microbiome and dysbiosis determines health and disease. Eur Spine J 29:1621–1640. https://doi.org/10.1007/s00586-020-06446-z
Munawar N, Ahsan K, Muhammad K, Ahmad A, Anwar MA, Shah I, Al Ameri AK, Al Mughairbi F (2021) Hidden role of gut microbiome dysbiosis in schizophrenia: Antipsychotics or psychobiotics as therapeutics? Int J Mol Sci. https://doi.org/10.3390/ijms22147671
Zaiss MM, Joyce Wu HJ, Mauro D, Schett G, Ciccia F (2021) The gut-joint axis in rheumatoid arthritis. Nat Rev Rheumatol 17:224–237. https://doi.org/10.1038/s41584-021-00585-3
Walter G, Vernier M, Pinelli PO, Million M, Coulange M, Seng P, Stein A (2014) Bone and joint infections due to anaerobic bacteria: an analysis of 61 cases and review of the literature. Eur J Clin Microbiol Infect Dis Off Publ Eur Soc Clin Microbiol 33:1355–1364. https://doi.org/10.1007/s10096-014-2073-3
Fernández-Murga ML, Olivares M, Sanz Y (2020) Bifidobacterium pseudocatenulatum CECT 7765 reverses the adverse effects of diet-induced obesity through the gut-bone axis. Bone 141:115580. https://doi.org/10.1016/j.bone.2020.115580
Zaiss MM, Jones RM, Schett G, Pacifici R (2019) The gut-bone axis: how bacterial metabolites bridge the distance. J Clin Invest 129:3018–3028. https://doi.org/10.1172/jci128521
Vallianou NG, Geladari E, Kounatidis D (2020) Microbiome and hypertension: Where are we now? J Cardiovasc Med (Hagerstown) 21:83–88. https://doi.org/10.2459/jcm.0000000000000900
Santisteban MM, Kim S, Pepine CJ, Raizada MK (2016) Brain-gut-bone marrow axis: implications for hypertension and related therapeutics. Circ Res 118:1327–1336. https://doi.org/10.1161/circresaha.116.307709
Matei DE, Menon M, Alber DG, Smith AM, Nedjat-Shokouhi B, Fasano A, Magill L, Duhlin A, Bitoun S, Gleizes A, Hacein-Bey-Abina S, Manson JJ, Rosser EC, Klein N, Blair PA, Mauri C (2021) Intestinal barrier dysfunction plays an integral role in arthritis pathology and can be targeted to ameliorate disease. Med (N Y) 2:864-883.e869. https://doi.org/10.1016/j.medj.2021.04.013
Luck H, Tsai S, Chung J, Clemente-Casares X, Ghazarian M, Revelo XS, Lei H, Luk CT, Shi SY, Surendra A, Copeland JK, Ahn J, Prescott D, Rasmussen BA, Chng MH, Engleman EG, Girardin SE, Lam TK, Croitoru K, Dunn S, Philpott DJ, Guttman DS, Woo M, Winer S, Winer DA (2015) Regulation of obesity-related insulin resistance with gut anti-inflammatory agents. Cell Metab 21:527–542. https://doi.org/10.1016/j.cmet.2015.03.001
Cani PD, Bibiloni R, Knauf C, Waget A, Neyrinck AM, Delzenne NM, Burcelin R (2008) Changes in gut microbiota control metabolic endotoxemia-induced inflammation in high-fat diet-induced obesity and diabetes in mice. Diabetes 57:1470–1481. https://doi.org/10.2337/db07-1403
Cani PD, Amar J, Iglesias MA, Poggi M, Knauf C, Bastelica D, Neyrinck AM, Fava F, Tuohy KM, Chabo C, Waget A, Delmée E, Cousin B, Sulpice T, Chamontin B, Ferrières J, Tanti JF, Gibson GR, Casteilla L, Delzenne NM, Alessi MC, Burcelin R (2007) Metabolic endotoxemia initiates obesity and insulin resistance. Diabetes 56:1761–1772. https://doi.org/10.2337/db06-1491
Everard A, Geurts L, Caesar R, Van Hul M, Matamoros S, Duparc T, Denis RG, Cochez P, Pierard F, Castel J, Bindels LB, Plovier H, Robine S, Muccioli GG, Renauld JC, Dumoutier L, Delzenne NM, Luquet S, Bäckhed F, Cani PD (2014) Intestinal epithelial MyD88 is a sensor switching host metabolism towards obesity according to nutritional status. Nat Commun 5:5648. https://doi.org/10.1038/ncomms6648
