Abstract
Clustering is one of the most fundamental unsupervised learning tasks with numerous applications in various fields. Clustering methods based on neural networks, called deep clustering methods, leverage the representational power of neural networks to enhance clustering performance. ClusterGan constitutes a generative deep clustering method that exploits generative adversarial networks (GANs) to perform clustering. However, it inherits some deficiencies of GANs, such as mode collapse, vanishing gradients and training instability. In order to tackle those deficiencies, the generative approach of implicit maximum likelihood estimation (IMLE) has been recently proposed. In this paper, we present a clustering method based on generative neural networks, called neural implicit maximum likelihood clustering, which adopts ideas from both ClusterGAN and IMLE. The proposed method has been compared with ClusterGAN and other neural clustering methods on both synthetic and real datasets, demonstrating promising results.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Unsupervised learning has gained wide interest due to the emergence of big data collections and the cost of label acquisition. Clustering is one of the most important and fundamental unsupervised learning tasks with numerous applications in computer science and many other scientific fields [1, 2]. It is defined as the process of partitioning a set of objects into groups (called clusters) so that the data in the same group share common characteristics and differ from data in other groups. While the clustering definition is simple, it is a hard machine learning problem. Its difficulty arises from several factors, e.g., data prepossessing and representation, clustering criterion, optimization algorithm, and parameter initialization. In addition, clustering evaluation is also challenging due to the unsupervised nature of the problem [3, 4].
Due to its particular importance, clustering is a well-studied problem with numerous proposed approaches. Generally, they can be classified as hierarchical (divisive or agglomerative), model-based (e.g., k-means [5], mixture models [3]) and density-based (e.g., DBSCAN [6], DensityPeaks [7]). Most methods are effective when the data space is low dimensional and not complex. Various feature extraction and feature transformation methods have been proposed to map the original complex data to a more “cluster-friendly” feature space as a prepossessing step to address those limitations. Some of the methods include Principal Component Analysis [8], Non-negative Matrix Factorization [9], Spectral methods [10], and Minimum Density Hyperplanes [11].
Neural networks have been employed for clustering in the context of deep learning. Deep neural networks have been used to learn rich and useful data representations from data collections without heavily relying on human-engineered features. They can improve the performance of supervised and unsupervised learning tasks because of their excellent nonlinear mapping capability and flexibility [12,13,14,15,16]. Although clustering has not initially been the primary goal of deep learning, several clustering methods have been proposed that exploit the representational power of neural networks; thus, the deep clustering category of methods has emerged [17,18,19,20,21]. Such methods aim to improve the quality of clustering results by appropriately training neural networks to transform the input data and generate cluster-friendly representations [22,23,24,25,26].
In this work, we propose a neural clustering method called Neural Implicit Maximum Likelihood Clustering (NIMLC). It is a generative clustering method that relies on the recently proposed method of Implicit Maximum Likelihood Estimation (IMLE) [27]. This is an alternative approach to GANs [28]: given a set of data objects, the IMLE method uses a generator network that takes random input vectors and learns to produce synthetic samples. By minimizing an appropriate objective, the network is trained so that the distribution of samples resembles the data distribution. It has been shown that this training procedure maximizes the likelihood of the dataset without explicitly computing the likelihood.
In analogy with the ClusterGan [29] method which exploits the GAN methodology to perform clustering, we have developed the NIMLC method, which relies on the IMLE methodology to perform clustering. NIMLC utilizes two neural networks, the generator and the encoder. In contrast to ClusterGAN, the discriminator network is not needed. The generator network is fed by appropriately selected random samples (latent vectors) z belonging to K clusters and is trained to produce synthetic samples that resemble the objects of dataset X. The encoder network provides the partition of the dataset X into K clusters by learning the inverse map from the data space X to the latent space Z. Training of both networks is achieved by minimizing an appropriately defined objective function that involves the IMLE loss (data generation) and the reconstruction loss for the latent vectors z.
Note that the IMLE method does not suffer from mode collapse, vanishing gradients or training instability that are frequently encountered in GAN training. Moreover, it does not require large datasets for training. Our aim is to exploit those nice IMLE properties for solving clustering problems through the development of the proposed NIMLC method.
The organization of the paper is the following. In Sect. 2, related work is presented. In Sect. 3, the IMLE method is first described and then the proposed NIMLC clustering method is presented and explained. Section 4 presents comparative experimental results on various datasets, while Sect. 5 provides conclusions and directions for future research.
2 Related work
Neural network methods can be distinguished into two main categories when used for clustering [23]. The first and broader category is based on autoencoders [30, 31] and aim to transform the data into “cluster-friendly” space latent space representations [17, 32,33,34,35,36,37,38,39,40,41,42,43]. The second category relies on generative neural networks, like GANs [44,45,46,47,48] and Variational Autoencoders [49,50,51].
2.1 Autoencoder based clustering
Inspired by the t-SNE [52] algorithm, the Deep Embedding Clustering (DEC) [53] method has been proposed that optimizes both the reconstruction objective and a clustering objective. DEC transforms the data in the embedded space using an autoencoder and then optimizes a clustering loss defined by the KL divergence between two distributions P and Q: Q is the soft clustering assignment of the data based on the distances in the embedded space between data points and cluster centers, and P is an adjusted target distribution aiming to enhance the clustering quality by leveraging the soft cluster assignments. The initial cluster centers are computed by the k-means algorithm. The Improved Deep Clustering with local structure preservation (IDEC) [54] has also been proposed to improve data representation by maintaining their local structure. The optimized objective function is
where f and g are the encoder and decoder, respectively (with learnable parameters), n is the number of data points and K the number of clusters, while \(\lambda \ge 0\) is the regularization parameter that balances the reconstruction loss and the clustering error.
