Abstract
Convolutional Neural Networks (CNN) with different architectures have shown promising results in skin cancer diagnosis. However, CNN has a high computational cost, which makes the need for a light version of CNN a desirable step. This version can be used on small devices, such as mobile phones or tablets. A light version can be created using pruning techniques. In this study, iterative magnitude pruning (IMP) is utilized. This method depends on pruning the network iteratively. The IMP method is applied on AlexNet with transfer learning (TL) and data augmentation. The proposed IMP AlexNet with TL is applied on three different skin cancer datasets which are PAD-UFES-20, MED-NODE, and PH2 dataset. The datasets used are a combination of smartphone, dermoscopic, and non-dermoscopic images. Different CNN versions are applied on the same datasets for comparison with IMP AlexNet. The CNNs used are VGG-16, ShuffleNet, SqueezNet, DarkNet-19, DarkNet-53, and Inception-v3. The proposed IMP AlexNet achieved accuracies of 97.62%, 96.79%, and 96.75%, with accuracy losses of 1.53%, 2.3%, and 2.2%, respectively, compared to the original AlexNet. In addition, the proposed IMP AlexNet requires less running time and memory usage than the traditional AlexNet. The average running time for IMP AlexNet is 0.45 min, 0.28 min, and 0.3 min, for PAD-UFES-20, MED-NODE, and PH2 datasets, respectively. The average RAM usage with IMP AlexNet is 1.8 GB, 1.6 GB, and 1.7 GB, respectively. IMP AlexNet accelerates the average running time by approximately 15 times that of the traditional AlexNet and reduces the average RAM used by 40%.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Deep neural networks (DNN) play a major and influential role in many aspects of scientific research, such as medical diagnosis [1, 2], remote sensing [3], agriculture [4], and different fields of research.
Skin cancer diagnosis is one of the DNN applications in the medical field. Skin cancer incidence is on the rise dramatically. According to the World Health Organization, the number of newly diagnosed cases worldwide in 2020 is 324,635 and the number of deaths is 57,043 [5, 6]. According to the official website of the Skin Cancer Foundation, the number of deaths caused by melanoma is expected to increase by 4.4 percent in 2023. In addition, it is estimated that the number of diagnosed cases of melanoma will reach 186,680 by the end of 2023 in the USA [7]. Studies conducted in different hospitals in Africa found that skin cancer diagnosis accounted for 13% of the total number of diagnosed malignancies [8].
DNN achieves promising results in skin cancer diagnosis in different image types, such as dermoscopic images [9], smartphone images [10], and non-dermoscopic images [11]. Researchers have faced a problem with DNN, which has a high computational cost. Skin cancer applications using DNN require expensive environments [9, 12]. During running, it takes a very long time to obtain results and has high memory usage.
DNN with its original version cannot be used in small devices, mobile phones, tablets, or even normal computers in clinics and hospitals. Therefore, to use a skin cancer detection application using DNN, some adjustments must be made to create a light version of the DNN. This light version can then be used in small devices or in devices without high capabilities. Using smartphones and tablets enables us to benefit from computing and communication features in one device that can be light and easy to carry in a pocket, allowing easy access and use in times of need.
Creating a light version of DNN directs research toward pruning techniques. Pruning is the process of eliminating the least influential parameters from a current network. The goal of the pruning process is to increase the efficiency of the network while maintaining its accuracy. Then, the computational cost required for running the neural network is reduced.
To the best of our knowledge, there is a shortage of pruning DNN research on skin cancer detection. Most previous work on pruning in medical image applications concentrates on magnetic resonance imaging (MRI), Computed Tomography (CT), ultrasound images [13], microscopic images [14], and X-ray images [15].
The proposed technique is Iterative Magnitude Pruning (IMP), which is applied on AlexNet because it has the highest performance in skin cancer detection research [16] and achieves the highest accuracy in the lowest running time. It is shown that when IMP is applied to AlexNet, the running time and memory usage are reduced without a significant loss in accuracy.
The proposed method is tested on three different skin cancer datasets. The results are compared with those of traditional AlexNet and six different CNNs. This comparison proves the robustness of IMP AlexNet compared to other CNNs.
The sections in this paper are arranged as follows: Sect. 2 discusses previous work on pruning techniques, Sect. 3 explains the proposed algorithm of IMP AlexNet, Sect. 4 demonstrates the results and discussions, and the final section provides the conclusion and future work.
2 Related work
Pruning techniques were used in different applications of DNN based on previous studies. Some are general research that uses datasets of different objects. Other previous studies are interested in specific applications, for example, remote sensing. Few studies have been conducted in medical imaging applications.
2.1 Pruning methods used in general applications:
Studies that use datasets with both general or different objects such as the method in [17]. An acceleration technique for CNN has been proposed in this study. Where they apply pruning on filters in CNN. The pruned filters have little effect on the accuracy of the output. The method was applied on VGG-16 and Resnet-110 and achieved accuracy close to the original on the CIFAR-10 dataset [18].
The research in [19] proposed an asymptotic soft filter pruning (ASFP) technique. In the first step, the pruned filters are updated during the retraining phase, then more filters are pruned asymptotically during the training phase. The technique is applied on VGGnet and ResNet using CIFAR-10 [18]. The accuracy of ASFP on VGGnet was 93.37%, while the original net was 93.58%. The accuracy of ResNet was 93.12%, while the original ResNet was 93.59%.
