Abstract
Riverbank filtration systems are important drinking water resources. Aquifers of riverbank filtration systems are subjected to considerable dynamics concerning the quantity and quality of the infiltrating water. The microbiological quality is mainly jeopardized by faecal contamination of the main river. Besides, water quality can be impacted by growth of natural water-borne bacteria due to the input of nutrients resulting in the proliferation of opportunistic pathogens, impairment of odour and taste or bio-corrosion. The occurrence of such phenomena indicates a biological instability. For highly dynamic riverbank filtration systems, it is thus of high relevance to assess the biological stability of the groundwater resource.
In the present study, we applied a holistic, two-tiered concept of in situ and predictive methods to assess the biostability of the aquifer in a bank filtration system of the Danube River. We applied traditional cultivation-based and selected cultivation-independent methods—including cultivation on yeast extract and R2A agar, determination of total cell counts via fluorescence microscopy and flow cytometry, leucine incorporation and 16S rRNA gene amplicon sequencing—at critical control points along the infiltration path from the river to the abstraction well.
The concentration of organic nutrients and the hydrological variability were the main controlling factors driving the biological stability of the groundwater body. Wells situated at greater distance displayed significantly lower dissolved organic carbon concentrations and a dampened hydrological influence in comparison to the well situated next to the river. Apparent discrepancies between the methods used indicated a different indicator function of the cultivation-based and cultivation-independent approaches. For complex systems, we thus recommend this new holistic concept for assessing biostability by combining in situ as well as predictive parameters and using cultivation-based and cultivation-independent methods.
Zusammenfassung
Flussuferfiltrationssysteme sind wichtige Trinkwasserressourcen. Die Aquifere von Uferfiltrationssystemen sind einer erheblichen Dynamik in Bezug auf die Menge und Qualität des infiltrierenden Wassers unterworfen. Die mikrobiologische Qualität wird hauptsächlich durch die fäkale Verunreinigung des Hauptflusses gefährdet. Darüber hinaus kann die Wasserqualität durch das Wachstum natürlicher wasserbürtiger Bakterien aufgrund von Nährstoffeinträgen beeinträchtigt werden, was zur Vermehrung opportunistischer Krankheitserreger, zur Beeinträchtigung von Geruch und Geschmack oder zur Biokorrosion führen kann. Das Auftreten solcher Phänomene deutet auf eine biologische Instabilität hin. Für hochdynamische Uferfiltrationssysteme ist es daher von großer Bedeutung, die biologische Stabilität der Grundwasserressource zu bewerten.
In der vorliegenden Studie wurde ein ganzheitliches, zweistufiges Konzept aus In-situ- und Vorhersage-Methoden angewandt, um die biologische Stabilität des Aquifers in einem Uferfiltrationssystem der Donau zu bewerten. Wir wendeten traditionelle kultivierungsbasierte und ausgewählte kultivierungsunabhängige Methoden – einschließlich Kultivierung auf Hefeextrakt und R2A-Agar, Bestimmung der Gesamtzellzahl mittels Fluoreszenzmikroskopie und Durchflusszytometrie, Leucin-Inkorporation und 16S rRNA-Genamplikon-Sequenzierung – an kritischen Kontrollpunkten entlang des Infiltrationspfads vom Fluss zum Entnahmebrunnen an.
Die Konzentration der organischen Nährstoffe und die hydrologische Variabilität waren die wichtigsten Einflussfaktoren für die biologische Stabilität des Grundwasserkörpers. Brunnen in größerer Entfernung wiesen deutlich niedrigere Konzentrationen an gelöstem organischem Kohlenstoff und einen gedämpften hydrologischen Einfluss im Vergleich zum Brunnen in Flussnähe auf. Offensichtliche Diskrepanzen zwischen den verwendeten Methoden deuten auf eine unterschiedliche Indikatorfunktion der kultivierungsbasierten und kultivierungsunabhängigen Ansätze hin. Für komplexe Systeme empfehlen wir daher dieses neue ganzheitliche Konzept zur Bewertung der Biostabilität durch die Kombination von In-situ- und Vorhersage-Parametern und die Verwendung kultivierungsbasierter und kultivierungsunabhängiger Methoden.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Riverbank filtration systems are important drinking water resources (Ray 2002; Kühn and Müller 2000). In Europe, many countries rely on groundwater abstraction from groundwater aquifers along the Danube River (Kirschner et al. 2015). In Austria, such porous aquifers along the Danube serve as the main or back-up drinking water resource for more than 2 million people (e.g. cities of Vienna, Linz, Tulln a. o.). In contrast to deep groundwater resources, aquifers of riverbank filtration systems are subjected to considerable dynamics concerning the quantity and quality of the infiltrating water, caused by fluctuating river water levels, heavy precipitation events or varying pollution levels. In order to provide safe drinking water of the best chemical and microbiological quality to consumers, European water utilities are obliged to adopt a complete risk-based approach on water safety (EU 2020). This approach consists of three components, including the implementation of a water safety plan (WSP) according to WHO guidelines (WHO 2005) and the assessment of potential risks stemming from distribution systems. A WSP includes the comprehensive assessment of the drinking water resource and production line (“from the source to the tap”) and the identification of critical control points for water quality determination, including chemical and microbiological analysis.