Similar to DEC, the Deep Clustering Network (DCN) [55] jointly learns the embeddings and the cluster assignments by optimizing the k-means clustering loss on the embedded space (Eq. 2). The optimized objective function is:
where M is a matrix that contains the k cluster centers in the embedded space, and \(s_i\) is the cluster assignment vector for data point \(x_i\) which has only one nonzero element.
2.2 Generative neural clustering
The second category of neural clustering methods includes techniques that are based on models for synthetic data generation and are typically based on the GAN [28] methodology. ClusterGan [29] is the most well-known method of this category that utilizes the GANs methodology to achieve data clustering and synthetic data generation. ClusterGan takes as input a set of random input vectors z that belong to K clusters. An input vector of cluster k is defined as \(z = (z_{n}, z_{c})\) where \(z_{n} \sim {\mathcal {N}}(0, \sigma ^{2} I_{d_n})\) and \(z_{c}=e_k\) where \(e_k\) is kth standard unit vector of length K. Besides the generator \({\mathcal {G}}\) and the discriminator network \({\mathcal {D}}\), ClusterGAN includes an additional network, the encoder \({\mathcal {E}}\) that provides the cluster assignments of its input x. ClusterGan trains the typical generator-discriminator architecture jointly with the encoder to achieve clustering and synthetic data generation by optimizing the objective function 3:
where \({\mathcal {H}}(.,.)\) is the cross-entropy loss, \(\beta _{n}\) and \(\beta _{c}\) the regularization coefficients q(.) the quality function, given as \(q(x) = log(x)\) for vanilla GAN [28], and \(q(x) = x\) for Wasserstein GAN (WGAN) [56].
3 Neural implicit maximum likelihood clustering
The proposed NIMLC method relies on the data generation capabilities of the IMLE algorithm, which is summarized next.
3.1 Implicit maximum likelihood estimation
Given a dataset \(X=\{x_1,\ldots , x_n\}\) of d-dimensional vectors, the IMLE algorithm [27] trains a generative neural network \({\mathcal {G}}_{\theta }\) with m inputs, d outputs and parameter vector (weights) \(\theta\). This generator takes as input a random vector \(z \in {\mathbb {R}}^{m}\) usually sampled from an m-dimensional Normal distribution and produces a sample \(s^{\theta } \in {\mathbb {R}}^{d}\), i.e., \(s^{\theta }={\mathcal {G}}_{\theta }(z)\) (see Fig. 2a). IMLE trains the generator to generate synthetic samples \(s^{\theta }\) that resemble the real data \(x_i\). It is a simple generative method that, under certain conditions, implicitly maximizes the likelihood of the dataset, although the IMLE objective does not explicitly contain any log-likelihood term, and training neural networks using maximum likelihood is considered a difficult task [57].
In each IMLE iteration, a sampling procedure takes place where a set of L random input vectors \(z_i\) (called latent vectors) are drawn from the Normal distribution \(z_{i} \sim {\mathcal {N}}(0, \sigma ^{2}I_m)\) and used for the computation of the corresponding synthetic samples \(s_{i}^{\theta }={\mathcal {G}}_{\theta }(z_i)\) (\(i=1, \ldots , L\)). Then, for each real data example \(x_{i}\) \((i=1, \ldots , N)\), its representative sample \(r_{i}^{\theta } \in S^{\theta }\) is determined through nearest neighbor search (NNS) in \(S^{\theta }\) based on Euclidean distance, i.e., \(r_{i}^{\theta } = NNS(x_{i}, S^{\theta })\). The generator parameters \(\theta\) are updated in order to minimize the following IMLE objective function:
Figure 1 provides an illustration of the IMLE behavior.
The IMLE method exhibits several nice properties: it does not suffer from mode collapse, vanishing gradients, or training instability, unlike popular deep generative methods such as, for example, GANs [28]. Mode collapses do not occur since the loss ensures that each data example is represented by at least one sample. Gradients do not vanish because the gradient of the distance between a data example and its representative sample does not become zero unless they coincide. Training is stable because the IMLE estimator is the solution to a simple minimization problem. Finally, it can be used both in the case of small and large datasets.
3.2 Cluster friendly input distribution
In the original IMLE method, the random input (latent) vectors z belong to a single cluster since they are drawn from a multivariate m-dimensional Normal distribution. This is not convenient for clustering. If we assume that the input vectors z are drawn from a mixture model, i.e., from K distinct distributions, then a clustering of the original dataset X could be obtained: each data point \(x_i\) can be assigned to the cluster to which its corresponding input vector \(z_i\) belongs to. Therefore in the proposed method, the single Normal distribution is replaced by K non-overlapping distributions, with the kth distribution responsible for the generation of the subset \(Z_k\) of input vectors assigned to cluster k. The most obvious first choice is a mixture of K m-dimensional Gaussian distributions. However, this choice requires the specification of the means and covariances of K Gaussian distributions so that they are well separated.
A more sophisticated mechanism for generating m-dimensional random vectors that belong to K disjoint clusters has been proposed in ClusterGan [29], where input vector z consists of two parts, i.e., \(z=(z_n, z_c)\). The first part \(z_n\) is random vector (of dimension \(d_n\)) drawn from the Gaussian distribution: \(z_{n} \sim {\mathcal {N}}(0, \sigma ^{2} I_{d_n})\). The second part \(z_c\), is deterministic and specifies the cluster k to which z is assigned. Specifically, \(z_c\) is the one-hot encoding of the corresponding cluster k. Thus, for K clusters, the dimension of \(z_c\) is equal to K and, if z belongs to the kth cluster, then \(z_c=e_k\) where \(e_k\) is the kth standard unit vector. Note that \(\sigma\) should be set to a small value so that clusters do not overlap.
