Abstract
Purpose
Inflammatory and oxidative stress upregulates matrix metalloproteinase (MMP) activity, leading to intervertebral disc degeneration (IDD). Gene therapy using human tissue inhibitor of metalloproteinase 1 (hTIMP1) has effectively treated IDD in animal models. However, persistent unregulated transgene expression may have negative side effects. We developed a recombinant adeno-associated viral (AAV) gene vector, AAV-NFκB-hTIMP1, that only expresses the hTIMP1 transgene under conditions of stress.
Methods
Rabbit disc cells were transfected or transduced with AAV-CMV-hTIMP1, which constitutively expresses hTIMP1, or AAV-NFκB-hTIMP1. Disc cells were selectively treated with IL-1β. NFκB activation was verified by nuclear translocation. hTIMP1 mRNA and protein expression were measured by RT-PCR and ELISA, respectively. MMP activity was measured by following cleavage of a fluorogenic substrate.
Results
IL-1β stimulation activated NFκB demonstrating that IL-1β was a surrogate for inflammatory stress. Stimulating AAV-NFκB-hTIMP1 cells with IL-1β increased hTIMP1 expression compared to unstimulated cells. AAV-CMV-hTIMP1 cells demonstrated high levels of hTIMP1 expression regardless of IL-1β stimulation. hTIMP1 expression was comparable between IL-1β stimulated AAV-NFκB-hTIMP1 cells and AAV-CMV-hTIMP1 cells. MMP activity was decreased in AAV-NFκB-hTIMP1 cells compared to baseline levels or cells exposed to IL-1β.
Conclusion
AAV-NFκB-hTIMP1 is a novel inducible transgene delivery system. NFκB regulatory elements ensure that hTIMP1 expression occurs only with inflammation, which is central to IDD development. Unlike previous inducible systems, the AAV-NFκB-hTIMP1 construct is dependent on endogenous factors, which minimizes potential side effects caused by constitutive transgene overexpression. It also prevents the unnecessary production of transgene products in cells that do not require therapy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
An estimated 84% of adults experience low back pain during their lifetime [1]. Of the many causes for low back pain, intervertebral disc degeneration (IDD) is a common contributor [2]. IDD results from an imbalance between catabolism and anabolism of the disc extracellular matrix (ECM) [3]. Loss of the ECM proteoglycan leads to dehydration and fibrosis of the nucleus pulposus (NP), which commonly results in formation of fissures in the annulus fibrosis (AF) [4]. These disc fissures are not only responsible for the abnormal biomechanical changes seen in IDD, but are also associated with neoinnervation, which has been implicated in pain [5].
The pathways that regulate intervertebral disc matrix homeostasis provide potential targets for gene therapy. For example, Le Maitre et al. [6] delivered an interleukin-1 (IL-1) receptor antagonist through a gene therapy approach and noted inhibition of matrix degradation. Recently, our lab introduced human tissue inhibitor of metalloproteinase 1 (hTIMP1) and bone morphogenetic protein (BMP) into the NP of rabbits and demonstrated delayed disc degeneration in a rabbit annulotomy model of IDD [7]. hTIMP1 has demonstrated promise as a therapeutic factor to ameliorate IDD. hTIMP1 inhibits the activities of most of the known matrix metalloproteinases (MMPs), the catabolic enzymes involved in degrading the ECM and also facilitates collagen and aggrecan expression in the disc [8,9,10,11].
Despite these encouraging results, the treatment of nonlethal diseases, such as IDD, with gene therapy presents unique safety concerns. Specifically, previous gene therapy studies have used cytomegalovirus (CMV) promoters to drive transgene expression, resulting in constitutively elevated protein expression that may have negative metabolic consequences. For example, Wallach et al. [12] noted severe neurological changes and significant histological changes in rabbits after intradural injection of gene constructs that resulted in elevated levels of TGF-β1. To avoid such unfavourable sequelae, controlled inducible transgene expression systems have been developed. We previously demonstrated tetracycline-inducible and RheoSwitch-inducible expression systems that successfully modulated transgene expression in NP cells [13, 14]. Unfortunately, these systems would be difficult to use clinically since gene expression requires an exogenous activator ligand.