The microbiological quality of drinking water from riverine resources in developed countries is predominantly jeopardized by faecal transmitted microorganisms (i.e. pathogenic bacteria, viruses, parasites) from point-source faecal pollution input of treated wastewater or combined sewer overflows (e.g. Kirschner et al. 2017; Demeter et al. 2021). However, drinking water quality can also be impacted by the unwanted growth of water-borne bacteria due to the input and availability of inorganic and organic nutrients potentially leading to the proliferation of opportunistic pathogens, impairment of odour and taste or bio-corrosion. The occurrence of such phenomena indicates a biological instability of the distributed water. For riverbank filtration systems with fluctuating nutrient and pollution levels, it is thus of high relevance to assess the biological stability of the groundwater resource before distribution in addition to faecal pollution surveillance.
Traditional microbiological methods to assess biological stability rely on cultivation-based approaches such as the determination of the number of heterotrophic bacteria (heterotrophic plate counts, Reasoner 1990), the determination of the assimilable organic carbon (AOC) after Van der Kooij (1992) and the determination of the growth potential after Wricke et al. (2002). Alternatively, the determination of the biodegradable portion of the organic carbon (BDOC) has been established (Servais et al. 1987). Modern and holistic concepts include the application of cultivation-independent approaches considering a combination of “in situ” and “predictive” methods (Prest et al. 2016; Fillinger et al. 2019). Cultivation-independent in situ methods comprise the direct determination of (i) total bacterial cell counts via epifluorescence microscopy (Servais et al. 1992b) or flow cytometry (Hammes et al. 2008), (ii) the viability and activity of the bacterial populations via flow cytometry (Hammes et al. 2011) or solid-phase cytometry (Riepl et al. 2011), ATP measurements (Hammes et al. 2010) or the incorporation of radiolabelled substrates (Servais et al. 1992a) and (iii) the bacterial community composition by high throughput sequencing approaches (Vierheilig et al. 2015; Fiedler et al. 2018). Such methods can also be applied for the predictive approach to assess biostability (determination of the growth potential of the bacterial community), i.e. batch growth experiments in the lab where the water is analysed before distribution, and the tests are used to predict the extent of growth that could potentially occur during water distribution (Prest et al. 2016).
In the present study, we applied this modern, two-tiered concept of in situ and predictive methods for a riverbank filtration system of the Danube River in Austria. We compared the traditional cultivation-based with a selection of cultivation-independent approaches to assess the biostability of the porous groundwater resource at critical control points along the groundwater flow-path from the river to the drinking water abstraction well. Using this approach, we could evaluate the meaningfulness of the different methods concerning biostability and comprehensively assess the biostability of the groundwater resource of the investigated riverbank filtration system. The used cultivation independent methods such as total cell counts, leucine incorporation and 16S rRNA gene sequencing fundamentally extend the traditional approach and allow for the first time a detailed look into the inner life of the drinking water “microcosm”, at the level of total bacterial community biomass, activity and composition. With this, a more comprehensive and efficient problems solution is ensured.
2 Study design and methodology
As described above, a two-tiered approach was applied to study the biostability of a porous groundwater resource within a riverbank filtration system of the Danube River in Austria.
For assessment of in situ parameters of biostability, water samples were taken in approximate one-month intervals over a 20 month period (n = 22; Oct 2014–May 2016) along a sequence of 4 groundwater wells situated in increasing distance to the river (20 to 536 m) and following a groundwater gradient from the river to the abstraction well. Groundwater for all microbiological analyses was collected in combusted (muffle furnace, 550 °C, 4 h) 5‑L glass bottles after pumping each well for 45 min (van Driezum et al. 2017). Basic physico-chemical parameters (level of water table, pH, water temperature, electrical conductivity, oxygen content) were immediately determined with portable meters. Danube discharge and precipitation data were obtained from a publicly available website (https://www.noel.gv.at/wasserstand/#/de/Messstellen). Additional clean 1‑L plastic bottles were filled for chemical analysis after rinsing three times with sample water. The samples were immediately brought to the laboratory and aliquoted for the different parameters.