Khosravi A, Yáñez A, Price JG, Chow A, Merad M, Goodridge HS, Mazmanian SK (2014) Gut microbiota promote hematopoiesis to control bacterial infection. Cell Host Microbe 15:374–381. https://doi.org/10.1016/j.chom.2014.02.006
Kwon O, Lee S, Kim JH, Kim H, Lee SW (2015) Altered gut microbiota composition in Rag1-deficient mice contributes to modulating homeostasis of hematopoietic stem and progenitor cells. Immune Netw 15:252–259. https://doi.org/10.4110/in.2015.15.5.252
Luo Y, Chen GL, Hannemann N, Ipseiz N, Krönke G, Bäuerle T, Munos L, Wirtz S, Schett G, Bozec A (2015) Microbiota from obese mice regulate hematopoietic stem cell differentiation by altering the bone niche. Cell Metab 22:886–894. https://doi.org/10.1016/j.cmet.2015.08.020
Nouh MR, Eid AF (2015) Magnetic resonance imaging of the spinal marrow: Basic understanding of the normal marrow pattern and its variant. World J Radiol 7:448–458. https://doi.org/10.4329/wjr.v7.i12.448
Hansen MS, Frost M (2021) Alliances of the gut and bone axis. Semin Cell Dev Biol. https://doi.org/10.1016/j.semcdb.2021.06.024
Clowes JA, Hannon RA, Yap TS, Hoyle NR, Blumsohn A, Eastell R (2002) Effect of feeding on bone turnover markers and its impact on biological variability of measurements. Bone 30:886–890. https://doi.org/10.1016/s8756-3282(02)00728-7
Gribble FM, Reimann F (2017) Signalling in the gut endocrine axis. Physiol Behav 176:183–188. https://doi.org/10.1016/j.physbeh.2017.02.039
Nauck MA, Meier JJ (2016) The incretin effect in healthy individuals and those with type 2 diabetes: physiology, pathophysiology, and response to therapeutic interventions. Lancet Diabetes Endocrinol 4:525–536. https://doi.org/10.1016/s2213-8587(15)00482-9
Fournier DE, Kiser PK, Shoemaker JK, Battié MC, Séguin CA (2020) Vascularization of the human intervertebral disc: a scoping review. JOR Spine 3:e1123. https://doi.org/10.1002/jsp2.1123
Kishikawa T, Maeda Y, Nii T, Motooka D, Matsumoto Y, Matsushita M, Matsuoka H, Yoshimura M, Kawada S, Teshigawara S, Oguro E, Okita Y, Kawamoto K, Higa S, Hirano T, Narazaki M, Ogata A, Saeki Y, Nakamura S, Inohara H, Kumanogoh A, Takeda K, Okada Y (2020) Metagenome-wide association study of gut microbiome revealed novel aetiology of rheumatoid arthritis in the Japanese population. Ann Rheum Dis 79:103–111. https://doi.org/10.1136/annrheumdis-2019-215743
Scher JU, Abramson SB (2011) The microbiome and rheumatoid arthritis. Nat Rev Rheumatol 7:569–578. https://doi.org/10.1038/nrrheum.2011.121
Ikegami A, Wake H (2021) Microglial regulation of blood brain barrier, the neuro-immunological interface. Brain Nerve 73:913–919. https://doi.org/10.11477/mf.1416201861
Wedderkopp N, Thomsen K, Manniche C, Kolmos HJ, Secher Jensen T, Leboeuf Yde C (2009) No evidence for presence of bacteria in modic type I changes. Acta Radiol 50:65–70. https://doi.org/10.1080/02841850802524485
Thevaranjan N, Puchta A, Schulz C, Naidoo A, Szamosi JC, Verschoor CP, Loukov D, Schenck LP, Jury J, Foley KP, Schertzer JD, Larché MJ, Davidson DJ, Verdú EF, Surette MG, Bowdish DME (2017) Age-associated microbial dysbiosis promotes intestinal permeability, systemic inflammation, and macrophage dysfunction. Cell Host Microbe 21:455-466.e454. https://doi.org/10.1016/j.chom.2017.03.002