Another concern with gene therapy is that viral vectors, specifically adenovirus (AV) vectors, may trigger severe immune responses that can have devastating consequences, as demonstrated by the death of a patient treated for an enzymatic deficiency with an AV vector [15]. Additionally, the immune response degrades the adenoviral episome, which leads to a discontinuation of therapeutic protein production [16, 17]. To mitigate the immune response, adeno-associated virus (AAV) vectors have become more popular. AAV vectors differ from AV vectors in that they require a helper virus to be expressed, which limits expression of AAV gene products and the resulting cellular mediated immune response [18].
The aim of this study was to establish a gene expression system that produces hTIMP1 protein only under conditions of stress. We achieved this using a novel transgene regulation system, AAV-NFκB-hTIMP1. The system was built on an AAV backbone with hTIMP1 expression regulated by a mini-promoter containing NFκB elements, which is turned on by inflammatory and oxidative stress and turned off upon diminishment of cellular stress [19].
Materials and methods
Overview of the study
Rabbit disc cells were transfected with AAV-CMV-hTIMP1 or AAV-NFκB-hTIMP1 plasmid DNA (pAAV-CMV-hTIMP1 or pAAV-NFκB-hTIMP1), or transduced with AAV-CMV-hTIMP1 or AAV-NFκB-hTIMP1 viruses (vAAV-CMV-hTIMP1 or vAAV-NFκB-hTIMP1). Disc cells were treated with IL-1β to induce inflammatory stress to activate the NFκB signaling pathway. Reverse transcription polymerase chain reaction (RT-PCR) and enzyme-linked immunosorbent assay (ELISA) were used to determine the levels of expression of hTIMP1 messenger RNA (mRNA) and protein, respectively. To test the temporal expression of hTIMP1 in response to stress, pAAV-NFκB-hTIMP1 transfected cells or vAAV-NFκB-hTIMP1 transduced cells were either transiently stimulated with IL-1β for 30 min—after which IL-1β was removed—or continuously stimulated with IL-1β for 96 h. hTIMP1 expression was measured in culture media and cell lysate at 0, 0.5, 12, 24, 48, 72, and 96 h after IL-1β treatment. The study design is summarized below (Fig. 1).
a Identification of an inducible transgene expression system in response to IL-1β. Rabbit disc cells were transfected or transduced with AAV-CMV-hTIMP1 or AAV-NFκB-hTIMP1. Cells were selectively exposed to IL-1β. RT-PCR and ELISA measured mRNA and protein expression 24 h after IL-1β exposure. b Demonstration of the time course of the hTIMP1 transgene expression. Rabbit disc cells were transfected or transduced with AAV-NFκB-hTIMP1. Cells were selectively exposed to IL-1β for either 30 min or continuously. RT-PCR and ELISA were used to measure mRNA and protein expression at different time points after stimulation
Construction of plasmids and production of viruses
The full length of functional hTIMP1 gene (0.62 kb) was derived from our previously published construct, AAV-CMV-hTIMP1 [7]. hTIMP1 was cloned into a regulated AAV vector driven by the NFκB promoter, AAV-NFκB-hTIMP1. Both AAV-NFκB-hTIMP1 and AAV-CMV-hTIMP1 gene constructs were utilized, with the latter serving as a control for constitutive hTIMP1 expression (Fig. 2). Plasmids were purified through CsCl2 gradient ultracentrifugation according to previously published protocols [20].
Recombinant adeno-associated viral (AAV) vectors. Schematic illustration of two AAV vectors containing hTIMP1 gene (624 bp). Vectors were controlled by either the human CMV immediate-early promoter/enhancer (PCMV) or the human NFκB regulatory elements (PNFκB) and then, followed by the simian virus 40 polyadenylation signal (PSV40) in the self-complementary AAV vector. ∆ITR, truncated inverted terminal repeat; LTR, long terminal repeat
Recombinant AAV viral vector stocks were produced according to the three-plasmid co-transfection method as described by Xiao et al. [21]. The viral stocks were obtained by CsCl2 gradient centrifugation. The titers of viral preparations used in the study were between 1 × 1012 and 1013 particles/mL.
Disc cell culture
AF and NP cells were isolated from New Zealand white rabbits immediately after sacrifice and cultured in F-12, 10% fetal bovine serum, 1% penicillin–streptomycin at 37 °C, and 5% CO2 until 90% confluence, as previously described [22]. Passage 1 cells were seeded on 12-well plates and expanded in monolayer for three days in normoxic conditions for subsequent transfection and transduction as described below.