Cultivation-based and cultivation-independent methods were applied. Heterotrophic plate counts (HPC) were determined after incubation at 22 ± 1 °C for 7 days on standard yeast extract agar (YEA, OENORM EN ISO 6222 1999) and at 27 ± 2 °C for 7 days on R2A agar (Reasoner and Geldreich 1985). Total cell counts (TCC) were determined by epifluorescence microscopy (van Driezum et al. 2018). Bulk bacterial activity was measured via the incorporation of radio-labelled 3H‑leucine into bacterial biomass for 24 h (LI) according to the protocol published in van Driezum et al. (2018). All chemical analysis of organic and inorganic nutrients (dissolved organic carbon, nitrate, ammonium) was performed according to standard methods (van Driezum et al. 2018).
For the predictive assessment of biostability, the growth potential of the natural bacterial community was determined in batch growth experiments over a period of 6 months (n = 10; Nov 2015–May 2016). Triplicate 250 ml aliquots were filled without air bubbles in combusted (550 °C, 4 h) and tightly sealed narrow-neck glass bottles with ground glass-stoppers and stored in the dark at 10 ± 1 °C (a representative groundwater temperature) for 7 days. Special care was taken during filling and handling the bottles to avoid contamination with organic matter. At day 7, the bottles were treated in an ultrasonic bath for 5 min, then opened and the water sample was aliquoted for the determination of the different parameters. Bacterial colonies were determined by plating various dilutions (0.001 to 1 ml) on R2A agar and incubating the plates at 27 ± 2 °C for 7 days. Total cell counts were determined via flow cytometric analysis with an Attune NxT flow cytometer (Thermo Fisher Scientific, Schrammel et al. 2018). For one representative batch growth experiment (Nov 2015), the composition of the total bacterial community was determined by 16S rRNA gene amplicon sequencing according to the protocol of Savio et al. (2019). Fig. 1 gives an overview on the different methods applied in this study.
Visualization of the different in situ and predictive parameters to assess the biostability of groundwater used in this study. Cultivation-based methods (left panel): heterotrophic plate counts as a standardized parameter. Sampling is followed by incubation on growth media for a defined time and by enumerating the number of colonies on the plates. Cultivation-independent methods (right panel): total bacterial cell counts, incorporation of radiolabeled substrates (leucine), and 16S rRNA gene amplicon sequencing as cultivation-independent diagnostic tools. Sampling is followed by (i) staining the bacteria with SYBR-Green and counting in a flow cytometer or fluorescence microscope (total cell count), (ii) incubation with radio-labelled leucine for 24 h and measurement of radioactivity in the bacterial proteins via liquid scintillation counting (leucine incorporation) and (iii) filtration, DNA extraction, amplification of the 16S rRNA gene by PCR and high through-put sequencing (16S-rRNA gene amplicon sequencing)
3 Results
3.1 In situ monitoring
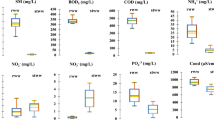
Over the whole investigation period, both cultivation-based in situ methods, i.e. HPC counts on YEA and R2A medium, followed the same pattern with an absolute peak at the end of January, 2016 (Fig. 2). Incubation on R2A agar resulted in significantly higher values than incubation on YEA (paired T‑test; average difference of 43 colony forming units (CFU)/ml; T = 3.06; p < 0.01) but log-transformed data of both methods were highly significantly inter-correlated (rho = 0.97; p < 0.001; Table 1). For both media, colony counts markedly decreased with increasing distance to the river (rho = −0.85 and −0.87 for YEA and R2A respectively, P < 0.001). Highest values were observed for the well located 20 m away from the river with values ranging from 22 to 1080 CFU/ml determined with YEA and from 28 to 2180 CFU/ml determined with R2A. In the abstraction well located 536 m away from the river, colony counts ranged from 0 to 9 CFU/ml only for YEA and from 0 to 28 CFU/ml on R2A (Fig. 2).
Seasonal variability of cultivation-based (left panel: heterotrophic plate counts on R2A and yeast extract agar, YEA) and cultivation-independent methods (right panel: total cell counts, TCC and leucine incorporation rates, LI) in four groundwater wells from a river bank abstraction system of the Danube River in Austria during the investigation period (Oct. 2014–May 2016). The wells are situated in increasing distance from the river; the names of the wells indicate their distance (in m) to the river. Bars indicate the standard deviation of three (colony counts) or four (LI) replicate measurements. No LI measurements available on 3 dates in 2016, no HPC measurements available on one date in 2014
Both cultivation-independent in situ assays showed highly similar patterns as the cultivation assays. Both, total cell counts and leucine incorporation rates peaked at the beginning of the investigation period and at the end of January 2016. Log-transformed TCC and LI data were highly significantly inter-correlated (rho = 0.82, p < 0.001) and highly significantly correlated with the cultivation assays, with correlation coefficients ranging from 0.81–0.90 (p < 0.001, see Table 1).