In summary, in order to generate an input vector \(z=(z_c,z_n)\) belonging cluster k, we set the \(z_c\) part equal to the one-hot encoding of k and draw the \(z_n\) part from \({\mathcal {N}}(0, \sigma ^{2} I_{d_n})\). By sampling an equal number of vectors for each cluster k, the set of random input vectors Z is created at each iteration which is partitioned into disjoint subsets \(Z_k\), each one containing the random input vectors for cluster k (\(k=1,\ldots ,K\)).
Additionally, since \(s^{\theta } = {\mathcal {G}}_{\theta }(z)\), the set \(S^{\theta }\) of computed samples is partitioned into K disjoint clusters \(S_{k}^{\theta }\). Consequently, the original dataset X can be partitioned into K clusters by assigning each \(x_i\) to the cluster of its representative \(r_{i}^{\theta }\), i.e., if \(r_{i}^{\theta } \in S_{k}^{\theta }\) then \(x_i\) is assigned to cluster k.
3.3 The IMLE loss from a clustering perspective
If we examine the IMLE objective function, we can observe its similarities with the k-means clustering loss. Specifically, if we generate exactly K samples \(S^{\theta }_K=\{s_1^{\theta }, \ldots , s_K^{\theta }\}\) in each training epoch, where K is the number of clusters, we can treat those synthetic samples as cluster representatives (centroids). In this case, the IMLE objective coincides with the k-means objective (\(\mathbbm {1}_{C_k}\) is the indicator function):
and IMLE can be considered as a clustering procedure that trains the generator to produce the cluster centers. The major difference between k-means and IMLE is that the k-means updates the centroids directly in order to minimize the clustering loss; on the contrary, the IMLE method updates the parameters \(\theta\) of the generator.
An issue to be considered is how to specify the k input vectors \(z_k\) that will be used to generate the K samples so that each sample represents a different cluster. Since \(z_k=(z_{nk}, z_{ck})\), a straightforward solution is to set \(\sigma = 0\), thus \(z_{nk}=0\) for all k and \(z_{ck}=e_k\) for all \(k=1,\ldots ,K\). Then by feeding those \(z_k\) vectors as inputs to the generator, the synthetic samples \(s_k\) are provided as outputs which can be treated as cluster representatives. Training the generator this way using IMLE, we observed clustering behavior similar to k-means and that the generated K samples resembled the average data point of each cluster.
3.4 The NIMLC architecture
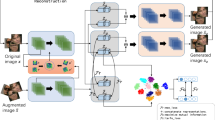
The proposed NIMLC approach is a modification of the IMLE method in order to achieve not only synthetic data generation but also clustering of the original dataset X. NIMLC combines ideas from IMLE and ClusterGAN. More specifically, it exploits the IMLE generator network that is fed with clustered input vectors z that follow the \((z_n, z_c)\) representation proposed in ClusterGAN. Additionally, it employs a second network called encoder (originally proposed in ClusterGAN) that is trained to provide the cluster assignment for a data point x. It should be noted that, unlike ClusterGAN, NIMLC does not make use of a discriminator network since it is based on IMLE for synthetic data generation. The NIMLC architecture is presented in Fig. 2b.
The generator \({\mathcal {G}}\) is trained to produce synthetic samples that resemble the real data \(x_i\) by minimizing the IMLE objective (Eq. 4). It provides a mapping from the latent space to the data space. The encoder \({\mathcal {E}}\) is trained jointly with the generator to implement the inverse mapping from the data space to the latent space. Thus for an input x, it provides estimates of \({\hat{z}}_n\) and \({\hat{z}}_c\). The latter (\({\hat{z}}_c\)) is computed using the softmax activation function (with K outputs) and provides a soft clustering assignment of the input x into K clusters.
In summary, the NIMLC architecture feeds an input vector \(z=(z_n, z_c)\) to the generator, which produces a synthetic sample \(s={\mathcal {G}}(z)\). This sample is subsequently fed to the encoder, which provides the output \({\hat{z}}={\mathcal {E}}(s)\). Note that the NIMLC network is actually an autoencoder since it takes an input z and provides as output an estimate \({\hat{z}}\) of z. After training, the encoder implements a clustering model providing soft clustering assignments \({\hat{z}}_c\) for any data point x.
3.5 The NIMLC objective function
The objective function used to train the NIMLC architecture consists of two parts. The first part concerns the generative process and is the IMLE error equal to \(\sum \nolimits _{i=1}^n ||r^{\theta _{\mathcal {G}}}_i - x_i||^2\) (Eq. 4). Since NIMLC is an autoencoder, the second part of the objective function is the reconstruction loss of the autoencoder. This loss can be split into two terms. The first term is the reconstruction loss for the \(z_n\) part: \(\sum \nolimits _{i=1}^n ||z_{ni} - {\hat{z}}_{ni}||^2\). The second term is the reconstruction loss for the \(z_c\) part. Since \(z_c\) has the form of one-hot vector and \({\hat{z}}_c\) are probability vectors provided by the softmax function, the cross-entropy \({\mathcal {H}}(z_{c}, {\hat{z}}_{c})\) between \(z_c\) and \({\hat{z}}_c\) is used as a loss function.
The complete objective function is presented below, where \(\beta _n\) and \(\beta _c\) are hyperparameters adjusting the importance of each term.
It should be noted that the first term depends only on the parameters \(\theta _{\mathcal {G}}\) of the generator, while the rest two terms depend on the parameters of both the generator \(\theta _{\mathcal {G}}\) and the encoder \(\theta _{\mathcal {E}}\).