The method in [20] depends on the iterative pruning technique that is applied on DenseNet. The method aims to reduce network complexity by removing nodes and filters with the lowest value near zero. The average value of removed parameters is determined by all training samples used. It was demonstrated that 90% of the parameters can be deleted without any significant loss in accuracy. The datasets used are the MNIST [21], CIFAR-10 [18], and Tiny ImageNet [22].
Mask Soft filter pruning (M-SFP) is the method proposed in [23]. The method is applied to ResNet-56. The method keeps the weights without zeroing the values. This is done by creating a mask for the feature map which corresponds to the features that will be pruned. The method achieved an accuracy of 93.9% with an accuracy reduction of 0.17%. The used datasets are CIFAR-10 and CIFAR-100 [18].
A study proposed a model of the pruning method for ANNs based on iterative magnitude pruning [24]. The method aims to reduce the epochs number of the intermediate iterations of IMP in the re-training process. The study applied the method to VGG-19 and used the CIFAR-10 dataset [18], achieving an accuracy of 90.6%.
A new technique was proposed in [25] for pruning pre-trained models layer-by-layer with a predefined compression ratio. The technique involves computing a relevance measure to identify the most critical units, and then pruning the channels with less information. The method was applied to VGG-16, ResNet-20, and ResNet-32, resulting in an accuracy drop of 0.86%, 0.12%, and 0.02%, respectively, on the CIFAR-10 dataset [18].
A pruning algorithm described in [26] removes weights from a network based on their gradients and magnitudes against the test dataset. The algorithm was applied to MobileNet and resulted in a 3.8% accuracy drop on the CIFAR-10 dataset [18].
2.2 Pruning methods in specific applications
The study in [27] proposed a method that uses an ensemble learning machine to achieve high accuracy in classifying different hyperspectral images. The method selects classifiers with robust complementarity and adds them iteratively to the ensemble. The ensemble is then pruned based on the accuracy array of the ensemble. If the validation accuracy of the ensemble doesn’t change after several iterations, the iterations are stopped to save computational time. The accuracy achieved by the algorithm ranges from 94 to 97%.
In [28], a filter pruning model is proposed for remote sensing image classification. The method involves removing filters that cannot learn semantic meanings in proportion to a predefined pruning rate. The study applies the method on VGG-16, VGG-19, and AlexNet using the UC Merced dataset [29] and the NWPU-RESISC45 dataset [30]. The results show a reduction in accuracy by 0.4%, 0.4%, and 0.45%, respectively.
A new method called Iterative Network Pruning with Uncertainty Regularization for Lifelong Sentiment Classification (IPRLS) was presented in [31]. The method is an iterative pruning method that removes frequent parameters in large deep networks to free up space for new tasks. The BERT [32] (bidirectional transformers for language understanding) model is used as the base model for sentiment classification, and the method is applied to 16 popular datasets (books, DVDs, magazines, …etc.). The average accuracy achieved ranges from 80 to 91%.
The Stack Attention-Pruning method is a technique proposed in [33] that is applied to Graph Convolutional Networks (GCN) for image classification in remote sensing. The method involves pruning and removing pixels that are lowly correlated to each other and constructing a refined graph of neighborhood-correlated pixels. The method achieved accuracy ranging from 96.7% to 97.3% on two public datasets, Indian Pines [34] and Salinas [35].
2.3 Pruning methods in medical applications
Pruning applications in medical diagnosis are limited, but in this sub-section, some examples of pruning in medical diagnosis are shown.
In a study on Pap smear image classification, a pruning technique called adaptive pruning deep transfer learning was proposed [14]. The model used in the study was divided into 10 convolutional layers and three fully connected layers. Due to the limited number of images, transfer learning was applied to use a pre-trained model. The next step was to prune the convolutional layer by removing some convolutional kernels that may affect the target task. The proposed method was tested on 389 cervical Pap smear images and achieved an accuracy of more than 98%.
The STAMP algorithm is a pruning model that allows simultaneous training and pruning of a U-Net architecture for medical image segmentation [13]. The model is based on filter ranking, where filters are pruned based on their ranking scores. The model has been shown to improve network performance while reducing the size of the U-Net by more than 85% in terms of parameters. The STAMP algorithm has been applied to various medical image datasets, including Brain MRI images [36], Cardiac MRI images [37], Spleen CT images [38], Prostate MRI images [39], and Brain ultrasound datasets [40].
The proposed algorithm in [41] is based on DNN deepening and pruning. The model is presented for the diagnosis of medical images. It is divided into two phases. The first phase is deepening, in which a DNN is allowed to grow by adding residual blocks iteratively on top of the created DNN without ever removing a previously added block. After reaching a suitable size of DNN the pruning phase starts. In the pruning phase, redundant parameters are deleted. The method is applied on ResNet and approximately maintains the same accuracy as the original networks. The proposed algorithm achieves 80.4% accuracy, while the original networks of ResNet achieve accuracy ranges from 80.2% to 80.7% when both methods are applied on the ISIC 2016 dataset [42].