For both, TCC and LI, highest variability was observed in the two wells situated next to the Danube River (Fig. 2) and significantly decreased with increasing distance to the river (rho = −0.82 and −0.88 for TCC and LI respectively, p < 0.001). Cell counts ranged from 134,500 to 328,000 cells ml−1 in well 20 m and from 58,000 to 307,000 cells ml−1 in well 33 m. In the groundwater abstraction well 536 m, cell count variability was markedly dampened and ranged from 48,000 to 95,500 cells ml−1. Similarly, LI rates ranged from 0.7 to 23.9 fmol ml−1 day−1 in well 20 m and from 0.31 to 9.9 fmol ml−1 day−1 in well 33 m. More stable conditions with much lower incorporation rates were observed for the groundwater abstraction well 536 m, with values ranging from 0.07 to 0.41 fmol ml−1 day−1.
3.1.1 Correlation with environmental parameters
When pooling the whole data set, all biostability in situ parameters were, next to the negative correlation with the distance from the river, highly significantly correlated with the amount of organic matter (DOC; rho = 0.52–0.57, p < 0.001; Table 2). No other environmental variable showed a significant relationship. Neither inorganic nitrogen compounds, nor precipitation events nor the seasonal temporal cycle had significant impact on the in situ biological stability parameters in the investigated system.
However, when only the data from well 20 m, situated in closest proximity to the Danube, was considered in correlation analysis, a significant positive correlation of discharge with all in situ parameters (TCC: rho = 0.49, p < 0.05; LI: rho = 0.59, p < 0.01; R2A: rho = 0.57, p < 0.01; YEA: rho = 0.60, p < 0.01) was observed. For all other wells with a higher distance to the Danube no significant relationship of the biostability parameters with discharge was observed.
3.2 Predictive biostability assessment
For the series of batch growth experiments for predictive biostability assessment, determination of total cell counts by flow cytometry and cultivation on R2A medium were applied. The increase in cell counts and colony counts was determined after incubation at 10 °C for 7 days. With the cell-based approach, a gradient of increasing biological instability with increasing distance to the river (20 m = 33 m > 292 m > 536 m) and significant correlation with distance (rho = 0.79, p < 0.001) was observed. As for the in situ parameters, values peaked at the end of January, 2016. A smaller peak was observed at the end of March, 2016 in well 20 and 33 m, where no samples of well 292 and 536 m are available. In contrast, with the cultivation-based approach well 292 m displayed the highest biological instability, with peaks at the end of January and middle of April, 2016. Despite its vicinity to the Danube, well 20 m had HPC values at a partly similar low level as well 536 m. Despite of these discrepancies, HPC correlated significantly with TCC (rho = 0.49, p < 0.01). No correlation with the distance from the Danube was observed (see Fig. 3).
Seasonal variability of heterotrophic plate counts on R2A medium and total cell counts (TCC) in batch growth experiments performed during the investigation of 4 groundwater wells in a river bank filtration system of the Danube River in Austria. The wells are situated in increasing distance from the river; the names of the wells indicate their distance (in m) to the river. Bars indicate the standard deviation of three replicate measurements
3.2.1 Correlation with in situ biostability and environmental parameters
R2A colony counts from batch growth experiments did not significantly correlate with any in situ indicator of biostability or any environmental variable, except with YEA colony counts (Table 3). In contrast, TCC counts from batch growth experiments correlated significantly with all in situ biostability parameters as well as with the distance from the river and with DOC (Table 3).
Surprisingly, a weak significant negative correlation occurred with groundwater temperature. No significant relationship to precipitation and discharge or to the inorganic nitrogen compounds was observed. When only the data from well 20 m, situated in closest proximity to the Danube, was considered in correlation analysis, no significant correlation of predictive biostability parameters with discharge occurred, either. In contrast, in situ biostability parameters did show significant correlation with discharge in this well (see 3.1.1). The smaller sample size for the predictive parameters and the fact that sampling was not guided by discharge, most likely prevented the detection of a significant positive relationship.
3.2.2 Determination of the bacterial community composition by 16S rRNA gene amplicon sequencing
Discrepancies between the cell-based and cultivation-based approaches for predictive biostability assessment were observed (see above). As both approaches target different levels of the groundwater bacterial community (total community vs. the culturable, copiotrophic subpopulation), specific disagreements between the methods can be expected. With 16S rRNA gene amplicon sequencing, the black box of bacterial community composition can be opened and allows explaining the observed differences. For one representative sampling date (Nov 30, 2015), the groundwater samples collected from the four wells were analysed for the bacterial community composition via 16S rRNA gene amplicon sequencing, both at the time of sampling (T0) and after 7 days of incubation of the batch growth experiments (T7) (Fig. 4). The results indicate that in the two wells with high distance to the Danube, the bacterial communities at T0 are rather similar to each other. At the class level (Fig. 4a), sequence reads of representatives of Omnitrophia make up approximately 40% of all bacterial reads in both wells, other classes show a comparable relative abundance. In contrast, in the wells next to the Danube Omnitrophia are much less abundant and a considerable amounts of Actinobacteria (specifically in well 33 m) and Brocadiae (specifically in well 20 m) are present. Interestingly, gamma-Proteobacteria (Fig. 4a) and specifically those belonging to the family Pseudomonadaceae (Fig. 4b), comprising well-known biofilm forming and opportunistic pathogenic species (e.g. Pseudomonas aeruginosa) were not detectable at T0, but showed massive abundance in all 4 wells at T7 of the batch growth experiments. This happened specifically in well 292 m, where in one replicate after 7 days of incubation, more than 50% of the total population was composed of Pseudomonas spp. In the abstraction well 536 m, Hydrogenophaga (belonging to the family Comamonadaceae) became another dominating bacterial group, while in well 33 and 20 m members of the families Methylophilaceae and Oxalobacteriaceae became dominant, respectively.