3.6 Slow paced learning
A critical hyperparameter of the NIMLC method is the standard deviation \(\sigma\) of the noise distribution used to generate the random part \(z_n\) of the input vectors. As mentioned earlier, when training the model with \(\sigma = 0\), we have a very strict case with one generated sample per cluster. This sample can be considered as the representative of the corresponding cluster, and the obtained clustering results are on par with those of k-means. On the other hand, if \(\sigma\) is relatively large (e.g., \(\sigma = 0.15\)), the random input vectors per cluster are not very close. Therefore it is possible for the generator to map the inputs of the same cluster to different regions in the data space, which negatively affects clustering performance. Moreover, we have observed that it is difficult to specify an appropriate value for \(\sigma\).
In order to tackle this problem we propose the following procedure:
-
Start training with a small value of sigma, preferably \(\sigma = 0\).
-
In each training epoch increase \(\sigma\) by a small amount \(\Delta \sigma\).
-
Stop increasing when a max value \(\sigma _{max}\) is reached.
The intuition is that this slow-paced training procedure ([58, 59]) strives to learn and cluster the “easier” data points first, like those that are close to the cluster centers and then tries to learn and cluster “more difficult” data points away from the cluster centers. Thus, we initially start to explore the clustering solution space with no variability in the input space (\(\sigma = 0\)). This way, we enforce only K samples to be generated and used to train the model. Then, at each training epoch, we add variability to the inputs by slowly increasing \(\sigma\) in order to incrementally capture complicated structures in the dataset.
Figure 3 provides an illustration of the generated samples for the Moons synthetic dataset as training proceeds and \(\sigma\) gradually increases. It is clear that the model progressively succeeds in learning more complex data structures, generating high-quality samples and providing the correct clustering solution.
3.7 The NIMLC algorithm
The NIMLC method is summarized in Algorithm 1. At each epoch, a set of input vectors \(Z=\{z_1, \ldots , z_L\}\) is generated, belonging to K clusters \(Z_k\), \(k=1,\ldots ,K\) of equal size. Each input vector \(z_i = (z_{ni}, z_{ci})\) of \(Z_k\) is computed by sampling \(z_{ni}\) from \({\mathcal {N}}(0, \sigma ^{2} I_{d_{n}})\) and setting \(z_{ci}=e_k\). We then feed the generator with the set of input vectors Z and the set of synthetic samples \(S^{\theta _{\mathcal {G}}} = \{{s}_1^{\theta _{\mathcal {G}}}, \ldots , {s}_{L}^{\theta _{\mathcal {G}}}\}\) are generated at its output, i.e., \(s_i^{\theta _{\mathcal {G}}} = {\mathcal {G}}(z_i)\). Then for each data batch \(X_b\), we compute the nearest synthetic sample \(r_i\) for each \(x_i \in X_b\), ie. \(r_i^{\theta _{\mathcal {G}}} = NNS(x_i, S^{\theta _{\mathcal {G}}})\). Next, each \(r_i\) is fed as input to the encoder that produces the reconstruction \(\hat{z_i} = {\mathcal {E}}(r_i^{\theta _{\mathcal {G}}})\), where \(\hat{z_i} = ({\hat{z}}_{ni}, {\hat{z}}_{ci})\). Then we update the parameters of the generator and the encoder using the gradients of the objective function (algorithm 1 steps 11 and 12). Finally, before proceeding to the next epoch, the standard deviation \(\sigma\) is updated.

It should be noted that using IMLE for clustering has been introduced in our previous work [60]. However, that method did not make use of the encoder network. Instead, a two-stage nearest neighbor search was used to perform cluster assignments. Specifically, in the first stage, the centroid \(c_k\) of each subset \(S^\theta _k\) was computed, and then the \(x_i\) was assigned to the cluster l whose centroid \(c_l\) is nearest to \(x_i\) based on Euclidean distance. Additionally, in the second stage, instead of determining the representative sample for \(x_i\) through the nearest neighbor search over the entire set of samples \(S^\theta\), the nearest neighbor search was executed only to the specific subset \(S^\theta _l\) that contains the samples of cluster l. The NIMLC method proposed herein includes two substantial improvements that lead to considerable performance enhancement. The first is the use of the encoder network that directly provides the cluster assignment for a given input x, while the second is the gradual increase in the noise variance \(\sigma\) that allows for slow-paced learning. Additionally, we exploit the generalization capability of the encoder network to cluster those data points that the generator could not learn sufficiently.
A computational overhead of our approach compared to deep clustering methods is related to nearest neighbor search. The training process involves several epochs where the algorithm must find closest synthetic sample to each original data point, resulting in an \({\mathcal {O}}(NL)\) overhead in distance calculations. However, we observed that recalculating the nearest neighbors in every training epoch is unnecessary; reusing them for 5 to 10 epochs can significantly reduce the training time without compromising clustering performance.
4 Experiments
In order to evaluate the proposed clustering method (NIMLC), we conducted an experimental study using several synthetic and real datasets. We have compared NIMLC against ClusterGan [29] and the two most popular deep clustering methods, namely DCN [55] and DEC [53]. We also provide results using k-means [5, 61] and the density-based method of i-DivClu-D [62].
4.1 Synthetic datasets
We have used three synthetic two-dimensional datasets (Table 1) with known ground truth and different structures in order to assess the clustering capability of our method. The Gaussians dataset consists of four clusters (Fig. 4a), while the Moons (Fig. 4b) and the Rings (Fig. 4c) consist of two clusters. The Gaussians dataset is easier to cluster compared to the other two datasets, whose structure is more complex. It should be emphasized that it is difficult for a parametric method to be able to cluster both cloud-shaped (Gaussians) and ring-shaped (Rings) datasets.
