Abstract
Study design
Investigation of the automation of radiological features from magnetic resonance images (MRIs) of the lumbar spine.
Objective
To automate the process of grading lumbar intervertebral discs and vertebral bodies from MRIs.
Summary of background data
MR imaging is the most common imaging technique used in investigating low back pain (LBP). Various features of degradation, based on MRIs, are commonly recorded and graded, e.g., Modic change and Pfirrmann grading of intervertebral discs. Consistent scoring and grading is important for developing robust clinical systems and research. Automation facilitates this consistency and reduces the time of radiological analysis considerably and hence the expense.
Methods
12,018 intervertebral discs, from 2009 patients, were graded by a radiologist and were then used to train: (1) a system to detect and label vertebrae and discs in a given scan, and (2) a convolutional neural network (CNN) model that predicts several radiological gradings. The performance of the model, in terms of class average accuracy, was compared with the intra-observer class average accuracy of the radiologist.
Results
The detection system achieved 95.6% accuracy in terms of disc detection and labeling. The model is able to produce predictions of multiple pathological gradings that consistently matched those of the radiologist. The model identifies ‘Evidence Hotspots’ that are the voxels that most contribute to the degradation scores.
Conclusions
Automation of radiological grading is now on par with human performance. The system can be beneficial in aiding clinical diagnoses in terms of objectivity of gradings and the speed of analysis. It can also draw the attention of a radiologist to regions of degradation. This objectivity and speed is an important stepping stone in the investigation of the relationship between MRIs and clinical diagnoses of back pain in large cohorts.
Level of Evidence: Level 3.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Back pain is one of the most important causes of life-long disability worldwide [1], resulting in enormous medical and social costs. Although around 85% of cases have no clear diagnoses [2, 3], many studies indicate that degeneration of the intervertebral disc is involved [4, 5]. MRI classifications of features of disc degeneration have become a major diagnostic tool, even though many with disc degeneration features are asymptomatic [6].
The uncertain association between radiological features of disc degeneration and back pain may be due to the definitions of the features themselves. Indeed, while numerous studies have investigated possible causes of disc degeneration, interpretation of the results is complicated by lack of a standardized MRI disc degeneration phenotype [7]. Improvements in the consistency, accuracy and objectivity of measurement of radiological features would improve understanding back pain in general. It would also aid clinical reporting. To this end, a number of studies have initiated work on automated systems for grading MRIs. To date, only a system for generating Pfirrmann scores has been developed [8], which requires human input.
Here we aim to automate the grading process of Spinal MRIs for all radiological features scored routinely. A simple pipeline of our approach can be seen in Fig. 1. This automation of predicting or determining radiological scores from the scans has three key benefits: (1) the results are generated consistently; (2) radiologists can concentrate their attention and expertise on alerts and potentially problematic areas; (3) it would help researchers to measure cohorts containing large amounts of lumbar MRI data.
Overview of the system. The input to the processing pipeline is a T2 sagittal MRI (including all slices) and the outputs are the predictions of the radiological features. Shown in a is the MR volume, which can vary in resolution and number of slices. The MR volume is first processed by the vertebrae detection system where we detect and label each lumbar vertebra in the volume as shown in b where the red boxes are the bounding regions of detected vertebrae (5). We detect the vertebrae instead of the discs as they are inherently easier to detect in the MR volume. From these vertebrae the intervertebral disc region can be easily extracted from its adjacent pair of vertebrae shown in (6) c. The image used to illustrate c is from the mid-sagittal slice but in actuality the disc region is volumetric. For each given MR volume, we process and analyze six intervertebral discs. The final process in the pipeline is where the disc volume is classified by a classifier (d), where we predict radiological features (10) e
Materials and methods
Dataset
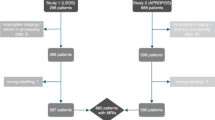
This study is based on a cohort collected during the Genodisc Project. The primary selection for recruitment to Genodisc was “patients who seek secondary care for their back pain or spinal problem”. Genodisc sourced MRI scans from centers in UK, Hungary, Slovenia and Italy. The scans from the study were not standardized, came from routine care in a number of different centers using different machines, and resulted in scans which varied in acquisition protocol. In this study, we excluded subjects whose MRI scan was of poor quality or in a non-DICOM format and used only the T2 sagittal scans. The scans were annotated with various radiological scores (global, the whole spine, and local, per disc) by a single expert experienced spinal radiologist (IMcC).