Hernandez CJ, Guss JD, Luna M, Goldring SR (2016) Links between the microbiome and bone. J Bone Miner Res 31:1638–1646. https://doi.org/10.1002/jbmr.2887
Schirmer M, Smeekens SP, Vlamakis H, Jaeger M, Oosting M, Franzosa EA, Ter Horst R, Jansen T, Jacobs L, Bonder MJ, Kurilshikov A, Fu J, Joosten LAB, Zhernakova A, Huttenhower C, Wijmenga C, Netea MG, Xavier RJ (2016) Linking the human gut microbiome to inflammatory cytokine production capacity. Cell 167:1125-1136.e1128. https://doi.org/10.1016/j.cell.2016.10.020
Risbud MV, Shapiro IM (2014) Role of cytokines in intervertebral disc degeneration: pain and disc content. Nat Rev Rheumatol 10:44–56. https://doi.org/10.1038/nrrheum.2013.160
Lyu FJ, Cui H, Pan H, Mc Cheung K, Cao X, Iatridis JC, Zheng Z (2021) Painful intervertebral disc degeneration and inflammation: from laboratory evidence to clinical interventions. Bone Res 9:7. https://doi.org/10.1038/s41413-020-00125-x
Zhang C, Gullbrand SE, Schaer TP, Lau YK, Jiang Z, Dodge GR, Elliott DM, Mauck RL, Malhotra NR, Smith LJ (2020) Inflammatory cytokine and catabolic enzyme expression in a goat model of intervertebral disc degeneration. J Orthop Res 38:2521–2531. https://doi.org/10.1002/jor.24639
De Luca P, de Girolamo L, Kouroupis D, Castagnetta M, Perucca Orfei C, Coviello D, Coco S, Correa D, Brayda-Bruno M, Colombini A (2020) Intervertebral disc and endplate cells response to IL-1β inflammatory cell priming and identification of molecular targets of tissue degeneration. Eur Cell Mater 39:227–248. https://doi.org/10.22203/eCM.v039a15
Rooks MG, Garrett WS (2016) Gut microbiota, metabolites and host immunity. Nat Rev Immunol 16:341–352. https://doi.org/10.1038/nri.2016.42
Shi N, Li N, Duan X, Niu H (2017) Interaction between the gut microbiome and mucosal immune system. Mil Med Res 4:14. https://doi.org/10.1186/s40779-017-0122-9
Mazmanian SK, Liu CH, Tzianabos AO, Kasper DL (2005) An immunomodulatory molecule of symbiotic bacteria directs maturation of the host immune system. Cell 122:107–118. https://doi.org/10.1016/j.cell.2005.05.007
Zheng D, Liwinski T, Elinav E (2020) Interaction between microbiota and immunity in health and disease. Cell Res 30:492–506. https://doi.org/10.1038/s41422-020-0332-7
Weiss GA, Hennet T (2017) Mechanisms and consequences of intestinal dysbiosis. Cell Mol Life Sci 74:2959–2977. https://doi.org/10.1007/s00018-017-2509-x
Yoo JY, Groer M, Dutra SVO, Sarkar A, McSkimming DI (2020) Gut Microbiota and immune system interactions. Microorganisms. https://doi.org/10.3390/microorganisms8101587
Hampton HR, Chtanova T (2019) Lymphatic migration of immune cells. Front Immunol 10:1168. https://doi.org/10.3389/fimmu.2019.01168
Gruber HE, Ingram JA, Hoelscher G, Zinchenko N, Norton HJ, Hanley EN Jr (2008) Brain-derived neurotrophic factor and its receptor in the human and the sand rat intervertebral disc. Arthritis Res Ther 10:R82. https://doi.org/10.1186/ar2456
Park EH, Moon SW, Suh HR, Hochman S, Lee MG, Kim YI, Jang IT, Han HC (2019) Disc degeneration induces a mechano-sensitization of disc afferent nerve fibers that associates with low back pain. Osteoarthr Cartil 27:1608–1617. https://doi.org/10.1016/j.joca.2019.07.010
Huygen F, Kallewaard JW, Nijhuis H, Liem L, Vesper J, Fahey ME, Blomme B, Morgalla MH, Deer TR, Capobianco RA (2020) Effectiveness and safety of dorsal root ganglion stimulation for the treatment of chronic pain: a pooled analysis. Neuromodulation 23:213–221. https://doi.org/10.1111/ner.13074