4.2 Real datasets
We further evaluated the method by including real datasets in our experimental study. For all datasets the number of clusters was set equal to the number of classes. As a pre-processing step, we used min-max normalization to map the attributes of each dataset to the [0, 1] interval in order to prevent attributes with large ranges from dominating the distance calculation and avoid numerical instabilities in the computation [63]. The descriptions of the datasets that we included in our study are given below. For a summary, refer to Table 2.
-
10x_73k [64] dataset consists of 73,233 RNA-transcripts belonging to 8 different cell types. The dataset is sparse since the data matrix has about 40% zero values. Hence, we selected the 720 genes with the highest variances across the cells to reduce the data dimensionality similar to [29].
-
Australian [65] two-class dataset is composed of 690 credit card applications. A 14-dimensional feature vector describes each sample.
-
CMU [65] contains grayscale facial images of twenty individuals captured with varying poses, expressions, and the presence or absence of glasses. The images are available in several resolutions, but for the purpose of our study, we have utilized the \(128 \times 120\) resolution images.
-
Dermatology [65] is a six-class dataset containing 366 patient records that suffer from six different types of Eryhemato-Squamous disease. Each patient is described by a 34-dimensional vector containing clinical and histopathological features.
-
E. coli [65] includes 336 proteins from the E. coli bacterium, and seven attributes, calculated from the amino acid sequences, are provided. Proteins belong to eight classes according to their cellular localization sites.
-
Iris [65] dataset contains three classes of 50 instances each, where each class refers to a type of iris plant. Each sample is described by a 4-dimensional vector, corresponding to the length and width of the sepals and petals in centimeters.
-
Olivetti [66] is a face database of 40 individuals with ten 64\(\times\)64 grayscale images per individual. For some individuals, the images were taken at different times, varying the lighting, facial expressions (open/closed eyes, smiling/not smiling), and facial details (glasses/no glasses). All the images were taken against a dark homogeneous background with the subjects in an upright, frontal position (with tolerance for some side movement).
-
Optical Recognition of Handwritten Digits [65] dataset (ORHD) comprises a set of handwritten digits, with ten classes corresponding to each digit from 0 to 9. The resolution of each image is 8x8. For our experiment, we utilized the test set of this dataset, which consists of 1797 images.
-
Pendigits [65] dataset consists of 250 writing samples from 44 different writers, a total of 10,992 written samples. Each sample is a 16-dimensional vector containing pixel coordinates associated with a label from ten classes.
-
United States Postal Service [67] dataset (USPS) is a collection of hand-written digits consisting of 7291 grayscale images. The dataset is organized into ten classes, each representing a digit from 0 to 9. Each digit is represented by a set of images, each of size \(16\times 16\) pixels.
-
Wine [65] three-class dataset consists of 178 samples of chemical analysis of wines. A 13-dimensional feature vector describes each sample.
4.3 Evaluation measures
It is important to mention that since clustering is an unsupervised problem, we ensured that all algorithms were unaware of the true clustering of the data. In order to evaluate the results of the clustering methods, we use standard external evaluation measures [68], which assume that ground truth clustering is available. For all algorithms, the number of clusters is set to the number of ground-truth categories [25] and assumes ground truth that cluster labels coincide with class labels. The first evaluation measure is clustering accuracy (ACC):
where \({\textbf {1}}(x)\) is the indicator function, \(y_{i}\) is the ground-truth label, \(c_{i}\) is the cluster assignment generated by the clustering algorithm, and m is a mapping function which ranges over all possible one-to-one mappings between assignments and labels. This measure finds the best matching between cluster assignments from a clustering method and the ground truth. It is worth noting that the optimal mapping function can be efficiently computed by the Hungarian algorithm [69]. The second evaluation measure is purity (PUR). The same equation formulates purity as clustering accuracy (Eq. 7), but their key difference is in the mapping function m. In this case, the mapping function of m greedily assigns clustering labels to ground truth categories in each cluster in order to maximize purity. The third evaluation measure is the normalized mutual information (NMI) defined as [70]:
where Y denotes the ground-truth labels, C denotes the clusters labels, I is the mutual information measure and H the entropy. The final evaluation metric is the adjusted Rand Index (ARI) [71, 72], which computes a similarity measure between two clustering solutions defined as the proportion of object pairs that are either assigned to the same cluster in both clusterings or to different clusters in both clusterings.
4.4 Implementation details
Both the generator and the encoder were trained using the Adam optimizer [73] with learning rate \(n = 3\times 10^{-4}\) and coefficients \(b_{1} = 0.5\) and \(b_{2} = 0.9\). We set \(b_n = b_c = 1\), and \(\Delta \sigma = 5 \times 10^{-5}\) in all experiments. Additionally, the number of samples was set equal to 100 and 200 for small and big datasets, respectively. We used the same architectures for the two networks as the ClusterGan [29]. Specifically, the dimension of \(z_{c}\) is the set equal to the number of clusters. We used Leaky Relu activations (LRelu) with leak = 0.2 and Batch Normalization (BN). We used the same number of hidden layers and hidden neurons for all datasets. We present the detailed generator and encoder architectures in Tables 3 and 4, respectively.
For ClusterGan, we used the proposed architecture and hyperparameters. In the case of the DCN and DEC, an extensive search for an autoencoder model was required in order to obtain good results. We chose symmetrical encoder and decoder networks to simplify the architecture search problem. We resorted to an encoder architecture with three layers: \(d-[2d,3d]-d_{z}\), where d is the data space dimension and \(d_{z}\) is the latent space dimension. All layers are fully connected.