3 Proposed model
The IMP method has been applied to AlexNet, and the resulting IMP AlexNet has been tested on three different datasets. The performance of the proposed model has been compared with different CNN versions using different optimizers to test its robustness and performance.
3.1 Dataset and pre-processing
The research uses three datasets to compare the model’s performance on different datasets. The first dataset is PAD-UFES-20 [43], which is composed of 2298 smartphone images for six different skin cancer types. In this research, two classes are used which are naevus and melanoma with 244 and 52 images, respectively.
The second used dataset is the MED-NODE dataset [44], which consists of 170 non-dermoscopic images (simple digital images) for two classes. 70 images for the melanoma class and 100 for the naevus class.
The third used dataset is the PH2 Dataset [45]. It consists of 200 dermoscopic images, 160 for naevus, and 40 for melanoma. Samples from the used datasets are shown in Fig. 1.
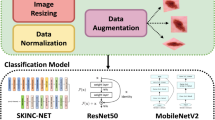
During the pre-processing phase, it is necessary to resize all images to a fixed size before inputting them to the CNN. The input size for each CNN version varies, with AlexNet and SqueezNet requiring an input size of 227 × 227 × 3, VGG-16 and ShuffleNet requiring 224 × 224 × 3, DarkNet-19 and DarkNet-53 requiring 256 × 256 × 3, and Inception-V3 requiring 299 × 299 × 3.
3.2 Data augmentation
The augmentation methods used are random rotation with rang [− 5, 5], random x reflection, random y reflection with 50% probability, random x shear with range [− 0.05, 0 05], random y shear with range [− 0.05, 0.05], random x scale with range [0.5, 1], random y scale with range [0.5, 1], random X translation with range [− 5, 5], and random Y translation with range [− 5, 5], Table 1 shows the change in number of images for each dataset after applying data augmentation techniques.
3.3 Transfer learning
Transfer learning is a popular approach in deep learning that involves reusing a pre-trained model on a new problem. This approach is useful in situations where a lot of data is needed to train a neural network from scratch, but access to that data is not always available. Transfer learning can train deep neural networks with comparatively little data, which is very useful in the data science field since most real-world problems typically do not have millions of labelled data points to train such complex models.
By applying transfer learning to a new task, one can achieve significantly higher performance than training with only a small amount of data. Transfer learning can save time and resources from having to train a new model from scratch for every new task. It can also help with computational costs by taking the conceivable parts of pre-trained CNN models and applying these parts to a new task problem. This is shown in Fig. 2.
In this research, the TL technique is applied to all pre-trained CNN models used, including IMP AlexNet as used before in [16]. The pre-trained CNN models are loaded without the last three layers, which are the fully connected layer, the SoftMax layer, and the classification layer for 1000 classes. Then, new layers are added on top of the pre-trained CNNs to adjust them to skin cancer classification tasks. The new layers include a new fully connected layer, a new SoftMax layer, and a new classification layer to classify two classes, which are melanoma and naevus.
3.4 Iterative magnitude pruning model
The Iterative Magnitude Pruning idea is a method of pruning neural networks that assign scores to the connections of the network based on their absolute value, which corresponds to their relative effect on the trained network accuracy.
The hierarchy of the IMP AlexNet model is shown in Fig. 3. The steps of IMP start after pre-processing and augmenting the input dataset and applying transfer learning on AlexNet. In the beginning, the importance of each connection is determined by assigning a score to each one, and the scores indicate the connection’s relative effect on the target accuracy. The relative effect for each connection can be computed by the function dlupdate in Matlab. Then, these obtained scores are sorted.
A threshold is used in pruning, any connection with scores less than this threshold is removed. The threshold can be calculated using the following equation.
The threshold is computed by Eq. 1: The iteration scheme is an array of points in the range from zero to the target sparsity value, X is the number of the current iteration of the model, and A is the size of the scores array.
Sparsification is a technique used to identify and remove unnecessary connections in a neural network without affecting its accuracy. After several trials, it was found that a target sparsity value of 0.90 is the optimal value for achieving high performance.
The iterative process of creating a pruning mask and removing connections with scores less than the calculated threshold is repeated until the highest performance is reached. The number of iterations used is ten, after which there is no significant change in performance. The pseudocode for IMP can be found in Algorithm 1.

Figures 4 and 5 show the difference between the original network and the pruned network after using dlupdate function from Matlab.
3.5 Pretrained convolutional neural networks
The study used several CNNs, including VGG-16, ShuffleNet, SqueezNet, DarkNet-19, DarkNet-53, and Inception-v3, to perform binary classification of melanoma and naevus. The CNNs followed the same steps, which included pre-processing and data augmentation of the dataset, constructing the network, and applying transfer learning by replacing the last three layers with new layers for binary classification. The processing of CNN models is shown in Fig. 6.
4 Experimental results and discussions
This section discusses the experimental environment and results of the proposed IMP AlexNet and the CNNs used in the comparison. All training options and system specifications are kept constant.
After several trials with different optimizers, it is found that the ‘Adam’ optimizer achieves the highest performance as shown in Table 3. The training options used with all optimizers are as follows: the minibatch size used is 32, the number of epochs is 50, the L2 regularization used value is 0.01, the initial learning rate used is 0.0001, and the value used for learn rate drop factor is 0.3.