Transfection of pAAV-NFκB-hTIMP1 and pAAV-CMV-hTIMP1
pAAV-NFκB-hTIMP1 and pAAV-CMV-hTIMP1 were transfected into rabbit AF (rAF) and rabbit NP (rNP) cells using Xfect Transfection Reagent (Clontech Laboratories, Inc., Palo Alto, CA) according to the manufacturer’s instructions. Briefly, rAF and rNP cells were seeded into 12-well plates at 25 k cells/well with a complete cell culture medium and cultured to 50% confluence. Xfect Polymer (0.75 μl/well) was mixed with pAAV-CMV-hTIMP1 or pAAV-NFκB-hTIMP1 (2.5 μg/well) and incubated at room temperature for 10 min to allow nanoparticle complexes to form. Transfection complexes were added to cells and incubated in a 37 °C/CO2 incubator.
Transduction of vAAV-NFκB-hTIMP1 and vAAV-CMV-hTIMP1
rAF and rNP cells were seeded into 12-well plates at 25 k cells/well with a complete cell culture medium and cultured to 50% confluence. 10–15 µl virus stock of vAAV-CMV-hTIMP1 or vAAV-NFκB-hTIMP1 was added to each well. Plates were placed in a 37 °C/CO2 incubator.
NFκB activation
Nuclear translocation of NFκB was detected using a fluorescent microscope at time points 0, 0.5, 12, 24, 48, and 72 h after IL-1β treatment. NFκB was visualized by immunofluorescent labelling using an anti-p65 antibody (Santa Cruz Biotechnology, Santa Cruz, CA). Nuclear translocation of NFκB was quantified using a customized MATLAB (The MathWorks, Inc, Natick, MA) program to analyze the images. The experiment was conducted in triplicate.
Inducible transgene expression
To assess whether transgene expression was inducible by IL-1β, cells were divided into six groups: cells alone without IL-1β, cells alone with IL-1β, pAAV-CMV-hTIMP1 without IL-1β, pAAV-CMV-hTIMP1 with IL-1β, pAAV-NFκB-hTIMP1 without IL-1β, and pAAV-NFκB-hTIMP1 with IL-1β. If indicated, 10 ng/mL of IL-1β was added at 24 h post-transfection. Supernatants and cells were collected for RT-PCR, ELISA, and MMP activity assay after treatment with IL-1β for 24 h. This experiment was repeated for cells that were transduced with vAAV-CMV-hTIMP1 or vAAV-NFκB-hTIMP1.
Time course of transgene expression
To assess the time course of the hTIMP1 transgene expression, cells transfected with pAAV-NFκB-hTIMP1 were divided into three groups: cells not treated with IL-1β, cells treated with IL-1β transiently for 30 min, and cells treated continuously with IL-1β for 96 h. If indicated, 10 ng/mL of IL-1β was added at 24 h post-transfection. Supernatant and cells were collected for RT-PCR and ELISA at 0, 0.5, 12, 24, 48, 72, and 96 h after IL-1β treatment. This experiment was repeated for cells that were transduced with vAAV-NFκB-hTIMP1.
hTIMP1 mRNA expression
For mRNA isolation, cells in PBS were detached from the plate by mechanical disruption. RLT lysis buffer containing 1% β-mercaptoethanol was added to the cells. The resultant solution was passed through a QIAshredder (Qiagen, Valencia, CA), and mRNA was isolated using an RNA extraction kit (Qiagen, Valencia, CA) with a DNase I step to remove genomic material. hTIMP1 mRNA expression was measured by RT-PCR using primer pair 5′-TGGCTTCTGGCATCCTGTTGTTG-3′ and 5′-CGCTGGTATAAGGTGGTCTGGTTG-3′. Relative changes in mRNA were quantified using the ΔΔCt method using glyceraldehyde 3-phosphate dehydrogenase (GAPDH) to normalize and compare between groups [23].
hTIMP1 protein expression
hTIMP1 protein expression in cell culture supernatants were measured by hTIMP1 ELISA Duoset (R&D Systems, Minneapolis, MN) according to the manufacturer’s instructions. Briefly, 2 × 106 cells were seeded into 60 mm dishes. After all cells attached, culture medium was replaced with fresh medium for 24 h. Cell culture supernatants were harvested after centrifugation to remove cell debris and diluted to be within the range of the standard curve.