Composition of the bacterial communities in groundwater samples of the 4 wells of the investigated riverbank filtration system of the Danube in Austria. a Composition at class level, b composition at family level. T0: samples at time of sampling, T7: samples after 7 days of batch incubation at 10 °C for predictive biostability assessment. Bars in different colours represent the relative abundance of sequence reads (in %) of the dominant bacterial groups. (R1–R3 three biological replicates)
4 Discussion
4.1 Elucidating the factors driving biological stability in the Danube riverbank filtration system
In the investigated riverbank filtration system, a dominant complex of factors drives the biological stability of the groundwater body. This complex comprises the concentration of organic nutrients (DOC) and the hydrological variability of the river, both of them depending on the distance of the groundwater wells to the river following the groundwater gradient from the river to the abstraction well. Except for the cultivation-based predictive biostability approach, all in situ and predictive biostability parameters declined with increasing distance to the Danube. Wells situated further from the river displayed significantly lower DOC concentrations and also a dampened hydrological influence in comparison to well 20 m situated next to the river. Only in this well in situ biological stability parameters correlated with river discharge. It is well known that the availability of organic carbon often represents the limiting factor of bacterial growth in raw drinking water (e.g. Van der Kooij 1995; Prest et al. 2016), aside from water temperature (Prest et al. 2016). In the system investigated here, the dynamics in water temperature was no obvious controlling factor, although a weak negative correlation between TCC development and temperature could be observed in the batch growth cultures. Inorganic nutrients or precipitation did also not appear to be decisive factors concerning biostability. It has been reported recently (van Driezum et al. 2018) that the bacterial community in the groundwater body of this investigated riverbank filtration system is massively influenced by flood levels. During such flood events, river water with high nutrient and bacterial load infiltrates into the aquifer, leading to a spontaneous increase in bacterial numbers and to a detachment of bacteria from desiccated subsurface particles.
4.2 The new holistic concept to assess biostability and specific assets and drawbacks of the different biostability parameters
In this study, a new holistic concept, adapted from Prest et al. (2016), was applied to assess the biostability of a groundwater body in a riverbank filtration system. This concept is based on the use of (i) in situ parameters and (ii) predictive parameters of biostability. As the widely used cultivation-based microbiological parameters only target a sub-part of the bacterial communities (van Nevel et al. 2017), cultivation-independent methods to quantify total bacterial numbers and activity were additionally used. Moreover, to open the black box of bacterial community composition, 16S rRNA gene amplicon sequencing was performed. With this method, discrepancies between cultivation-based and cultivation-independent results concerning the biostability of the investigated groundwater could be resolved.
Standard cultivation-based in situ assessment of biostability (heterotrophic plate counts) sometimes fails to lead to statistically sound results. The number of colonies can be too low to display sufficient statistical variability and the bacterial groups growing on the agar used (YEA or R2A) are not representative for the total bacterial community, as only a minor proportion is culturable. Therefore, in many systems, heterotrophic plate counts and total bacterial cell counts lead to uncoupled results (Van Nevel et al. 2017). In this study, we used R2A in addition to the standard YEA medium and prolongation of the incubation time to 7 days, in order to enhance the sensitivity of the cultivation-based assay. By this, the results of both cultivation-based assays were highly correlated with the results from the cultivation-independent methods (total bacterial cell counts and activity). This indicates that the adapted cultivation method is sensitive enough to resolve variations and that it can be used as a useful in situ method for studying biostability in this system. R2A medium increased the retrieval of culturable heterotrophic bacteria, but also cultivation on YEA delivered sufficient resolution. The reason for the observed high compliance of the cultivation-based and cultivation-independent in situ methods in this system is most likely the fact that the dominating factor influencing this system is the vicinity to and the large influence of the river. By the input of nutrients and copiotrophic bacteria from the river into the groundwater body, there is a strong co-correlation between the total bacterial community and these bacterial sub-populations. Thus both, the cultivation-based and the cultivation-independent parameters characterizing the bacterial community of this system react in the same way.