Ahmadi S, Razazan A, Nagpal R, Jain S, Wang B, Mishra SP, Wang S, Justice J, Ding J, McClain DA, Kritchevsky SB, Kitzman D, Yadav H (2020) Metformin reduces aging-related leaky gut and improves cognitive function by beneficially modulating gut microbiome/goblet cell/mucin axis. J Gerontol A Biol Sci Med Sci 75:e9–e21. https://doi.org/10.1093/gerona/glaa056
Enss ML, Cornberg M, Wagner S, Gebert A, Henrichs M, Eisenblätter R, Beil W, Kownatzki R, Hedrich HJ (2000) Proinflammatory cytokines trigger MUC gene expression and mucin release in the intestinal cancer cell line LS180. Inflamm Res 49:162–169. https://doi.org/10.1007/s000110050576
Oh S, Lillehoj HS, Lee Y, Bravo D, Lillehoj EP (2019) Dietary antibiotic growth promoters down-regulate intestinal inflammatory cytokine expression in chickens challenged With LPS or co-infected with eimeria maxima and clostridium perfringens. Front Vet Sci 6:420. https://doi.org/10.3389/fvets.2019.00420
Mejías-Luque R, Lindén SK, Garrido M, Tye H, Najdovska M, Jenkins BJ, Iglesias M, Ernst M, de Bolós C (2010) Inflammation modulates the expression of the intestinal mucins MUC2 and MUC4 in gastric tumors. Oncogene 29:1753–1762. https://doi.org/10.1038/onc.2009.467
Hansson GC (2020) Mucins and the microbiome. Annu Rev Biochem 89:769–793. https://doi.org/10.1146/annurev-biochem-011520-105053
Liu Y, Yu X, Zhao J, Zhang H, Zhai Q, Chen W (2020) The role of MUC2 mucin in intestinal homeostasis and the impact of dietary components on MUC2 expression. Int J Biol Macromol 164:884–891. https://doi.org/10.1016/j.ijbiomac.2020.07.191
Deplancke B, Gaskins HR (2001) Microbial modulation of innate defense: goblet cells and the intestinal mucus layer. Am J Clin Nutr 73:1131s–1141s. https://doi.org/10.1093/ajcn/73.6.1131S
Burger-van Paassen N, Vincent A, Puiman PJ, van der Sluis M, Bouma J, Boehm G, van Goudoever JB, van Seuningen I, Renes IB (2009) The regulation of intestinal mucin MUC2 expression by short-chain fatty acids: implications for epithelial protection. Biochem J 420:211–219. https://doi.org/10.1042/bj20082222
Mack DR, Michail S, Wei S, McDougall L, Hollingsworth MA (1999) Probiotics inhibit enteropathogenic E. coli adherence in vitro by inducing intestinal mucin gene expression. Am J Physiol 276:G941-950. https://doi.org/10.1152/ajpgi.1999.276.4.G941
Pelaseyed T, Bergström JH, Gustafsson JK, Ermund A, Birchenough GM, Schütte A, van der Post S, Svensson F, Rodríguez-Piñeiro AM, Nyström EE, Wising C, Johansson ME, Hansson GC (2014) The mucus and mucins of the goblet cells and enterocytes provide the first defense line of the gastrointestinal tract and interact with the immune system. Immunol Rev 260:8–20. https://doi.org/10.1111/imr.12182
Grondin JA, Kwon YH, Far PM, Haq S, Khan WI (2020) Mucins in intestinal mucosal defense and inflammation: learning from clinical and experimental studies. Front Immunol 11:2054. https://doi.org/10.3389/fimmu.2020.02054
Mortensen PB, Clausen MR (1996) Short-chain fatty acids in the human colon: relation to gastrointestinal health and disease. Scand J Gastroenterol Suppl 216:132–148. https://doi.org/10.3109/00365529609094568
Wenzel TJ, Gates EJ, Ranger AL, Klegeris A (2020) Short-chain fatty acids (SCFAs) alone or in combination regulate select immune functions of microglia-like cells. Mol Cell Neurosci 105:103493. https://doi.org/10.1016/j.mcn.2020.103493