MMP enzymatic activity
MMP activity was assayed using 520 MMP Fret Substrate XI (AnaSpec, Fremont, CA). Cell-conditioned media (CM) samples were divided into two sets: 1 mM APMA was added to one set of APMA-activated CM samples and PBS was added to the second set of non-activated control CM samples. Immediately after adding APMA/PBS, 520 MMP Fret Substrate XI solution was applied to all samples and fluorescence (Ex/Em = 355/460 nm) was measured on a BioTek microplate reader every 5 min over 1 h. MMP activities were determined as the differences between sample fluorescence measurements and those of corresponding vehicle controls. A linear fit model was used to determine the rate of the cleavage reaction. Cleavage rates were compared both among samples and between APMA-activated and non-activated sample treatments.
Statistical analysis
Student’s t-test with significance set to p < 0.05 was used to identify differences in mRNA expression, protein expression, and MMP enzymatic activity between control cells, cells with the AAV-CMV-hTIMP1 plasmid, and cells with the AAV-NFκB-hTIMP1 plasmid. Student’s t-test with significance set to p < 0.05 was used to identify differences in mRNA expression and protein expression in cells transfected with pAAV-NFκB-hTIMP1 or cells transduced with vAAV-NFκB-hTIMP1 that were untreated, treated with IL-1β for 30 min, or treated with IL-1β continuously for 96 h.
Results
NFκB activation in response to inflammatory stimulus in disc cells
The transgene was designed to be regulated by the NFκB signalling pathway, which responds to inflammatory stress. To confirm that IL-1β was an appropriate surrogate for inflammatory stress, NP cells were exposed to IL-1β and nuclear translocation of NFκB from the cytoplasm was assessed. In IL-1β treated cells, NFκB was activated—indicated by increased NFκB nuclear translocation—30 min after IL-1β treatment (Fig. 3). NFκB activation remained, albeit at modestly decreased levels, from 12 to 72 h after IL-1β treatment, which is consistent with the dynamics of NFκB activation reported previously [24, 25]. These findings demonstrated that the NFκB signalling pathway can be robustly activated by the pro-inflammatory cytokine IL-1β in disc cells, which justifies the use of IL-1β to investigate the expression of the hTIMP1 transgene from AAV-NFκB-hTIMP1.
NFκB activation by IL-1β in NP cells. Immunofluorescence of p65, a subunit of NFκB, was measured at 0, 0.5, 12, 24, 48, and 72 h after cells were treated with 10 ng/mL of IL-1β. Cells exhibited complete translocation of p65 from the cytoplasm to the nuclei 30 min after IL-1β treatment. p65 nuclear localization gradually decreased at the later time points
Inflammatory stress upregulates hTIMP1 gene and protein expression in disc cells harbouring the AAV- NFκB-hTIMP1 DNA construct
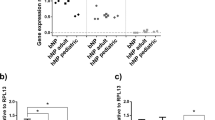
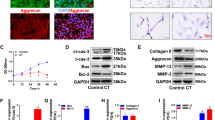
To evaluate regulated expression of hTIMP1 in our new transgene system, rAF and rNP cells were transfected with either pAAV-NFκB-hTIMP1 or pAAV-CMV-hTIMP1. pAAV-CMV-hTIMP1 served as a positive control for high production of hTIMP1 from the constitutively expressed CMV promoter. As expected, pAAV-CMV-hTIMP1 transfected disc cells highly expressed hTIMP1 mRNA and protein in an unregulated manner, regardless of the absence or presence of IL-1β (Fig. 4). hTIMP1 mRNA expression as measured by RT-PCR from pAAV-NFκB-hTIMP1 transfected disc cells was negligible in the absence of a stressful stimulus, i.e. IL-1β, in both rAF (Fig. 4a) and rNP cells (Fig. 4b). With IL-1β treatment, hTIMP1 mRNA expression increased five- to six-fold in pAAV-NFκB-hTIMP1 transfected disc cells to a level comparable to the constitutive expression seen in pAAV-CMV-hTIMP1 transfected disc cells. hTIMP1 protein levels as determined by ELISA mirrored those of hTIMP1 mRNA (Fig. 4c, d). Negative control cells, which were not transfected, did not express any hTIMP1 mRNA or protein. These results suggest that hTIMP1 expression from pAAV-NFκB-hTIMP1 is upregulated only under conditions of stress.