In all, we obtained images of 12,018 individual discs, six discs per patient, and their scores. Some scans contained fewer than six discs but the majority showed the complete lumbar region. To train and test the performance of our system, patients were grouped into two different sets, 90% in a training set of 1806 patients, and 10% in an independent sample of 203 patients. The test set, used to test the accuracy or concurrent validity of the automated ratings, was compared to the reference standard of expert ratings of the experienced radiologist.
System overview
An overview of the system is shown in Fig. 1. The system uses routine MRI scans acquired from a DICOM file stored on a standard laptop computer. The first step in the analysis is the delineation of the vertebral bodies and then the discs. The discs are then analyzed for the desired radiological features, and then classified. Here the automatically generated classification was compared with the radiologist’s score of each feature.
Intervertebral disc localization
The radiological scores for analysis of the discs are tied to each intervertebral disc, with the six discs per patient (T12-Sacrum) usually visible in standard clinical MRI protocols; these discs have to be accurately detected. In the first part of the study, we followed a conventional image analysis approach that detects vertebral bodies from T12 to S1 [9, 10]. From these detected vertebral bodies, a more suitable region is defined and annotated, i.e., T12–L1 to L5–S1, for each spine (Fig. 2).
Detection process. a Bounding regions, in red overlaid on the mid-sagittal slice of the scan, b the three views of the whole 3D volumes of the bounding region, and c the resulting extracted disc regions (only mid-sagittal slice is shown in the examples but in actuality each disc region consists of multiple slices)
The detection regions are in the form of 3D bounding volumes in the scan where each volume includes a disc and the surrounding upper and lower endplate regions. These volumes are normalized to reduce the signal inhomogeneity across a scan, and are centered on the detected middle slice for each disc to reduce lateral shifts (for example, from a scoliosis) (Fig. 3). Examples of the output regions are shown in Fig. 4.
Examples of the radiological features on examples of discs. Pfirrmann Grading and Disc Narrowing are graded on the mid-sagittal slices, while the other radiological features can appear anywhere in the volume. The automatic system operates on all slices of the input scan. Both Pfirrmann Grading and Disc Narrowing have multiple gradings, 1 to 5 for Pfirrmann and 1 to 4 for disc narrowing, which are shown in the example. However, the other radiological features are binary, i.e., the discs are labeled as either normal or pathological, and the examples shown are pathological examples for each feature
Radiological scores classification
In the second part of the study, a classifier, which predicts the radiological features, is then trained using the detected regions as the input, and the prior determinations of the radiological features from the experienced radiologist’s assessments as the output. Since each intervertebral level/disc possesses eight radiological scores, preferably the classifier used must be able to simultaneously predict them without human intervention. To this end, we opted for a convolutional neural network (cnn), which can both learn without feature crafting (human input), and classify multiple scores at once. Hence, there is no need to create individual descriptors for the classifier suited for each radiological score. This method is the current state-of-the-art approach in machine learning, and employs deep learning [11]. This is the use of multiple layers of abstraction to describe the relationship between the raw input data [12]. Another advantage of using a CNN model as a classifier is the possibility of ease of troubleshooting predictions of the model. For each prediction of a specific radiological score, there exists a corresponding probability that suggests the degree of confidence of the prediction of the model.
Radiological features
This study has focused on six main radiological features that can be seen in part or totally on sagittal T2 images (Fig. 5): (1) Pfirrmann grading, (2) disc narrowing, (3) spondylolisthesis, (4) central canal stenosis, (5) endplate defects, and (6) marrow signal variations (Modic changes).
a Example of the detected region of the vertebrae and the corresponding assessments of the mid-sagittal slice of an MRI. The red boxes are the detected vertebrae regions and the blue boxes are the extracted disc regions passed through to the classifier. b L2–L3 and L5–S1 disc volume examples from a and their resulting predictions computed from the disc volumes. Likewise, d the L1–L2 and L5–S1 disc volume examples from c and the predictions
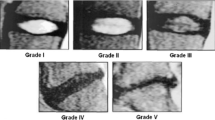
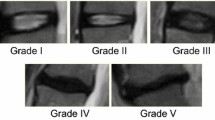
‘Pfirrmann grading’ classifies disc degeneration using criteria of disc signal heterogeneity, brightness of the nucleus and disc height into 5 grades [13]. ‘Disc narrowing’ is defined as a multi-class measurement of the disc heights; 4 grades. In this study, ‘spondylolisthesis’ is a binary measure of the vertebral slip. ‘Central canal stenosis’ is the constriction of the central canal, in the region adjacent to each intervertebral disc. The radiologist’s score is based on assessment of both sagittal and axial images, so we have only studied a binary ‘present’ or ‘absent’ stenosis. ‘Endplate defects’ are any deformities of the endplate regions, both upper and lower, with respect to the intervertebral disc. ‘Marrow signal variations’ can be viewed as either Type 1 or Type 2 Modic changes, as both T1 and T2 scans are needed to differentiate the two types. Both types of Modic changes manifest as visible signal variations at the endplate extending into the vertebral body, observed on a T2 scan [14]. Table 1 shows a summary of the grading of each radiological feature and Fig. 5 shows the examples of each radiological score and some of the output examples of the system.