NIMLC and ClusterGan methods were executed for 5000 epochs, while DCN and DEC required 300 to 500 epochs of pretraining and 100 epochs of training with the clustering objective. Furthermore, for the methods that depend on initialization, we executed the neural approaches of NIMLC, ClusterGan, DCN, and DEC three times and the k-means algorithm ten times with k-means\(\texttt {++}\) [74] initialization. Average performance results are provided. The i-DivClu-D method is deterministic and requires the number of nearest neighbors as a hyperparameter. In our experiments we set its value at the minimum number that resulted in a connected graph.
4.5 Results on synthetic datasets
In Table 5, we provide the average clustering performance of the compared methods for the synthetic datasets. All methods performed well when the dataset consisted of spherical, well-separated data clusters, as happens in the Gaussians dataset. In the Moons and Rings datasets, ClusterGan and DEC had a similar clustering performance as the k-means algorithm, while the DCN method performed better. On the other hand, the NIMLC method could perfectly solve the Moons dataset and had by a significant margin the best clustering performance on Rings, which is the most difficult of the three synthetic datasets. It should be stressed that the NIMLC method presents the unique capability of solving both the Gaussian and the Rings datasets by training the same neural architecture. It should be noted that the density-based i-DivClu-D method demonstrated perfect clustering performance in all three synthetic datasets.
4.6 Results on real datasets
Table 6 shows the clustering performance of the compared methods on tabular datasets, while Table 7 displays performance results on image datasets.
The NIMLC method achieved excellent clustering performance on 10x_73k, Australian, CMU, Olivetti-Faces, and Pendigits, outperforming all other methods. Moreover, on Dermatology, E. coli, Iris, ORHD, USPS, and Wine datasets, the NIMLC method demonstrated comparable results with the best-performing method. It should be stressed that the high dimensionality of data and the limited number of training samples resulted in training failures for ClusterGan, DCN, and DEC on CMU and Olivetti-Faces datasets. In contrast, the NIMLC method was able to learn these datasets effectively despite the small number of samples. Compared to k-means and the density-based clustering method (i-DivClu-D) our method also provides better or comparative results in all cases with the superiority being more clear on 10x_73k, Pendigits, Oliveti-Faces and USPS datasets. In summary, experimental results indicate that the proposed method constitutes a neural-based clustering method that performs well both for low dimensional and high dimensional data without requiring large number of samples as happens with typical deep clustering methods. Therefore it constitutes a viable alternative for both conventional and deep clustering approaches.
5 Conclusions
We have proposed the NIMLC clustering method that is based on neural network training. NIMLC is a generative clustering approach that relies on the IMLE generative methodology to perform clustering. The NIMLC brings ideas from the ClusterGAN algorithm into the IMLE framework to overcome some of the GAN deficiencies. The method is based on a simple training objective, does not suffer from training instabilities, and performs well on small datasets. Experimental comparison against several deep clustering methods and conventional clustering mathods illustrates the potential of the approach. A notable characteristic of the method is that, as shown in the experiments with synthetic data, it is able to cluster both cloud-shaped and ring-shaped data using the same hyperparameter setting.
Future research could focus on a more detailed experimental investigation of the performance of NIMLC and its use in real-world applications. Additionally, we aim to consider a modification of the method where the number of samples could vary at each epoch. In the same spirit, we could explore the possibility of adopting a self-paced approach similar to the one proposed in [59] for the online tuning of the parameter \(\Delta \sigma\). Finally, it is interesting to study how the NIMLC approach could be integrated into a general methodology for estimating the true number of clusters in the dataset.
Data availability
\(\bullet\) The UCI datasets are available at the official UCI page: archive.ics.uci.edu. \(\bullet\) The 10x_73k dataset is available at the following github repository: github.com/eugenelin1/DRA. \(\bullet\) The Olivetti-Faces and ORHD datasets are available at the official page of scikit-learn: − https://scikit-learn.org/stable/modules/generated/sklearn.datasets.fetch_olivetti_faces.html − https://scikit-learn.org/stable/modules/generated/sklearn.datasets.load_digits.html\(\bullet\) The USPS dataset is available at the following page: https://www.csie.ntu.edu.tw/ cjlin/libsvmtools/datasets/multiclass.html#usps
References
Filippone M, Camastra F, Masulli F, Rovetta S (2008) A survey of kernel and spectral methods for clustering. Pattern Recognit 41(1):176–190
Jain AK (2010) Data clustering: 50 years beyond k-means. Pattern Recognit Lett 31(8):651–666
Bishop CM (2006) Pattern recognition. Mach Learn 128(9):66
Jain AK, Murty MN, Flynn PJ (1999) Data clustering: a review. ACM Comput Surv 31(3):264–323
MacQueen J (1967) Some methods for classification and analysis of multivariate observations. In: Proceedings of the fifth Berkeley symposium on mathematical statistics and probability, vol 1. Oakland, CA, USA, pp 281–297