Parada Venegas D, De la Fuente MK, Landskron G, González MJ, Quera R, Dijkstra G, Harmsen HJM, Faber KN, Hermoso MA (2019) Short chain fatty acids (SCFAs)-mediated gut epithelial and immune regulation and its relevance for inflammatory bowel diseases. Front Immunol 10:277. https://doi.org/10.3389/fimmu.2019.00277
Yan J, Takakura A, Zandi-Nejad K, Charles JF (2018) Mechanisms of gut microbiota-mediated bone remodeling. Gut Microbes 9:84–92. https://doi.org/10.1080/19490976.2017.1371893
Mackiewicz Z, Salo J, Konttinen YT, Kaigle Holm A, Indahl A, Pajarinen J, Holm S (2009) Receptor activator of nuclear factor kappa B ligand in an experimental intervertebral disc degeneration. Clin Exp Rheumatol 27:299–306
Rutges JP, Duit RA, Kummer JA, Oner FC, van Rijen MH, Verbout AJ, Castelein RM, Dhert WJ, Creemers LB (2010) Hypertrophic differentiation and calcification during intervertebral disc degeneration. Osteoarthr Cartil 18:1487–1495. https://doi.org/10.1016/j.joca.2010.08.006
Fujisaki J, Wu J, Carlson AL, Silberstein L, Putheti P, Larocca R, Gao W, Saito TI, Lo Celso C, Tsuyuzaki H, Sato T, Côté D, Sykes M, Strom TB, Scadden DT, Lin CP (2011) In vivo imaging of Treg cells providing immune privilege to the haematopoietic stem-cell niche. Nature 474:216–219. https://doi.org/10.1038/nature10160
Qiu X, Gui Y, Xu Y, Li D, Wang L (2015) DHEA promotes osteoblast differentiation by regulating the expression of osteoblast-related genes and Foxp3(+) regulatory T cells. Biosci Trends 9:307–314. https://doi.org/10.5582/bst.2015.01073
Zaiss MM, Axmann R, Zwerina J, Polzer K, Gückel E, Skapenko A, Schulze-Koops H, Horwood N, Cope A, Schett G (2007) Treg cells suppress osteoclast formation: a new link between the immune system and bone. Arthritis Rheum 56:4104–4112. https://doi.org/10.1002/art.23138
Yu M, D’Amelio P, Tyagi AM, Vaccaro C, Li JY, Hsu E, Buondonno I, Sassi F, Adams J, Weitzmann MN, DiPaolo R, Pacifici R (2018) Regulatory T cells are expanded by Teriparatide treatment in humans and mediate intermittent PTH-induced bone anabolism in mice. EMBO Rep 19:156–171. https://doi.org/10.15252/embr.201744421
Grant MP, Epure LM, Bokhari R, Roughley P, Antoniou J, Mwale F (2016) Human cartilaginous endplate degeneration is induced by calcium and the extracellular calcium-sensing receptor in the intervertebral disc. Eur Cell Mater 32:137–151. https://doi.org/10.22203/ecm.v032a09
Sun Y, Chen Q, Lin P, Xu R, He D, Ji W, Bian Y, Shen Y, Li Q, Liu C, Dong K, Tang YW, Pei Z, Yang L, Lu H, Guo X, Xiao L (2019) Characteristics of gut microbiota in patients with rheumatoid arthritis in Shanghai. China Front Cell Infect Microbiol 9:369. https://doi.org/10.3389/fcimb.2019.00369
Liang G, Bushman FD (2021) The human virome: assembly, composition and host interactions. Nat Rev Microbiol 19:514–527. https://doi.org/10.1038/s41579-021-00536-5
Baker JL, Bor B, Agnello M, Shi W, He X (2017) Ecology of the oral microbiome: beyond bacteria. Trends Microbiol 25:362–374. https://doi.org/10.1016/j.tim.2016.12.012
Funding
Open Access funding enabled and organized by CAUL and its Member Institutions.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, W., Lai, K., Chopra, N. et al. Gut-disc axis: A cause of intervertebral disc degeneration and low back pain?. Eur Spine J 31, 917–925 (2022). https://doi.org/10.1007/s00586-022-07152-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00586-022-07152-8