Statistical analysis
The performance measure used for validation was ‘class average accuracy’, this is generally used in image analysis systems for highly unbalances classifications as occurred in the Genodisc dataset [15], e.g., only 9% of discs showed upper marrow changes [16, 17]. For our benchmark, the average class intra-rater agreement was calculated from two separate sets of labels by the same radiologist on a subset of the dataset that consists of 121 patients [18]. We are essentially comparing the radiologist’s reliability against the reproducibility of our Model.
Results
Intervertebral disc localization
Figure 2 shows a typical result of the detection process summarized in Fig. 1. The bounding regions in red are overlaid on the mid-sagittal slice of the scan and the detected vertebrae are enclosed in red boxes.
The system achieved 95.6% vertebral body detection and labeling accuracy and managed to detect corners of the vertebral bodies with a maximum error of 2 mm [14]. The cases in which detection failed can be grouped into two main types: (1) corrupted/poor scan quality, and (2) presence of a transitional vertebra near the sacrum. Examples of problems with detection are shown in Fig. 3.
Radiological scores classification
The distribution of scores per disc for each radiological score can be seen in Table 2 (Genodisc data) and typical MRIs are shown in Fig. 4.
Figure 5 shows typical gradings of discs by the system for two separate spinal MRIs. Our system consistently achieved comparable performance when comparing the radiologist intra-rater agreement (agreement between the two sets of readings done by the radiologist at different times) and the accuracy of the system; see the second and third columns of Table 3. Reliability coefficients for repeated assessments by the radiologist (intra-rater) and the automated versus radiologist’s assessments can be seen in the fourth and fifth columns of Table 3.
Comparison of readings between radiologist and model
A comparison of the scores of the radiologist with the scores of the system is shown in a histogram (Fig. 6) for the same test spines. Figure 6a shows the relative gradings of Pfirrmann scores, and Fig. 6b shows the relative gradings for disc narrowing. Figure 6c shows a comparison of the binary readings for spondylolisthesis, central canal stenosis, endplate defects and marrow changes. While the comparisons are good, the main trend is that the model tends to predict more abnormal/pathological features than the radiologist.
a Pfirrmann grading; b disc narrowing; c binary radiological features. Histogram of the scores of the model compared with the radiologist. Pfirrmann grading and disc narrowing are tabulated in different sub-figures. The main trend is that the prediction from the model tends to predict more abnormal/pathological cases than the radiologist
Kappa statistics showed that the automated system achieved consistently comparable performance when comparing the radiologist intra-rater agreement (agreement between the two sets of readings done by the radiologist at different times) and the accuracy of the system (second and third columns of Table 3). Kappa values for repeated assessments by the radiologist (intra-rater) and the automated versus radiologist’s assessments can be seen in the fourth and fifth columns of Table 3.
We found that only 3.9% of the discs in the test set have differences of more than one Pfirrmann grade between our method’s determination and the radiologist’s. We found, on average, the difference between the intra-rater agreement and our model is around 0.4%.
Evidence hotspots
Figure 7 shows examples of evidence hotspots obtained by the automated method. For each prediction of a specific radiological score, there exists a corresponding heatmap, which shows where in the disc region the abnormality lies. These heatmaps, the “evidence hotspots”, can be seen to be highly specific to the score although they are only trained from labels indicating the presence and absence of a specific radiological feature such as marrow changes, i.e., a disc-specific grading instead of voxel-specific label [19]. Another advantage of using a CNN as a classifier is the ease of troubleshooting predictions of the model. These hotspots could be beneficial in aiding radiologists in assessing scans and can act as a validity check for the actual predictions of the CNN model.
