Abstract
Background
Genome-wide association studies have identified genes in the transforming growth factor-β (TGFβ) signaling pathway that are responsible for regulating carcinogenesis.
Methods
We searched for single-nucleotide polymorphisms (SNPs) located within 3′-untranslated regions (3′-UTRs) that might affect the ability of miRNAs to bind genes in the TGFβ pathway for further analysis. We used TaqMan technology to genotype these SNPs in a population-based case–control study of 1147 colorectal cancer patients and 1203 matched controls in a Chinese population.
Results
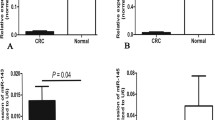
The rs1590 variant of TGFBR1 exhibited a significant association with colorectal cancer risk. Compared with individuals carrying the rs1590 TT genotype, individuals carrying the GT/GG genotypes had a decreased risk of colorectal cancer [odd ratio (OR) = 0.82, 95% confidence interval (CI) = 0.68–0.97], which was more evident among older individuals with a family history of cancer. Luciferase assays confirmed that the rs1590 T allele altered the capacity of miR-532-5p to bind TGFBR1.
Conclusions
Based on these findings, the rs1590 variant in the 3′-UTR of TGFBR1 may contribute to the susceptibility to colorectal cancer, predominantly by altering miR-532-5p binding.

Similar content being viewed by others
Abbreviations
- GWAS:
-
Genome-wide association study
- SNP:
-
Single-nucleotide polymorphism
- 3′-UTR:
-
3′-untranslated region
- HWE:
-
Hardy–Weinberg equilibrium
- OR:
-
Odd ratio
- CIs:
-
Confidence intervals
- TGFβ:
-
Transforming growth factor-β
References
Siegel RL, Miller KD, Fedewa SA, et al. Colorectal cancer statistics, 2017. CA Cancer J Clin. 2017;67:177–93.
Chen W, Zheng R, Baade PD, et al. Cancer statistics in China, 2015. CA Cancer J Clin. 2016;66:115–32.
de la Chapelle A. Genetic predisposition to colorectal cancer. Nat Rev Cancer. 2004;4:769–80.
Peters U, Bien S, Zubair N. Genetic architecture of colorectal cancer. Gut. 2015;64:1623–36.
Tenesa A, Dunlop MG. New insights into the aetiology of colorectal cancer from genome-wide association studies. Nat Rev Genet. 2009;10:353–8.
Ikushima H, Miyazono K. TGFbeta signalling: a complex web in cancer progression. Nat Rev Cancer. 2010;10:415–24.
Blobe GC, Schiemann WP, Lodish HF. Role of transforming growth factor beta in human disease. N Engl J Med. 2000;342:1350–8.
Valle L, Serena-Acedo T, Liyanarachchi S, et al. Germline allele-specific expression of TGFBR1 confers an increased risk of colorectal cancer. Science. 2008;321:1361–5.
Zhong R, Liu L, Zou L, et al. Genetic variations in the TGFbeta signaling pathway, smoking and risk of colorectal cancer in a Chinese population. Carcinogenesis. 2013;34:936–42.
Ryan BM, Robles AI, Harris CC. Genetic variation in microRNA networks: the implications for cancer research. Nat Rev Cancer. 2010;10:389–402.
Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116:281–97.
Ma L, Zhu L, Gu D, et al. A genetic variant in miR-146a modifies colorectal cancer susceptibility in a Chinese population. Arch Toxicol. 2013;87:825–33.
Agarwal V, Bell GW, Nam JW, et al. Predicting effective microRNA target sites in mammalian mRNAs. eLife. 2015;4:e05005.
Bardhan K, Liu K. Epigenetics and colorectal cancer pathogenesis. Cancers (Basel). 2013;5:676–713.
Markowitz SD, Bertagnolli MM. Molecular origins of cancer: molecular basis of colorectal cancer. N Engl J Med. 2009;361:2449–60.
Al-Tassan NA, Whiffin N, Hosking FJ, et al. A new GWAS and meta-analysis with 1000Genomes imputation identifies novel risk variants for colorectal cancer. Sci Rep. 2015;5:10442.
Bellam N, Pasche B. Tgf-beta signaling alterations and colon cancer. Cancer Treat Res. 2010;155:85–103.
Pasche B, Knobloch TJ, Bian Y, et al. Somatic acquisition and signaling of TGFBR1*6A in cancer. JAMA. 2005;294:1634–46.
Wang YQ, Qi XW, Wang F, et al. Association between TGFBR1 polymorphisms and cancer risk: a meta-analysis of 35 case-control studies. PLoS One. 2012;7:e42899.
Zhang X, Wu L, Sheng Y, et al. The association of polymorphisms on TGFBR1 and colorectal cancer risk: a meta-analysis. Mol Biol Rep. 2012;39:2567–74.
Song P, Zhu H, Zhang D, et al. A genetic variant of miR-148a binding site in the SCRN1 3′-UTR is associated with susceptibility and prognosis of gastric cancer. Sci Rep. 2014;4:7080.
Wang M, Du M, Ma L, et al. A functional variant in TP63 at 3q28 associated with bladder cancer risk by creating an miR-140-5p binding site. Int J Cancer. 2016;139:65–74.
Chang YY, Kuo WH, Hung JH, et al. Deregulated microRNAs in triple-negative breast cancer revealed by deep sequencing. Mol Cancer. 2015;14:36.
He H, Wang L, Zhou W, et al. MicroRNA expression profiling in clear cell renal cell carcinoma: identification and functional validation of key miRNAs. PLoS One. 2015;10:e0125672.
Lee H, Park CS, Deftereos G, et al. MicroRNA expression in ovarian carcinoma and its correlation with clinicopathological features. World J Surg Oncol. 2012;10:174.
Wang F, Chang JT, Kao CJ, et al. High Expression of miR-532-5p, a tumor suppressor, leads to better prognosis in ovarian cancer both in vivo and in vitro. Mol Cancer Ther. 2016;15:1123–31.
Song X, Wang Z, Jin Y, et al. Loss of miR-532-5p in vitro promotes cell proliferation and metastasis by influencing CXCL2 expression in HCC. Am J Transl Res. 2015;7:2254–61.
Acknowledgements
The authors alone are responsible for the content and writing of the article.
Author information
Authors and Affiliations
Contributions
Jinfei Chen and Meilin Wang conceived and designed the experiments. Dongying Gu, Shuwei Li, and Mulong Du wrote the paper. Cuju Tang, Haiyan Chu, and Na Tong contributed reagents/materials/analysis tools. Dongying Gu, Zhengdong Zhang, and Jinfei Chen recruited samples. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors disclose no potential conflicts of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gu, D., Li, S., Du, M. et al. A genetic variant located in the miR-532-5p-binding site of TGFBR1 is associated with the colorectal cancer risk. J Gastroenterol 54, 141–148 (2019). https://doi.org/10.1007/s00535-018-1490-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00535-018-1490-y