hTIMP1 mRNA and protein expression in disc cells transfected with plasmid vectors as measured by RT-PCR and ELISA. In cells transfected with NFκB promotor vectors, hTIMP1 expression was induced by IL-1β with little expression at baseline. In contrast, there was noted to be high hTIMP1 expression in cells with CMV promotor vectors regardless of stimulus. * = a value significantly different from that of other groups (p < 0.05)
Regulated expression of hTIMP1 in our new transgene system was also confirmed in rAF and rNP cells transduced with vAAV-NFκB-hTIMP1 or vAAV-CMV-hTIMP1. hTIMP1 gene and protein expressions in vAAV-NFκB-hTIMP1 transduced disc cells were similar to those observed in pAAV-NFκB-hTIMP1 transfected disc cells, i.e. negligible hTIMP1 expression at baseline and high expression of hTIMP1 with IL-1β stimulation. The negative controls had no expression of hTIMP1 and the positive controls under the constitutive promoter transduced with vAAV-CMV-hTIMP1 had high expression regardless of IL-1β stimulation (Fig. 5).
hTIMP1 mRNA and protein expression in disc cells transduced with viral vectors as measured by RT-PCR and ELISA. Cells transduced with viral vectors displayed similar pattern to cells transfected with plasmid vectors, confirming that expression was secondary to the NFκB-mediated promoter’s selective response to IL-1β. * = a value significantly different from that of other groups (p < 0.05)
Upregulated hTIMP1 gene expression from AAV-NFκB-hTIMP1 suppressed MMP enzymatic activity in disc cells
hTIMP1 has been used as a therapeutic to minimize pathologic proteolytic degradation of the ECM by MMPs. Hence, it is important to evaluate if hTIMP1 expression from the AAV-NFκB-hTIMP1 construct also regulates MMP activity in disc cells. Non-transfected disc cells exhibited a low level of baseline MMP enzymatic activity, which was upregulated three-fold in the presence of IL-1β. These results demonstrate that under condition of stress without the presence of MMP inhibitors such as hTIMP1, there is a significant increase in MMP activity in disc cells, as expected.
Upregulated hTIMP1 expression in the presence of IL-1β in pAAV-NFκB-hTIMP1 transfected disc cells dramatically blunted the observed increase in MMP enzymatic activity (Fig. 6). Similar suppression of MMP activity was seen in pAAV-CMV-hTIMP1 transfected disc cells in which hTIMP1 was constitutively expressed. Disc cells transfected with pAAV-NFκB-hTIMP1 in the absence of IL-1β also exhibited minimal MMP activity, confirming that the transfection itself did not result in increased MMP activity.
Downregulation of hTIMP1 expression following IL-1β withdrawal in disc cells harbouring the AAV-NFκB-hTIMP1 DNA construct
The AAV-NFκB-hTIMP1 construct was designed to express hTIMP1 transgene only under conditions of stress. hTIMP1 indeed was expressed in the presence of IL-1β stimulation and not in the absence of IL-1β (Figs. 4, 5). However, the question remained as to whether IL-1β-induced expression of hTIMP1 would diminish in AAV-NFκB-hTIMP1 containing disc cells once the stressful stimulus was removed. This idea was tested by exposing pAAV-NFκB-hTIMP1 transfected disc cells to IL-1β for 30 min, then removing IL-1β from the culture media and quantifying hTIMP1 expression at different time points following IL-1β withdrawal. As a positive control, AAV-NFκB-hTIMP1 transfected disc cells were treated with IL-1β continuously for 96 h. As a negative control, untreated AAV-NFκB-hTIMP1 transfected disc cells were included. rAF and rNP cells transfected with pAAV-NFκB-hTIMP1 and exposed to 30 min of IL-1β demonstrated peaked hTIMP1 mRNA and protein expression within 12–24 h compared to the negative control group. More importantly, the levels of hTIMP1 mRNA and protein progressively decreased in the transiently stimulated cells after 24 h and reached a minimum at 96 h after IL-1 withdrawal (Fig. 7).
In cells continuously stimulated with IL-1β for 96 h, hTIMP1 mRNA and protein expression were generally increased at all time points within the 24- to 96-h period when compared to the transiently stimulated group. The negative control group demonstrated consistently low expression of hTIMP1 in both AF and NP cells (Fig. 7).
It is possible that the reduced hTIMP1 expression after IL-1β withdrawal was not only due to decreased NFκB activation but also due to transient transfection of the plasmid, i.e. loss of plasmid from the cells over time. To address the possible loss of plasmids from cells due to transient transfection, we performed the same experiments except with transduction instead of transfection. Downregulation of hTIMP1 mRNA and protein expression following IL-1β withdrawal was confirmed in rAF and rNP cells transduced with vAAV-NFκB-hTIMP1 (Fig. 8). These results were almost identical results to those in cells transfected with pAAV-NFκB-hTIMP1, which implies that downregulation of hTIMP1 expression is due to withdrawal of inflammatory stress and not transient transfection of cells.