Examples of disc volumes (upper in each pair) and their corresponding evidence hotspots (lower in each pair). The leftmost and rightmost images are the second and eighth slice for each disc, out of the full volume of 9 slices. Going from top to bottom are: i upper endplate defects, ii lower endplate defects, iii upper marrow change, iv lower marrow change, v spondylolisthesis, and vi central canal stenosis. Pathological examples are shown for each radiological score/classification task, with endplate defects appearing as protrusions of the discs into the vertebral bodies, and marrow changes appearing as localized discolorations of the vertebral bodies near the vertebral endplates. Note that these hotspots localize extremely well, e.g., in the lower endplate defects example the hotspots appear only in the lower endplate even though there are defects on the upper endplate. These examples are randomly selected from different patients
Discussion
Here we developed an automated system for classifying MRI features of disc degeneration, based on a multi-centre clinical dataset (Genodisc) of 2008 lumbar scans. Our automated method takes around 1–2 min to process a scan. The bottleneck in this process is the detection system. For scans of adequate quality, the vertebral bodies and discs were detected accurately in 95.6% scans (Fig. 2) with detection failing only if scans were corrupted or of poor quality, or if transitional vertebrae were present (Fig. 3). The entirety of the disc regions of the scans were used as the input data for classification of the radiological features scored (Fig. 4) as shown by the examples in Fig. 5. No extra annotations were used for the classification task; it was dependent only on the assessments provided by the radiologist. We used the reference standard of a single expert radiologist, with repeat measurement on a randomly selected cohort. The best model trained by us achieves extremely good performance on all its trained tasks, consistently close to the performance of the radiologist (Table 3). On average, we found the difference between the intra-rater agreement and our model is around 0.4%, which suggests that the model is a close automated analog of the radiologist in terms of radiological reading. A novel feature of the Model is to identify ‘Evidence Hotspots’ that are the voxels that most contribute to the degradation scores (Fig. 7).
In this study, our system excelled at determining Pfirrmann and disc narrowing grades on a relevant data set of clinical images, both in terms of accuracies and reliability scores (Kappa values). The method currently only produced comparable performance in terms of accuracies (compared to intra-rater agreement), but not reliability scores, for the other radiological scores [spondylolisthesis, central canal stenosis, endplate changes (Table 3)]. We theorized that this arises because: (1) the system was trained to perform well on average class accuracy rather than reliability scores [15], and (2) there were relatively few discs with pathological features such as spondylolisthesis or endplate defects. Furthermore, since our system currently operates only on sagittal scans, assessments such as Central Canal Stenosis, which requires both axial and sagittal information, would tend to have a lower performance. We plan on adding the capability to process both axial and sagittal scans in the near future to see if we can improve upon the performance on reliability scores. We also anticipate that we could also use our method and validate it against other disc degeneration classification systems [20,21,22]. Others have reported results of automated image analysis of lumbar MRI scans, but not on the scale that we have reported here [8]. In addition, these systems require human supervision but in our method, there is no need to create individual descriptors for the classifier suited for each radiological score (Lootus et al. [9] and Castro-Mateos et al. [8]).
It is important to note that the gradings provided by the automated system are learnt from the gradings presented to it, i.e., they depend on the reference standard. If the system, trained on the same dataset, used an assessment of grading scores, which differed somewhat from those presented here, the grades provided by the automated system would differ accordingly. The grading scores are nevertheless objective and consistent. We thus think that the automated scoring system, through its speed, consistency and objectivity, would be of particular value in providing an objective set of MRI grading scores for phenotyping disc degeneration in studies involving large cohorts of spinal MRIs.
Conclusions
We have shown that radiological scores can be predicted to an excellent standard using only the disc-specific assessments as a reference set. The proposed method is quite general, and although we have implemented it here for sagittal T2 scans, it could easily be applied to T1 scans or axial scans, and for radiological features not studied here or indeed to any medical task where label/grading might be available only for a small region or a specific anatomy of an image. One benefit of automated reading is to produce a numerical signal score that would provide a scale of degeneration and so avoid an arbitrary categorization into artificial grades.