Ester M, Kriegel H-P, Sander J, Xu X (1996) A density-based algorithm for discovering clusters in large spatial databases with noise. In: Kdd, vol 96, pp 226–231
Rodriguez A, Laio A (2014) Clustering by fast search and find of density peaks. Science 344(6191):1492–1496
Wold S, Esbensen K, Geladi P (1987) Principal component analysis. Chemom Intell Lab Syst 2(1–3):37–52
Lee DD, Seung HS (1999) Learning the parts of objects by non-negative matrix factorization. Nature 401(6755):788–791
Ng AY, Jordan MI, Weiss Y (2002) On spectral clustering: analysis and an algorithm. In: Advances in neural information processing systems, pp 849–856
Pavlidis NG, Hofmeyr DP, Tasoulis SK (2016) Minimum density hyperplanes. J Mach Learn Res 6:66
LeCun Y, Bengio Y, Hinton G (2015) Deep learning. Nature 521(7553):436–444
Krizhevsky A, Sutskever I, Hinton GE (2012) Imagenet classification with deep convolutional neural networks. Adv Neural Inf Process Syst 25:1097–1105
Rifai S, Vincent P, Muller X, Glorot X, Bengio Y (2011) Contractive auto-encoders: explicit invariance during feature extraction. In: ICML
Nellas IA, Tasoulis SK, Plagianakos VP (2021) Convolutional variational autoencoders for image clustering. In: 2021 International conference on data mining workshops (ICDMW). IEEE, pp 695–702
Guo X, Liu X, Zhu E, Yin J (2017) Deep clustering with convolutional autoencoders. In: Neural information processing: 24th international conference, ICONIP 2017, Guangzhou, China, November 14–18, 2017, proceedings, Part II 24. Springer, pp 373–382
Song C, Liu F, Huang Y, Wang L, Tan T (2013) Auto-encoder based data clustering. In: Iberoamerican congress on pattern recognition. Springer, pp 117–124
McConville R, Santos-Rodriguez R, Piechocki RJ, Craddock I (2021) N2d:(not too) deep clustering via clustering the local manifold of an autoencoded embedding. In: 2020 25th International conference on pattern recognition (ICPR). IEEE, pp 5145–5152
Yang J, Parikh D, Batra D (2016) Joint unsupervised learning of deep representations and image clusters. In: Proceedings of the IEEE conference on computer vision and pattern recognition, pp 5147–5156
Caron M, Bojanowski P, Joulin A, Douze M (2018) Deep clustering for unsupervised learning of visual features. In: Proceedings of the European conference on computer vision (ECCV), pp 132–149
Ji X, Henriques JF, Vedaldi A (2019) Invariant information clustering for unsupervised image classification and segmentation. In: Proceedings of the IEEE/CVF international conference on computer vision, pp 9865–9874
Nutakki G, Abdollahi B, Sun W, Nasraoui O (2019) An introduction to deep clustering, pp 73–89. https://doi.org/10.1007/978-3-319-97864-2_4
Ren Y, Pu J, Yang Z, Xu J, Li G, Pu X, Yu PS, He L (2022) Deep clustering: a comprehensive survey. arXiv preprint arXiv:2210.04142
Aljalbout E, Golkov V, Siddiqui Y, Strobel M, Cremers D (2018) Clustering with deep learning: taxonomy and new methods. arXiv preprint arXiv:1801.07648
Min E, Guo X, Liu Q, Zhang G, Cui J, Long J (2018) A survey of clustering with deep learning: from the perspective of network architecture. IEEE Access 6:39501–39514
Nutakki GC, Abdollahi B, Sun W, Nasraoui O (2019) In: Nasraoui O, Ben N’Cir C-E (eds) An introduction to deep clustering. Springer, Cham, pp 73–89. https://doi.org/10.1007/978-3-319-97864-2_4
Li K, Malik J (2018) Implicit maximum likelihood estimation. arXiv preprint arXiv:1809.09087
Goodfellow I, Pouget-Abadie J, Mirza M, Xu B, Warde-Farley D, Ozair S, Courville A, Bengio Y (2014) Generative adversarial nets. In: Advances in neural information processing systems, pp 2672–2680
Mukherjee S, Asnani H, Lin E, Kannan S (2019) Clustergan: latent space clustering in generative adversarial networks. In: Proceedings of the AAAI conference on artificial intelligence, vol 33, pp 4610–4617
Hinton GE, Salakhutdinov RR (2006) Reducing the dimensionality of data with neural networks. Science 313(5786):504–507
Vincent P, Larochelle H, Lajoie I, Bengio Y, Manzagol P-A, Bottou L (2010) Stacked denoising autoencoders: learning useful representations in a deep network with a local denoising criterion. J Mach Learn Res 11(12):66
Huang P, Huang Y, Wang W, Wang L (2014) Deep embedding network for clustering. In: 2014 22nd International conference on pattern recognition. IEEE, pp 1532–1537
Peng X, Xiao S, Feng J, Yau W-Y, Yi Z (2016) Deep subspace clustering with sparsity prior. In: IJCAI, pp 1925–1931
Ji P, Zhang T, Li H, Salzmann M, Reid I (2017) Deep subspace clustering networks. Adv Neural Inf Process Syst 30:66
Ghasedi Dizaji K, Herandi A, Deng C, Cai W, Huang H (2017) Deep clustering via joint convolutional autoencoder embedding and relative entropy minimization. In: Proceedings of the IEEE international conference on computer vision, pp 5736–5745
Chen D, Lv J, Zhang Y (2017) Unsupervised multi-manifold clustering by learning deep representation. In: Workshops at the thirty-first AAAI conference on artificial intelligence
Li F, Qiao H, Zhang B (2018) Discriminatively boosted image clustering with fully convolutional auto-encoders. Pattern Recognit 83:161–173
Yang X, Deng C, Zheng F, Yan J, Liu W (2019) Deep spectral clustering using dual autoencoder network. In: Proceedings of the IEEE/CVF conference on computer vision and pattern recognition, pp 4066–4075
Ren Y, Wang N, Li M, Xu Z (2020) Deep density-based image clustering. Knowl Based Syst 197:105841