Discussion
Past IDD gene therapy studies used unregulated expression vectors with constitutively expressed transgenes. While these studies showed promise, there was concern regarding unregulated, constitutive expression of transgene proteins, particularly for growth factors where overexpression can cause undesirable side effects [12]. In addition, previous attempts to control transgene expression relied on an exogenously added bio-signal (e.g. tetracycline) to stimulate or inhibit expression. This study demonstrates the proof of concept of an intrinsic inducible transgene expression system to treat IDD by regulated production of hTIMP1 in disc cells, responsive to the internal environment in the disc, potentially improving the overall safety and effectiveness of intradiscal gene therapy. The NFκB promoter allows controlled hTIMP1 expression, i.e. hTIMP1 gene is turned on under conditions of stress and off in the absence of cellular stress, avoiding untoward side effects of constitutive expression and associated metabolic stress on disc cells. Compared to the traditional CMV-driven gene therapy strategy, this regulated system prevents bioenergetic overtaxing from chronic overexpression of the transgene and adverse effects of accumulated transgene protein products.
As demonstrated in Figs. 4 and 5, disc cells transfected or transduced with AAV-CMV-hTIMP1 with the constitutive CMV promoter resulted in unregulated constitutive expression of hTIMP1, while disc cells harbouring AAV-NFκB-hTIMP1 exhibited hTIMP1 expression only in the presence of IL-1β, demonstrating that the system is inducible. Furthermore, upregulation of transgene expression by IL-1β highlights the potential applicability of our regulated gene therapy strategy to treat inflammatory-related conditions, such as IDD, induced only under conditions of inflammation. Of note, previous measurements of IL-1β in degenerated human disc cells range from 34.9 to 135.5 pg/mL, which is well below the 10 ng/mL amount used in this experiment [24]. However, IL-1 levels as low as 100 pg/mL, similar to the levels seen in vivo in degenerated discs, are reported to activate inflammatory pathways, such as NFκB [25, 26]. Additionally, other pro-inflammatory cytokines that activate NFκB, such as TNF-α, are present in degenerated disc cells. Furthermore, NFκB activation is well documented in degenerative discs in vivo [27,28,29]. Thus, although confirmatory in vivo studies are needed, it is highly likely that expression of the hTIMP1 transgene in the AAV-NFκB-hTIMP1 vector system would be induced in degenerated discs in vivo.
Perhaps more importantly, our inducible system turns off when the source of stress is removed, providing an additional safety feature to prevent constitutive expression. After stimulation of disc cells by IL-1β both hTIMP1 mRNA expression and protein production prominently increase. In the transiently stimulated group, cells treated with IL-1β for only 30 min, hTIMP1 mRNA and protein levels started to decline 12 and 24 h after the first dose of IL-1β, respectively. In the continuously stimulated group, hTIMP1 levels were higher than that of the transiently stimulated group throughout 96 h. It should be mentioned that hTIMP1 expression did decrease in the continuously stimulated group. This decrease could be due to either degradation of the plasmid or the oscillatory nature of NFκB activation by IL-1β, whereby NFκB activates its own repressor resulting in activity that oscilates with time [30, 31]. While this characteristic may cause difficulty maintaining a constant amount of gene product, there is significantly more gene product being produced in those cells maintained in a pro-inflammatory state.
It should be noted that disc cells transfected or transduced with the NFκB-hTIMP1 plasmid exhibited some basal hTIMP1 expression compared to non-transfected or non-transduced cells. Such basal hTIMP1 expression accounted for decreased MMP activity in disc cells containing pAAV-NFκB-hTIMP1 when compared to non-transfected disc cells. It is likely that the basal hTIMP1 expression observed in disc cells transduced or transfected with AAV-NFκB-hTIMP1 was due to the cellular stress associated with the transduction or transfection procedure.
This study is limited to testing the novel regulated system in in vitro cell cultures. Future in vivo studies are required to demonstrate the clinical use of this inducible transgene system in restoration of disc matrix and amelioration of IDD. In vivo studies will also present new challenges, specifically how gene therapy will be delivered to disc cells. A minimally invasive procedure such as an injection would be preferable. However, there are complications associated with injections including volume constraints, damage to the AF, and cell leakage [32]. Despite these limitations, this study established a novel inducible system to regulate transgene expression of hTIMP1. The novel inducible system ensures that hTIMP1 expression is high only under conditions of stress, such as inflammation, that activates NFκB signalling. This inducible system facilitates the creation of new approaches towards effective and safe gene therapy to IDD and opens the door for future studies analyzing molecular therapeutics of disc degeneration. This study serves as a proof of concept. It is possible that genes other than hTIMP1 could be integrated into the inducible system extending its application beyond the treatment of IDD to other inflammatory disease states.