References
Vos T, Flaxman A, NaghavI M et al (2012) Global Health; Public Health Years lived with disability (YLDs) for 1160 sequelae of 289 diseases and injuries 1990–2010: a systematic analysis for the Global Burden of Disease Study 2010. Lancet 380:2163–2196
Palmer K, Walsh K, Bendall H et al (2000) Back pain in Britain: comparison of two prevalence surveys at an interval of 10 years. BMJ 320:1577
Deyo RA, Weinstein JN (2001) Low back pain. N Engl J Med 344:363–370
de Schepper E, Damen J, van Meurs J et al (2010) The association between lumbar disc degeneration and low back pain. Spine 35:531–536
Cheung K (2010) The relationship between disc degeneration, low back pain, and human pain genetics. Spine J 10:958–960
Brinjikji W, Luetmer P, Comstock B et al (2015) Systematic literature review of imaging features of spinal degeneration in asymptomatic populations. AJNR 36:811–816
Steffens D, Hancock M, Pereira LM, Kent P, Latimer J, Maher C (2016) Do MRI findings identify patients with low back pain or sciatica who respond better to particular interventions? A systematic review. Eur Spine J 25(4):1170–1187
Castro-Mateos I, Hua R, Pozo J et al (2016) Intervertebral disc classification by its degree of degeneration from T2-weighted magnetic resonance images. Europ Spine J 25:2721–2727
Lootus M, Kadir T, Zisserman A (2015) Automated Radiological Grading of Spinal MRI. Recent advances in computational methods and clinical applications for spine imaging 20:119–130
Lootus M, Kadir T, Zisserman A (2014) Vertebrae Detection and Labelling in Lumbar MR Images. Computational methods and clinical applications for spine imaging 17:219–230
LeCun Y, Bengio Y, Hinton G (2015) Deep learning. Nature 521:436–444
Jamaludin A, Lootus M, Kadir T et al (2016) Automatic intervertebral discs localization and segmentation: a vertebral approach. Comput Methods Clin Appl Spine Imaging. doi:10.1007/978-3-319-41827-8_9
Pfirrmann C, Metzdorf A, Zanetti M et al (2001) Magnetic resonance classification of lumbar intervertebral disc degeneration. SPINE 26:1873–1878
Jamaludin A, Kadir T, Zisserman A (2016) Automatic Modic changes classification in spinal MRI. Comput Methods Clin Appl Spine Imaging. doi:10.1007/978-3-319-41827-8_2
Maji S, Rahtu E, Kannala J et al (2013) Fine-grained visual classification of aircraft. arXiv: 1306.5151 (eprint)
Cohen J (1968) Weighted kappa: nominal scale agreement with provision for scaled disagreement or partial credit. Psychol Bull 70:213–220
Sim J, Wright C (2005) The kappa statistic in reliability studies: use, interpretation, and sample size requirements. Phys Ther 85:257–268
Lin LI-K (1989) A concordance correlation coefficient to evaluate reproducibility. Biometrics 45:255–268
Jamaludin A, Kadir T, Zisserman A (2016) Automatically pinpointing classification evidence in spinal MRIs. Med Image Comput Comput Assist Interv. doi:10.1007/978-3-319-46723-8_20
Schneiderman G, Flannigan B, Kingston S et al (1987) Magnetic resonance imaging in the diagnosis of disc degeneration: correlation with discography. Spine 12:276–281
Griffith J, Wang M, Wang YXJ, Antonio G et al (2007) Modified Pfirrmann grading system for lumbar intervertebral disc degeneration. Spine 32:E708–E712
Riesenburger R, Safain M, Ogbuji R et al (2015) A novel classification system of lumbar disc degeneration. J Clin Neurosci 22:346–351
Acknowledgements
This work was supported by the RCUK CDT in Healthcare Innovation (EP/G036861/1) and the EPSRC Programme Grant Seebibyte (EP/M013774/1). The Genodisc data were obtained during the EC FP7 Project (HEALTH-F2-2008-201626).
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Conflict of interest
The authors declared that they have no potential conflict of interest.
REC/IRB approvals
UK Research Ethics Committee approval 09/H0501/95 for The European Union Health Project on Genes and Disc Degeneration called ‘Genodisc’ (FP7 Health 2007A Grant Agreement No. 201626). Equivalent IRB/REC approvals were obtained in each recruiting country.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Jamaludin, A., Lootus, M., Kadir, T. et al. ISSLS PRIZE IN BIOENGINEERING SCIENCE 2017: Automation of reading of radiological features from magnetic resonance images (MRIs) of the lumbar spine without human intervention is comparable with an expert radiologist. Eur Spine J 26, 1374–1383 (2017). https://doi.org/10.1007/s00586-017-4956-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00586-017-4956-3