The traditional cultivation-based assay can easily be performed by routine water laboratories, but it takes up to 3 days before a result is available (Table 4). Using this method, one can also refer to legally anchored reference values; however, their absolute validity has been questioned (Van Nevel et al. 2017). The determination of total cell counts in drinking water has been established as a standard parameter in Switzerland (Schweizerisches Lebensmittelbuch 2012; Egli and Bucheli 2014) and can quickly be determined by flow cytometry (or epifluorescence microscopy) within a few hours after sampling (Table 4). This method has therefore been increasingly used for high temporal resolution water monitoring in recent years, specifically when determined with flow cytometry (Prest et al. 2013; Besmer et al. 2016). Moreover, the method can also be enhanced to distinguish between viable and dead or active and inactive cells (Hoefel et al. 2003). In contrast to the more conservative in situ biostability parameter TCC, activity parameters such as leucine incorporation show variability at a shorter time scale, as both input of external bacteria and internal regrowth are immediately reflected by higher activity rates. LI rates can be determined within 24 h and additionally allow the calculation of bacterial community growth rates and turnover (van Driezum et al. 2018). This method can only be performed in specialised laboratories due to the handling of radiolabelled material (Table 4). The reason why LI and TCC corresponded so well in this study, despite targeting different aspects of biostability (biomass vs activity) is the fact that the hydrology driven input of external bacteria dominates the internal growth phenomena (van Driezum et al. 2018) and the “slower” parameter TCC therefore reacts similarly as the “quicker” parameter LI.
Concerning the predictive biostability parameters, obvious differences between the cultivation-based and the cultivation-independent approach were observed, leading to different conclusions concerning the biostability of the investigated wells. While the TCC method coincides with the results of the in situ parameters, with increasing instability towards the Danube River and dependence on the concentration of organic matter, the cultivation on R2A medium indicated that well 292 m with a high distance to the river has the highest biological instability. 16S rRNA gene amplicon sequencing revealed that specifically in this well, representatives of the genus Pseudomonas massively increased in the batch growth experiments, a genus that includes important bacterial pathogens (e.g. Pseudomonas aeruginosa) and biofilm formers. It can thus be concluded that—under certain circumstances—the two different methodical approaches have a different indicator function concerning biostability. While TCC provide information on biostability problems concerning the bulk bacterial community, HPC might indicate biostability problems arising from specific bacterial subpopulations that are overlooked when the total population is analysed (Table 4). Thus, both approaches should be considered when biostability is to be assessed in a holistic way. Leucine incorporation is another method that could provide additional information concerning predictive biostability assessment (although not used in the growth experiments in this study). Despite small changes in total cell counts, a high LI activity could occur in batch growth experiments indicating a substantial transformation of the bacterial community in terms of community composition (Table 4).
5 Conclusions
In this study, we could successfully demonstrate the importance of using a holistic approach for the assessment of biostability. Depending on the specific problem (health problems due to growth of opportunistic pathogens, aesthetic problems due to the impairment of odour and taste or technical problems due to biocorrosion, fouling or clogging), a specifically adapted combination of cultivation-based and cultivation-independent in situ and predictive methods is likely to be most useful, as all the available traditional and emerging methods show theirs strength and limitations. The combination of TCC determined via flow cytometry, activity measurements via LI, and bacterial community analysis via 16S rRNA gene amplicon sequencing, which allows both the evaluation of biostability problems at the level of the bulk community and at the level of specific bacterial groups (Table 4) is a straight-forward and emerging, innovative approach. Specific groups of interest could even be analysed at higher resolution down to the species level, when group-specific primers (e.g. for the genus Pseudomonas) are applied (Pereira et al. 2018) for 16S rRNA gene amplicon sequencing. However, 16S rRNA gene amplicon sequencing is of limited value when it comes to the analysis at the strain level or the functional analysis of the bacterial populations, e.g. the presence of pathogenic or antimicrobial resistance genes. Aside from amplicon sequencing of functional genes, cultivation-based methods are still the gold standard to identify isolates down to the strain level and perform functional analysis of the organisms. In this respect, the use of HPC is an easy method that can be performed by standard laboratories, providing information on potential problems with copiotrophic bacteria. For specific situations, cultivation may have to be extended to specific bacteria of interest, e.g. opportunistic pathogens, biofilm formers or bacteria causing technical problems.