The proposed IMP AlexNet and the CNNs used are implemented on MATLAB 2021 64-bit. The system used has an Intel processor 2.21 GHz with core i7, 16GB RAM, and a Nvidia Geforce Gtx 1060 graphic card.
4.1 IMP AlexNet model evaluation
The performance of the IMP AlexNet model is compared with other models including traditional AlexNet, VGG-16, ShuffleNet, SqueezNet, DarkNet-19, DarkNet-53, and Inception-v3. The comparison is based on classification accuracy, average running time, and average used RAM.
The performance measures are computed for the testing dataset using the following equations: accuracy using Eq. 2, sensitivity (Recall) using Eq. 3, specificity using Eq. 4, precision using Eq. 5 [46], and F1score using Eq. 6 [47].
The variables in the equations mentioned earlier are tp for true positive, tn for true negative, fp for false positive, and fn for false negative. The equations use TPR for the true positive rate, TNR for the true negative rate, and PPV for the positive prediction value.
The IMP AlexNet model and the CNNs are run by following a specific process that involves loading the datasets, dividing them into training and testing sets with an 80/20 split, resizing the images according to the network used, applying data augmentation, and then applying the CNNs with transfer learning. The results of the models are the average of 10 repetitions of running. The confusion matrix for the three used datasets is shown in Table 2.
Table 3 presents the evaluation measures (Accuracy, Sensitivity, specificity, precision, and F1-score) of the proposed IMP AlexNet (presented in bold) compared to other CNNs. The traditional AlexNet achieved the best classification accuracy in the three datasets with 99.15%, 99.13%, and 99% for PAD-UFES-20, MED-NODE, and PH2 datasets, respectively. The proposed IMP AlexNet achieved 97.62%, 96.79%, and 96.75% for PAD-UFES-20, MED-NODE, and PH2 datasets, respectively. DarkNet-19 achieved results approximately close to the results of AlexNet with 99.1%, 98.84%, and 98.5% accuracy, but it needs more running time and memory usage as shown in Table 4.
Table 4 presents the performance measures of the compared CNN models. It shows the average accuracies, the average number of iterations in each run, the average running time of 10 repeated runs, the average RAM used by the models, and the average running time per image. The performance measures of IMP AlexNet are presented in bold, it is found that the proposed IMP AlexNet achieves 97.62% in an average running time of 0.45 min and the average RAM used is 1.8 GB on the PAD-UFES-20 dataset. When the MED-NODE dataset is used with the proposed IMP AlexNet, the average accuracy is 96.79% in an average running time of 0.28 min, and the average RAM used is 1.6 GB. On the PH2 dataset, IMP AlexNet achieved an average accuracy is 96.75% in an average running time of 0.3 min and the average RAM used is 1.7 GB.
According to Table 4, AlexNet and IMP AlexNet were not affected by the size of the dataset, as their running times with the three datasets were close to each other and achieved the highest accuracies in the table. However, DarkNet-53 and Inception-V3 showed differences in running times as the size of the dataset varied. When the size of the dataset increased, the running time increased. DarkNet-53 achieved 50.2, 42.7, and 43.6 min for PAD-UFES-20, MED-NODE, and PH2 datasets, respectively. Inception-V3 achieved 20.6, 15.3, and 16 min for the same datasets.
The running time of different neural networks was compared with the same number of iterations. For AlexNet and IMP AlexNet, the average number of iterations with PAD-UFES-20 was 350, and the running time was 5.7 and 0.45 min, respectively. On the other hand, for DarkNet-53, the number of iterations was 250, but the running time was 50.2 min. The study did not find a significant impact of the number of iterations on the running time.
The average running time per image is added to fairly compare the running time between the used CNN models. It is found that IMP AlexNet keeps the lowest running time per image and requires less than a second to classify an image in the three used datasets. Additionally, IMP AlexNet had the lowest RAM, making it a good candidate for transfer to a mobile application in future work.
A comparison between the accuracy achieved is held in Fig. 7, Group 1 refers to the traditional AlexNet, and Group 2 refers to the proposed IMP AlexNet. The traditional AlexNet has the highest accuracy compared to other CNNs, while the proposed IMP AlexNet results are slightly lower than the traditional AlexNet results. The accuracy reduction between the traditional AlexNet and the proposed IMP AlexNet is 1.53, 2.3, and 2.2 for PAD-UFES-20, MED-NODE, and PH2 datasets, respectively. Additionally, the running time and memory usage are reduced, as shown in Figs. 8 and 9.
In Fig. 8, it is observed that the average running time is reduced from the traditional AlexNet that achieved 5.7 min, 5 min, and 5.2 min to the proposed IMP AlexNet that achieved 0.45 min, 0.28 min, and 0.3 min for PAD-UFES-20, MED-NODE, and PH2 datasets, respectively.
In Fig. 9 the average used RAM is reduced from 2.8 GB, 2.2 GB, and 2.6 GB with traditional AlexNet to 1.8 GB,1.6 GB, and 1.7 GB with the proposed IMP AlexNet for PAD-UFES-20, MED-NODE, and PH2 datasets, respectively.