Affeldt S, Labiod L, Nadif M (2020) Spectral clustering via ensemble deep autoencoder learning (sc-edae). Pattern Recognit 108:107522
Guo X, Liu X, Zhu E, Zhu X, Li M, Xu X, Yin J (2019) Adaptive self-paced deep clustering with data augmentation. IEEE Trans Knowl Data Eng 32(9):1680–1693
Yang X, Deng C, Wei K, Yan J, Liu W (2020) Adversarial learning for robust deep clustering. Adv Neural Inf Process Syst 33:9098–9108
Wang J, Jiang J (2021) Unsupervised deep clustering via adaptive gmm modeling and optimization. Neurocomputing 433:199–211
Springenberg JT (2015) Unsupervised and semi-supervised learning with categorical generative adversarial networks. arXiv preprint arXiv:1511.06390
Zhou P, Hou Y, Feng J (2018) Deep adversarial subspace clustering. In: Proceedings of the IEEE conference on computer vision and pattern recognition, pp 1596–1604
Ghasedi K, Wang X, Deng C, Huang H (2019) Balanced self-paced learning for generative adversarial clustering network. In: Proceedings of the IEEE/CVF conference on computer vision and pattern recognition, pp 4391–4400
Mrabah N, Bouguessa M, Ksantini R (2020) Adversarial deep embedded clustering: on a better trade-off between feature randomness and feature drift. IEEE Trans Knowl Data Eng 6:66
Yang X, Yan J, Cheng Y, Zhang Y (2022) Learning deep generative clustering via mutual information maximization. IEEE Trans Neural Netw Learn Syst 6:66
Jiang Z, Zheng Y, Tan H, Tang B, Zhou H (2016) Variational deep embedding: an unsupervised and generative approach to clustering. arXiv preprint arXiv:1611.05148
Dilokthanakul N, Mediano PA, Garnelo M, Lee MC, Salimbeni H, Arulkumaran K, Shanahan M (2016) Deep unsupervised clustering with Gaussian mixture variational autoencoders. arXiv preprint arXiv:1611.02648
Yang L, Fan W, Bouguila N (2021) Deep clustering analysis via dual variational autoencoder with spherical latent embeddings. IEEE Trans Neural Netw Learn Syst 6:66
Van der Maaten L, Hinton G (2008) Visualizing data using t-sne. J Mach Learn Res 9(11):66
Xie J, Girshick R, Farhadi A (2016) Unsupervised deep embedding for clustering analysis. In: International conference on machine learning. PMLR, pp 478–487
Guo X, Gao L, Liu X, Yin J (2017) Improved deep embedded clustering with local structure preservation. In: Ijcai, pp 1753–1759
Yang B, Fu X, Sidiropoulos ND, Hong M (2017) Towards k-means-friendly spaces: simultaneous deep learning and clustering. In: International conference on machine learning. PMLR, pp. 3861–3870
Gulrajani I, Ahmed F, Arjovsky M, Dumoulin V, Courville A (2017) Improved training of Wasserstein Gans. arXiv preprint arXiv:1704.00028
Mohamed S, Lakshminarayanan B (2016) Learning in implicit generative models. arXiv preprint arXiv:1610.03483
Bengio Y, Louradour J, Collobert R, Weston J (2009) Curriculum learning. In: Proceedings of the 26th annual international conference on machine learning, pp 41–48
Kumar M, Packer B, Koller D (2010) Self-paced learning for latent variable models. Adv Neural Inf Process Syst 23:66
Vardakas G, Likas A (2022) Implicit maximum likelihood clustering. In: IFIP International conference on artificial intelligence applications and innovations. Springer, pp 484–495
Lloyd S (1982) Least squares quantization in pcm. IEEE Trans Inf Theory 28(2):129–137
Tasoulis S, Pavlidis NG, Roos T (2020) Nonlinear dimensionality reduction for clustering. Pattern Recognit 107:107508
Milligan GW, Cooper MC (1988) A study of standardization of variables in cluster analysis. J Classif 5(2):181–204
Zheng GX, Terry JM, Belgrader P, Ryvkin P, Bent ZW, Wilson R, Ziraldo SB, Wheeler TD, McDermott GP, Zhu J (2017) Massively parallel digital transcriptional profiling of single cells. Nat Commun 8(1):1–12
Dua D, Graff C (2017) UCI machine learning repository. http://archive.ics.uci.edu/ml
Frey BJ, Dueck D (2007) Clustering by passing messages between data points. Science 315(5814):972–976
Hull JJ (1994) A database for handwritten text recognition research. IEEE Trans Pattern Anal Mach Intell 16(5):550–554
Rendón E, Abundez I, Arizmendi A, Quiroz EM (2011) Internal versus external cluster validation indexes. Int J Comput Commun 5(1):27–34
Kuhn HW (2005) The Hungarian method for the assignment problem. Nav Res Logist 52(1):7–21
Estévez PA, Tesmer M, Perez CA, Zurada JM (2009) Normalized mutual information feature selection. IEEE Trans Neural Netw 20(2):189–201
Hubert L, Arabie P (1985) Comparing partitions. J Classif 2(1):193–218
Chacón JE, Rastrojo AI (2022) Minimum adjusted rand index for two clusterings of a given size. Adva Data Anal Classif 66:1–9
Kingma DP, Ba J (2014) Adam: a method for stochastic optimization. arXiv preprint arXiv:1412.6980
Arthur D, Vassilvitskii S (2006) k-means++: the advantages of careful seeding. Technical Report 2006-13, Stanford InfoLab. http://ilpubs.stanford.edu:8090/778/
Acknowledgments
We would like thank Spyridon Tzimas for his valuable contribution to the experimental part of this work.
This research was supported by project “Dioni: Computing Infrastructure for Big-Data Processing and Analysis” (MIS No. 5047222) co-funded by European Union (ERDF) and Greece through Operational Program “Competitiveness, Entrepreneurship and Innovation,” NSRF 2014-2020.
Funding
Open access funding provided by HEAL-Link Greece.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Vardakas, G., Likas, A. Neural clustering based on implicit maximum likelihood. Neural Comput & Applic 35, 21511–21524 (2023). https://doi.org/10.1007/s00521-023-08524-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00521-023-08524-x