References
Besmer M.D., Epting J., Page R.M., Sigrist J.A., Huggenberger P., Hammes F. (2016): Online flow cytometry reveals microbial dynamics influenced by concurrent natural and operational events in groundwater used for drinking water treatment. Scientific Reports 6:38462
Demeter K., Derx J., Komma J., Parajka J., Schijven J., Sommer R., Cervero-Arago S., Lindner G., Zoufal-Hruza C.M., Linke R., Savio D., Ixenmaier S.K., Kirschner A.K.T., Kromp H., Blaschke A.P., Farnleitner A.H. (2021): Modelling the interplay of future changes and wastewater management measures on the microbiological river water quality considering safe drinking water production. Science of the Total Environment 768: 144278
van Driezum I.H., Chik A.H.S., Jakwerth S., Lindner G., Farnleitner A.H., Sommer R., Blaschke A.P., Kirschner A.K.T. (2018): Spatiotemporal analysis of bacterial biomass and activity to understand surface and groundwater interactions in a highly dynamic riverbank filtration system. Science of the Total Environment 627:450–461
Van Driezum I.H., Derx J., Saracevic E., Kirschner A.K.T., Sommer R., Farnleitner A.H., Blaschke A.P. (2017): Does Pumping Volume Affect the Concentration of Micropollutants in Groundwater Samples? Groundwater: Monitoring and Remediation 37: 82–88
Egli T., Bucheli M. (2014): Wie viele Zellen sind im Trinkwasser? Aqua Gas 11: 90–98
EU (2020): European Drinking Water Directive 2020/2184 of the European Parliament and of the Council of 16 December 2020 on the quality of water intended for human consumption (recast). Official Journal of the European Union L435:1–62
Farnleitner A.H., Zibuschka F., Burtscher M.M., Lindner G., Reischer G. & Mach R.L. (2004): Eubacterial 16S-rDNA amplicon profiling: a rapid technique for comparison and differentiation of heterotrophic plate count communities from drinking water. International Jorunal of Food Microbiology 92:333–345
Fiedler C.J., Schönher C., Proksch P., Kerschbaumer D.J., Mayr E., Zunabovic-Pichler M., Domig K.J., Perfler R. (2018): Assessment of Microbial Community Dynamics in River Bank Filtrate Using High-Throughput Sequencing and Flow Cytometry. Frontiers in Microbiology 9: 2887
Fillinger L., Hug K., Trimbach A.M., Wang H., Kellermann C., Meyer A., Bendinger B., Griebler C. (2019): D‑A-(C) index: a practical approach towards the microbiological-ecological monitoring of groundwater ecosystems. Water Research 163:114902
Hammes F., Berney M., Egli T. (2011): Cultivation-independent assessment of bacterial viability. Advances in Biochemistry, Engineering and Biotechnology 124, 123–150
Hammes F., Berney M., Wang Y., Vital M., Koster O., Egli T. (2008): Flow cytometric total bacterial cell counts as a descriptive microbiological parameter for drinking water treatment processes. Water Research 42, 269–277
Hammes F., Goldschmidt F., Vital M., Wang Y., Egli T. (2010): Measurement and interpretation of microbial adenosine tri-phosphate (ATP) in aquatic environments. Water Research 44, 3915–3923
Hoefel D., Grooby W.L., Monis P.T., Andrews S., Saint C.P. (2003): Enumeration of water-borne bacteria using viability assays and flow cytometry: a comparison to culture-based techniques. Journal of Microbiological Methods 55: 585–597
Kirschner A.K.T., Kavka G.G., Reischer G.H., Sommer R., Blaschke A.P., Stevenson M., Vierheilig J., Mach R.L., Farnleitner A.H. (2015): Microbiological Water Quality of the Danube River: Status Quo and Future Perspectives. In: Liska I (ed) The Danube River Basin. The Handbook of Environmental Chemistry, vol 39: 439–468; Springer, Berlin, Heidelberg.
Kirschner A.K.T., Reischer G.H., Jakwerth S., Savio D., Ixenmaier S., Toth E., Sommer R., Mach R.L., Linke R., Eiler A., Kolarevic S., Farnleitner A.H. (2017): Multiparametric monitoring of microbial faecal pollution reveals the dominance of human contamination along the whole Danube River. Water Research 124:543–555.