Table 5 lists the improvements achieved by the proposed IMP AlexNet compared to other CNNs. The first column indicates the name of the compared method, the second column indicates how many times IMP AlexNet accelerates the ordinary methods, and the third column indicates the average reduction achieved in the used RAM. The first row shows how the traditional AlexNet is improved. It is found that the IMP AlexNet accelerates the average running time by around 15 times of the traditional AlexNet, and it saves average used RAM by 40%.
4.2 Influence of unbalanced classes
In this study, the unbalanced classes did not significantly affect the classification accuracy because the difference between the classes in the used datasets was not huge. On the other hand, the Isic-2020 dataset has a significant difference in the number of samples between the two classes, it is composed of 584 for malignant 32,542 and 32,542 for benign [48]. When we applied our model to Isic-2020, it achieved high accuracy (more than 90%) although the malignant class is sometimes totally misclassified.
The IMP AlexNet model’s confusion matrix shows that there is no effect of unbalanced classes. In the PAD-UFES-20 dataset, one image is misclassified in the melanoma class and two images are misclassified in the naevus class. In the MED-NODE dataset and PH2 dataset, only one image is misclassified in each class. Class imbalance can affect the accuracy of classification models. The confusion matrix provides more insight into the accuracy of a predictive model and which classes are being predicted correctly or incorrectly.
The F1-score is used to detect if the model is a good predictor or not because the F1-score is a combination of precision and recall as shown in Eq. (6). Precision computes the correct positive predictions that the model can make. Recall computes the correct positive samples of the dataset that the model can identify. A high F1-score indicates that both precision and recall are high, while a low F1-score indicates that either precision or recall (or both) are low. Like the case of the Isic-2020 dataset, it achieved very high accuracy but the F1 score is low. The F1-score is a useful metric for evaluating model performance, especially in cases where accuracy may be misleading, such as imbalanced.
So, we can say that the proposed IMP AlexNet is a good predictor when the melanoma class resembles 18% to 42% from the used dataset, which is the case in the used datasets. The melanoma class percentages from the whole dataset are 18%, 20%, and 42% for PAD-UFES-20, PH2, and MED-NODE respectively. Unlike the case in Isic-2020, the melanoma class percentage from the whole dataset is 2% which achieved F1-score ranges from 50 to 60%.
In Table 4, with Adam optimizer, you can find that IMP AlexNet achieved an F1-score greater than 90% for the three used datasets which are 93.59%, 95.87%, 90.94% for PAD-UFES-20, MED-NODE, and PH2 datasets, respectively. According to [49], f1-score values greater than 90% are considered to be very good and this is the case with f1-scores achieved by the proposed IMP AlexNet. Accuracy cannot be the only measure for evaluating the model performance, it must be accuracy alongside with f1-score to correctly evaluate the model.
4.3 Comparison between IMP AlexNet Model and prior work
In Table 6, a comparison is held between the proposed IMP AlexNet and the previous studies that use the same datasets that we used in our study. By looking for the accuracies achieved before in column 3, it is found that our model still has the highest classification accuracy among them. The traditional Alexnet and the proposed IMP AlexNet are presented in bold.
Our model not only achieved a high accuracy, it is also outperforming the state of the art. By inspecting the limitations of each study in column 5, you will find that our model solves these limitations. First, the input image in some previous model must be a binary image [10, 54], specific colour space [51], or has low resolution [53]. Unlike the case with the proposed IMP AlexNet, it accepts coloured images with the input resolution of AlexNet.
Second, some models in prior work are to some extent complicated. Some of them have a variable number of neurons [10]. Others have a large number of layers [11, 50, 52]. Some of them use a cluster-based algorithm which has high time complexity [44]. Unlike the case in IMP AlexNet which has eight layers only. This directly affects the running time of the model.
Third, our model can test different types of skin cancer images which are dermoscopic (PH2 Dataset), non-dermoscopic (MED-NODE), and smartphone images (PAD-UFES-20). This advantage is missing in most previous studies which test their models with only one type of dataset like the case in [44, 51,52,53,54].
Fourth, the specifications of the model are clearly stated which are the running time, RAM used, and performance measures. The model results are average for ten independent runs, but some previous studies didn’t mention whether the results are average for several runs or only one run, and others take the average for a few numbers of independent runs. Some of them didn’t mention the running time and RAM used [10, 44, 50, 51, 54]. Others didn’t mention f1-score in the evaluation measures which is the indicator of the model efficiency [51].
Finally, we can say that the model in our study is an integrated step for creating a mobile application system in future work, able to test a skin lesion in real-time using a mobile phone camera or any type of skin cancer images.
5 Conclusion and future work
A method called IMP AlexNet has been developed to create a light version of CNN that can be used on mobile devices or computers with limited capabilities. The IMP AlexNet was utilized on three different skin cancer datasets, which included smartphone images, dermoscopic images, and non-dermoscopic images.
To showcase the robustness of the proposed IMP AlexNet, the results were compared with those of traditional AlexNet and other CNN models. The comparison considered three main elements which are classification accuracy, average running time, and average used RAM.