Change history
07 March 2021
A Correction to this paper has been published: https://doi.org/10.1007/s00586-021-06783-7
References
Reyo RA, Tsui-Wu YJ (1987) Descriptive epidemiology of low-back pain and its related medical care in the United States. Spine 12:264–8
DePalma MJ, Ketchum JM, Saullo T (2011) What is the source of chronic low back pain and does age play a role? Pain Med 12:224–233
Vo NV, Hartman RA, Patil PR, Risbud MV, Kletsas D, Iatridis JC, Hoyland JA, Le Maitre CL, Sowa GA, Kang JD (2016) Molecular mechanisms of biological aging in intervertebral discs. J Orthop Res 34:1289–1306
Peng BG (2013) Pathophysiology, diagnosis, and treatment of discogenic low back pain. World J Orthop 4:42
Ito K, Creemers L (2013) Mechanisms of intervertebral disk degeneration/injury and pain: a review. Glob Spine J 3:145–152
Le Maitre CL, Hoyland JA, Freemont AJ (2007) Interleukin-1 receptor antagonist delivered directly and by gene therapy inhibits matrix degradation in the intact degenerate human intervertebral disc: an in situ zymographic and gene therapy study. Arthritis Res Ther 9:R83
Leckie SK, Bechara BP, Hartman RA, Sowa GA, Woods BI, Coelho JP, Witt WT, Dong QD, Bowman BW, Bell KM, Vo NV, Wang B, Kang JD (2012) Injection of AAV2-BMP2 and AAV2-TIMP1 into the nucleus pulposus slows the course of intervertebral disc degeneration in an in vivo rabbit model. Spine J 12:7–20
Kozaci LD, Guner A, Oktay G, Guner G (2006) Alterations in biochemical components of extracellular matrix in intervertebral disc herniation: role of MMP-2 and TIMP-2 in type II collagen loss. Cell Biochem Funct 24(5):431–436
Roberts S, Caterson B, Menage J, Evans EH, Jaffray DC, Eisenstein SM (2000) Matrix metalloproteinases and aggrecanase: their role in disorders of the human intervertebral disc. Spine J 25:3005–3013
Hubert MG, Vadala G, Sowa G, Studer RK, Kang JD (2008) Gene therapy for the treatment of degenerative disk disease. J Am Acad Orthop Surg 16:312–319
Liu Y, Yu T, Ma X, Xiang H, Hu Y, Chen B (2016) Lentivirus-mediated TGF-β3, CTGF and TIMP1 gene transduction as a gene therapy for intervertebral disc degeneration in an in vivo rabbit model. Exp Ther Med 11:1399–1404
Wallach CJ, Kim JS, Sobajima S, Lattermann C, Oxner WM, McFadden K, Robbins PD, Gilbertson LG, Kang JD (2006) Safety assessment of intradiscal gene transfer: a pilot study. Spine J 6:107–112
Vadala G, Sowa GA, Smith L, Hubert MG, Levicoff EA, Vincenzo D, Gilbertson LG, Kang JD (2007) Regulation of transgene expression using an inducible system for improved safety of intervertebral disc gene therapy. Spine 32:1381–1387
Sowa G, Westrick E, Pacek C, Coelho P, Patel D, Vadala G, Georgescu H, Vo N, Studer R, Kang J (2011) In vitro and in vivo testing of a novel regulatory system for gene therapy for intervertebral disc degeneration. Spine 36:E623–E628
Marshall E (1999) Gene therapy death prompts review of adenovirus vector. Science 286(4558):2244–2245
Yang Y, Nunes FA, Berencsi K, Furth EE, Gonczol E, Wilson JM (1994) Cellular immunity to viral antigens limits E1-deleted adenoviruses for gene therapy. Proc Natl Acad Sci 91:4407–4411
Tripathy SK, Black HB, Goldwasser E, Leiden JM (1996) Immune responses to transgene–encoded proteins limit the stability of gene expression after injection of replication–defective adenovirus vectors. Nat Med 2:545
Daya S, Berns KI (2008) Gene therapy using adeno-associated virus vectors. Clin Microbiol Rev 21:583–593
Lawrence T (2009) The nuclear factor NF-κB pathway in inflammation. Cold Spring Harb Perspect Biol 1:a001651
Wang B, Li J, Xiao X (2000) Adeno-associated virus vector carrying human minidystrophin genes effectively ameliorates muscular dystrophy in mdx mouse model. Proc Natl Acad Sci 97:13714–13719