Van der Kooij D. (1992): Assimilable organic carbon as an indicator of bacterial regrowth. Journal AWWA 84: 57–66
Van der Kooij D. (1995): Significance and Assessment of the Biological Stability of Drinking Water. Handbook of Environmental Chemistry 5: 89–102
Kühn W., Müller U. (2000): Riverbank Filtration: An Overview. Journal of the American Waterworks Association 92:60–69
Van Nevel S., Koetzsch S., Proctor C., Besmer M.D., Prest E., Vrouwenvelder J.S., Knezev A., Boon N., Hammes F. (2017): Flow cytometric bacterial cell counts challenge conventional heterotrophic plate counts for routine microbiological drinking water monitoring. Water Research 113: 191–206
OENORM EN ISO 6222 (1999): Water quality—Enumeration of culturable micro-organisms—Colony count by inoculation in a nutrient agar culture medium (ISO 6222:1999). Austrian Standards, Vienna, Austria [in German]
Pereira R.P.A., Peplies J., Mushi D., Brettar I., Höfle M.G. (2018): Pseudomonas-Specific NGS Assay Provides Insight Into Abundance and Dynamics of Pseudomonas Species Including P. aeruginosa in a Cooling Tower. Frontiers in Microbiology 9:1958
Prest E.I., Hammes F., Kötzsch S., van Loosdrecht M.C., Vrouwenvelder J.S. (2013): Monitoring microbiological changes in drinking water systems using a fast and reproducible flow cytometric method. Water Research 47: 7131–42
Prest E.I., Hammes F., van Loosdrecht M.C.M., Vrouwenvelder J.S. (2016): Biological Stability of Drinking Water: Controlling Factors, Methods, and Challenges. Frontiers in Microbiology 7: 45
Ray C. (2002): Riverbank filtration: understanding contaminant biogeochemistry and pathogen removal. NATO Science Series. IV Environmental Sciences 14, Kluwer Academic Publishers, Dordrecht/Boston/London
Reasoner D.J. (1990): Monitoring heterotrophic bacteria in potable water. In: Mc—Feters GA (Ed), Drinking Water Microbiology. Springer-Verlag, NewYork
Reasoner D.J., Geldreich E.E. (1985): A new medium for the enumeration and subculture of bacteria from potable water. Applied and Environmental Microbiology 49: 1–7.
Riepl M., Schauer S., Knetsch S., Holzhammer E., Farnleitner A.H., Sommer R., Kirschner A.K.T. (2011): Applicability of solid-phase cytometry and epifluorescence microscopy for rapid assessment of the microbiological quality of dialysis water. Nephrology Dialysis Transplantation 26:3640–3645
Savio S., Stadler P., Reischer G.H., Demeter K., Linke R.B., Blaschke A.P., Stadler H., Mach R.L., Kirschner A.K.T., Farnleitner A.H. (2019): Spring water of an alpine karst aquifer is dominated by a taxonomically stable but discharge-responsive bacterial community. Frontiers in Microbiology 10:28
Schrammel B., Cervero-Aragó S., Dietersdorfer E., Walochnik J., Lück C., Sommer R., Kirschner A.K.T. (2018): Differential development of Legionella sub-populations during short- and long-term starvation. Water Research 141:417–427
Schweizerisches Lebensmittelbuch (2012): Untersuchungsmethode 333.1: Bestimmung der Totalzellzahl und des quantitativen Verhältnisses der Zellen niedrigen bzw. hohen Nukleinsäuregehaltes in Süsswasser mittels Durchflusszytometrie. Bundesamt für Gesundheit, Bern
Servais P., Billen G., Hascoet M.C. (1987): Determination of the biodegradable fraction of dissolved organic matter in waters. Water Research 21: 445–450
Servais P., Billen G., Laurent P., Levi Y., Randon G. (1992a): Studies of BDOC and bacterial dynamics in the drinking water distribution system of the Northern Parisian suburbs. Revue des Sciences de L’eau 5: 473–488
Servais P., Laurent P., Randon G. (1992b): Measurement of bacterial biomass and activity in tap water. Revue des Sciences de L’eau 5:69
Vierheilig J., Savio D., Ley R.E., Mach R.L., Farnleitner A.H., Reischer G.H. (2015): Potential applications of next generation DNA sequencing of 16S rRNA gene amplicons in microbial water quality monitoring. Water Science and Technology. 72: 1962–1972
World Health Organization (2005): Water Safety Plans: Managing drinking-water quality from catchment to consumer. World Health Organization, Geneva, Switzerland, 234 pp
Wricke B., Petzoldt H., Korth A., Krüger M., Andrusch T., Böhm U., Häusler H. (2002): Minimierung der Desinfektionsnebenprodukte und der Wiederverkeimung im Verteilungsnetz von Fernwasserversorgungssystemen. Abschlussbericht zum BMBF-Vorhaben, Schriftenreihe des TZW, Heft 17
Acknowledgements
The work of this paper was conducted within the “Aquascreen”-project funded by the FTI-program of the province of Lower Austria (“FTI—Forschungs‑, Technologie- und Innovationsprogramm Niederösterreich”) as part of the project call “Ernährung—Medizin—Gesundheit” (project number WST3-F-5031298/001-2017). Further support was provided by Wiener Wasser (MA 31) via the project “Ground Water Resource Systems Vienna (GWRS)”.
Funding
Open access funding provided by Medical University of Vienna.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kirschner, A.K.T., Lindner, G., Jakwerth, S. et al. Assessing biological stability in a porous groundwater aquifer of a riverbank filtration system: combining traditional cultivation-based and emerging cultivation-independent in situ and predictive methods. Österr Wasser- und Abfallw 73, 490–500 (2021). https://doi.org/10.1007/s00506-021-00801-0
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00506-021-00801-0
Keywords
- Biostability
- Bank filtration
- Cultivation
- Cultivation independent
- Cell count
- Flow cytometry
- Bacterial growth
- High through-put sequencing