The proposed IMP AlexNet achieved high accuracies on three different skin lesion datasets: PAD-UFES-20, MED-NODE, and PH2. Specifically, the accuracies achieved were 97.62%, 96.79%, and 96.75%, respectively. These accuracies were achieved in the lowest average running time and the lowest average used RAM, which was 0.45, 0.28, and 0.3 min and 1.8, 1.6, and 1.7 GB, respectively. These results achieved the main goal of the research.
It is concluded that IMP AlexNet achieved its result with the lowest running time and used RAM. The previous observation outperforms the state of the art and makes the IMP AlexNet light version of CNNs that can be used as a mobile application in future work with accepted classification accuracy.
For future work, it is suggested the following: First, applying IMP AlexNet on datasets with multiclass skin cancer. Second, applying IMP on different CNNs for example DarkNet-19 because it achieves accuracy approximately close to AlexNet. Third, converting IMP AlexNet to a mobile application. Finally, parallel processing can be applied with the proposed IMP AlexNet which can improve the achieved results of IMP AlexNet.
Data availability
The datasets used during the current study are available online: PAD-UFES-20: https://data.mendeley.com/datasets/zr7vgbcyr2/1. MED-NODE: https://www.cs.rug.nl/~imaging/databases/melanoma_naevi/.PH2-dataset: https://www.fc.up.pt/addi/ph2%20database.html.
References
Prakash JA, Ravi V, Sowmya V, Soman KP (2022) Stacked ensemble learning based on deep convolutional neural networks for pediatric pneumonia diagnosis using chest X-ray images. Neural Comput Appl. https://doi.org/10.1007/s00521-022-08099-z
Sengar N, Joshi RC, Dutta MK, Burget R (2023) EyeDeep-Net: a multi-class diagnosis of retinal diseases using deep neural network. Neural Comput Appl. https://doi.org/10.1007/s00521-023-08249-x
Xu B (2021) Improved convolutional neural network in remote sensing image classification. Neural Comput Appl 33:8169–8180. https://doi.org/10.1007/s00521-020-04931-6
Uğuz S, Uysal N (2021) Classification of olive leaf diseases using deep convolutional neural networks. Neural Comput Appl 33(9):4133–4149. https://doi.org/10.1007/s00521-020-05235-5
Retrieved August 20, 2023: https://gco.iarc.fr/today/data/factsheets/cancers/16-Melanoma-of-skin-fact-sheet.pdf
Arnold M, Singh D, Laversanne M, Vignat J, Vaccarella S, Meheus F, Cust AE, de Vries E, Whiteman DC, Bray F (2022) Global burden of cutaneous melanoma in 2020 and projections to 2040. JAMA Dermatol 158(5):495–503. https://doi.org/10.1001/jamadermatol.2022.0160
Retrieved August 20, 2023: https://www.skincancer.org/skin-cancer-information/skin-cancer-facts/
Onyishi NT, Ohayi SR (2022) Prevalence of squamous and basal cell carcinomas in African albino skin cancer lesions: a systematic review and meta-analysis of proportion. J Skin Cancer. https://doi.org/10.1155/2022/5014610
Cabrejos-Yalán V, Rosales-Huamani J & Arenas-Ñiquin J (2022) Optimization of a deep learning model for skin cancer detection with magnitude-based weight pruning. In: World Conference on Information Systems and Technologies (pp 624–629). Springer International Publishing., Cham doi:https://doi.org/10.1007/978-3-031-04826-5_61
Moldovanu S, Obreja C, Biswas K, Moraru L (2021) Towards accurate diagnosis of skin lesions using feedforward back propagation neural networks. Diagnostics 11(6):936. https://doi.org/10.3390/diagnostics11060936
Hosny K, Kassem M (2022) Refined residual deep convolutional network for skin lesion classification. J Digit Imaging 35(2):258–280. https://doi.org/10.1007/s10278-021-00552-0
Pandey R, Uziel S, Hutschenreuther T, Krug S (2023) Towards deploying DNN models on edge for predictive maintenance applications. Electronics 12(3):639. https://doi.org/10.3390/electronics12030639
Dinsdale NK, Jenkinson M, Namburete AI (2022) STAMP: simultaneous training and model pruning for low data regimes in medical image segmentation. Med Image Anal. https://doi.org/10.1101/2021.11.26.470124
Wang P, Wang J, Li Y, Li L, Zhang H (2020) Adaptive pruning of transfer learned deep convolutional neural network for classification of cervical pap smear images. IEEE Access 8:50674–50683. https://doi.org/10.1109/ACCESS.2020.2979926
Rajaraman S, Siegelman J, Alderson PO, Folio LS, Folio LR, Antani SK (2020) Iteratively pruned deep learning ensembles for COVID-19 detection in chest X-rays. IEEE Access 8:115041–115050. https://doi.org/10.1109/access.2020.3003810
Medhat S, Abdel-Galil H, Aboutabl AE, Saleh H (2022) Skin cancer diagnosis using convolutional neural networks for smartphone images: a comparative study. J Radiat Res Appl Sci 15(1):262–267. https://doi.org/10.1016/j.jrras.2022.03.008