Xiao X, Li J, Samulski R (1998) Production of high-titer recombinant adeno-associated virus vector in the absence of helper adenovirus. J Virol 72:2224–2232
Sowa GA, Coelho JP, Vo N, Pacek C, Westrick E, Kang JD (2012) Cells from degenerative intervertebral discs demonstrate unfavourable responses to mechanical and inflammatory stimuli: a pilot study. Am J Phys Med Rehabil Acad Physiatr 91:846
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2− ΔΔCT method. Methods 25:402–408
Altun I (2016) Cytokine profile in degenerated painful intervertebral disc: variability with respect to duration of symptoms and type of disease. Spine J 16(7):857–861
Wuertz K, Vo N, Kletsas D, Boos N (2012) Inflammatory and catabolic signalling in intervertebral discs: the roles of NF-κB and MAP kinases. Eur Cell Mater 23:103–120
Kim JH, Studer RK, Sowa GA, Vo NV, Kang JD (2008) Activated macrophage-Like THP-1 cells modulate annulus fibrosus cell production of inflammatory mediators in response to cytokines. Spine 33(21):2253–2259
Nerlich AG, Bachmeier BE, Schleicher E, Rohrbach H, Paesold G, Boos N (2007) Immunomorphological analysis of rage receptor expression and NF-κB Activation in tissue samples from normal and degenerated intervertebral discs of various ages. Ann New York Acad Sci 1096:239–248
Nasto LA, Seo HY, Robinson AR, Tilstra JS, Clauson CL, Sowa GA, Ngo K, Dong Q, Pola E, Lee JY, Niedernhofer LJ, Kang JD, Robbins PD, Vo NV (2012) ISSLS prize winner inhibition of NF-κB activity ameliorates age-associated disc degeneration in a mouse model of accelerated aging. Spine 37(21):1819–25
Sun Z, Yin Z, Liu C, Tian J (2015) The changes in the expression of NF-κB in a degenerative human intervertebral disc model. Cell Biochem Biophys 72:115–122
Bartfeld S, Hess S, Bauer B, Machuy N, Ogilvie LA, Schuchhardt J, Meyer TF (2010) High-throughput and single-cell imaging of NF-κB oscillations using monoclonal cell lines. BMC Cell Biol 11:21
Savas T, Hughey JJ, Lee TK, Lipniacki T, Quake SR, Covert MW (2010) Single-cell NF-κB dynamics reveal digital activation and analogue information processing. Nature 466:267–271
Smith LJ, Silverman L, Sakai D, Le Maitre CL, Mauck RL, Malhotra NR, Lotz JC, Buckley CT (2018) Advancing cell therapies for intervertebral disc regeneration from the lab to the clinic: recommendations of the ORS spine section. JOR Spine 1(4):e1036
Acknowledgements
The work was supported in part by the National Institutes of Health (AG044376-01) to NV, the Orthopaedic Surgery Department, Ferguson Grant, the National Natural Science Foundation of China (81902236) to YH, and Shanghai Pujiang Program (18PJD027) to YH. We would like to acknowledge the National Institutes of Health supported microscopy resources in the Centre for Biologic Imaging. Specifically, the confocal microscope supported by grant number 1S10OD019973-01. Thank you to Dr. Paul Robbins for providing the NFkB-promoter construct, Thomas Lozito for MMP activity assay, and Jessa Darwin for providing editorial assistance on this manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflicts of interest
Dr. Shaw reports receiving personal fees from Ceterix Orthopaedics and grants from AO Spine and Lumbar Spine Research Society outside the submitted work. Dr. Kang reports he is a shareholder in ALung and Cardiorobotics and is a consultant for OnPointSurgical outside the submitted work. All other authors have no conflicts of interest to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Han, Y., Ouyang, Z., Wawrose, R.A. et al. ISSLS prize in basic science 2021: a novel inducible system to regulate transgene expression of TIMP1. Eur Spine J 30, 1098–1107 (2021). https://doi.org/10.1007/s00586-021-06728-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00586-021-06728-0