Li H, Kadav A, Durdanovic I, Samet H, Graf HP (2016) Pruning filters for efficient convnets. arXiv preprint arXiv:1608.08710. doi:https://doi.org/10.48550/arXiv.1608.08710
Krizhevsky A, Hinton G. (2009) Learning multiple layers of features from tiny images
He Y, Dong X, Kang G, Fu Y, Yan C, Yang Y (2019) Asymptotic soft filter pruning for deep convolutional neural networks. IEEE Trans Cybern 50(8):3594–3604. https://doi.org/10.1109/TCYB.2019.2933477
Tan C M J, Motani M (2020) Dropnet: reducing neural network complexity via iterative pruning. In: International Conference on Machine Learning 9356–9366.PMLR. doi:https://doi.org/10.48550/arXiv.2207.06646
LeCun Y, Cortes C, and Burges C. Mnist handwritten digit database. ATT Labs [Online]. Available: http://yann.lecun. com/exdb/mnist, 2, 2010
Tiny ImageNet: retrieved 2020 from https://tinyimagenet.herokuapp.com
Kim NJ, Kim H (2020) Mask-soft filter pruning for lightweight CNN inference. In: 2020 International SoC Design Conference (ISOCC) 316-317. IEEE. doi:https://doi.org/10.1109/ISOCC50952.2020.9333054
Zullich M, Medvet E, Pellegrino F A, Ansuini A. (2021) Speeding-up pruning for artificial neural networks: introducing accelerated iterative magnitude pruning. In: 2020 25th International Conference on Pattern Recognition (ICPR) 3868–3875. IEEE. doi:https://doi.org/10.1109/ICPR48806.2021.9412705
Alqahtani A, Xie X, Jones MW, Essa E (2021) Pruning CNN filters via quantifying the importance of deep visual representations. Comput Vis Image Underst 208:103220. https://doi.org/10.1016/j.cviu.2021.103220
Belay K (2022) Gradient and magnitude based pruning for sparse deep neural networks. In: Proceedings of the AAAI Conference on Artificial Intelligence vol 36(11) p 13126-13127. doi:https://doi.org/10.1609/aaai.v36i11.21699
Zhang Y, Cao G, Li X (2020) Multiview-based random rotation ensemble pruning for hyperspectral image classification. IEEE Trans Instrum Meas 70:1–14. https://doi.org/10.1109/TIM.2020.3011777
Guo X, Hou B, Ren B, Ren Z, Jiao L (2021) Network pruning for remote sensing images classification based on interpretable CNNs. IEEE Trans Geosci Remote Sens 60:1–15. https://doi.org/10.1109/TGRS.2021.3077062
Yang Y, Newsam S (2010) Bag-of-visual-words and spatial extensions for land-use classification. In: Proceedings of the 18th SIGSPATIAL international conference on advances in geographic information systems 270–279. doi:https://doi.org/10.1145/1869790.1869829
Cheng G, Han J, Lu X (2017) Remote sensing image scene classification: benchmark and state of the art. Proc IEEE 105(10):1865–1883. https://doi.org/10.1109/JPROC.2017.2675998
Geng B, Yang M, Yuan F, Wang S, Ao X, Xu R. (2021) Iterative network pruning with uncertainty regularization for lifelong sentiment classification. In: Proceedings of the 44th International ACM SIGIR conference on Research and Development in Information Retrieval 1229–1238. doi:https://doi.org/10.1145/3404835.3462902
Devlin J, Chang M W, Lee K, Toutanova K. (2018) Bert: pre-training of deep bidirectional transformers for language understanding. arXiv preprint arXiv:1810.04805. doi:https://doi.org/10.48550/arXiv.1810.04805
Liu N, Zhang B, Ma Q, Zhu Q, Liu X (2021) Stack attention-pruning aggregates multiscale graph convolution networks for hyperspectral remote sensing image classification. IEEE Access 9:44974–44988. https://doi.org/10.1109/ACCESS.2021.3061489
Bo C, Lu H, Wang D (2015) Hyperspectral image classification via JCR and SVM models with decision fusion. IEEE Geosci Remote Sens Lett 13(2):177–181. https://doi.org/10.1109/LGRS.2015.2504449
Liu S, Shi Q (2020) Multitask deep learning with spectral knowledge for hyperspectral image classification. IEEE Geosci Remote Sens Lett 17(12):2110–2114. https://doi.org/10.1109/LGRS.2019.2962768
Frisoni GB, Jack CR Jr, Bocchetta M, Bauer C, Frederiksen KS, Liu Y, Preboske G, Swihart T, Blair M, Cavedo E, Grothe MJ (2015) The EADC-ADNI harmonized protocol for manual hippocampal segmentation on magnetic resonance: evidence of validity. Alzheimers Dement 11(2):111–125. https://doi.org/10.1016/j.jalz.2014.05.1756
Tobon-Gomez C, Geers AJ, Peters J, Weese J, Pinto K, Karim R, Ammar M, Daoudi A, Margeta J, Sandoval Z, Stender B (2015) Benchmark for algorithms segmenting the left atrium from 3D CT and MRI datasets. IEEE Trans Med Imaging 34(7):1460–1473. https://doi.org/10.1109/TMI.2015.2398818
Simpson AL, Leal JN, Pugalenthi A, Allen PJ, DeMatteo RP, Fong Y, Gönen M, Jarnagin WR, Kingham TP, Miga MI, Shia J (2015) Chemotherapy-induced splenic volume increase is independently associated with major complications after hepatic resection for metastatic colorectal cancer. J Am Coll Surg 220(3):271–280. https://doi.org/10.1016/j.jamcollsurg.2014.12.008
Litjens G, Debats O, Ven W V D, Karssemeijer N, Huisman H. (2012) A pattern recognition approach to zonal segmentation of the prostate on MRI. In: International Conference on Medical Image Computing and Computer-Assisted Intervention 413–420. Springer, Berlin, Heidelberg. doi:https://doi.org/10.1007/978-3-642-33418-4_51
Papageorghiou AT, Ohuma EO, Altman DG, Todros T, Ismail LC, Lambert A, Jaffer YA, Bertino E, Gravett MG, Purwar M, Noble JA (2014) International standards for fetal growth based on serial ultrasound measurements: the Fetal Growth Longitudinal Study of the INTERGROWTH-21st Project. Lancet 384(9946):869–879. https://doi.org/10.1016/S0140-6736(14)61490-2
Fernandes FE, Yen GG (2020) Automatic searching and pruning of deep neural networks for medical imaging diagnostic. IEEE Trans Neural Netw Learn Syst 32(12):5664–5674. https://doi.org/10.1109/TNNLS.2020.3027308
Gutman D, Codella N C, Celebi E, Helba B, Marchetti M, Mishra N, Halpern A. (2016) Skin lesion analysis toward melanoma detection: A challenge at the international symposium on biomedical imaging (ISBI) 2016, hosted by the international skin imaging collaboration (ISIC). arXiv preprint arXiv:1605.01397. doi:https://doi.org/10.48550/arXiv.1605.01397
Pacheco AG, Lima GR, Salomao AS, Krohling B, Biral IP, de Angelo GG, Alves FC Jr, Esgario JG, Simora AC, Castro PB, Rodrigues FB (2020) PAD-UFES-20: A skin lesion dataset composed of patient data and clinical images collected from smartphones. Data Brief 32:106221. https://doi.org/10.1016/j.dib.2020.106221
Giotis I, Molders N, Land S, Biehl M, Jonkman MF, Petkov N (2015) MED-NODE: a computer-assisted melanoma diagnosis system using non-dermoscopic images. Expert Syst Appl 42(19):6578–6585. https://doi.org/10.1016/j.eswa.2015.04.034
Mendonça T, Ferreira PM, Marques JS, Marcal AR, Rozeira J (2013) PH 2-A dermoscopic image database for research and benchmarking. In: 2013 35th annual international conference of the IEEE engineering in medicine and biology society (EMBC) 5437-5440. IEEE. doi:https://doi.org/10.1109/EMBC.2013.6610779
Stojanović M, Apostolović M, Stojanović D, Milošević Z, Toplaović A, Mitić-Lakušić V, Golubović M (2014) Understanding sensitivity, specificity and predictive values. Vojnosanit Pregl 71(11):1062–1065. https://doi.org/10.2298/vsp1411062s
Adegun AA, Viriri S (2020) FCN-based DenseNet framework for automated detection and classification of skin lesions in dermoscopy images. IEEE Access 8:150377–150396. https://doi.org/10.1109/ACCESS.2020.3016651
Retrieved August 22, 2023: https://challenge2020.isic-archive.com/
Retrieved August 20, 2023: https://encord.com/blog/f1-score-in-machine-learning/
Mehr R, Ameri A (2022) Skin cancer detection based on deep learning. J Biomed Phys Eng 12(6):559. https://doi.org/10.31661/jbpe.v0i0.2207-1517
Waheed Z, Waheed A, Zafar M & Riaz F (2017) An efficient machine learning approach for the detection of melanoma using dermoscopic images. In: 2017 International conference on communication, computing and digital systems (C-CODE) (pp 316–319). IEEE. doi:https://doi.org/10.1109/C-CODE.2017.7918949
Rodrigues D, Ivo R, Satapathy S, Wang S, Hemanth J, Reboucas FP (2020) A new approach for classification skin lesion based on transfer learning, deep learning, and IoT system. Pattern Recogn Lett 136:8–15. https://doi.org/10.1016/j.patrec.2020.05.019
Astorino A, Fuduli A, Veltri P, Vocaturo E (2020) Melanoma detection by means of multiple instance learning. Interdiscip Sci: Comput Life Sci 12:24–31. https://doi.org/10.1007/s12539-019-00341-y
Mukherjee S, Adhikari A, Roy M (2020) Malignant melanoma detection using multi layer preceptron with visually imperceptible features and PCA components from MED-NODE dataset. Int J Med Eng Inform 12(2):151–168. https://doi.org/10.1504/IJMEI.2020.106899
Xu D, Tian Y (2015) A comprehensive survey of clustering algorithms. Ann Data Sci 2:165–193. https://doi.org/10.1007/s40745-015-0040-1
Funding
Open access funding provided by The Science, Technology & Innovation Funding Authority (STDF) in cooperation with The Egyptian Knowledge Bank (EKB).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Medhat, S., Abdel-Galil, H., Aboutabl, A.E. et al. Iterative magnitude pruning-based light-version of AlexNet for skin cancer classification. Neural Comput & Applic 36, 1413–1428 (2024). https://doi.org/10.1007/s00521-023-09111-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00521-023-09111-w