Abstract
Variation in disease risk underlying observed disease counts is increasingly a focus for Bayesian spatial modelling, including applications in spatial data mining. Bayesian analysis of spatial data, whether for disease or other types of event, often employs a conditionally autoregressive prior, which can express spatial dependence commonly present in underlying risks or rates. Such conditionally autoregressive priors typically assume a normal density and uniform local smoothing for underlying risks. However, normality assumptions may be affected or distorted by heteroscedasticity or spatial outliers. It is also desirable that spatial disease models represent variation that is not attributable to spatial dependence. A spatial prior representing spatial heteroscedasticity within a model accommodating both spatial and non-spatial variation is therefore proposed. Illustrative applications are to human TB incidence. A simulation example is based on mainland US states, while a real data application considers TB incidence in 326 English local authorities.
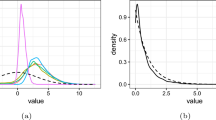
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Modelling variation in disease or other events underlying observed totals for geographic areas is important for detecting elevated rates (Beale et al. 2008). In disease mapping, the observations often consist of incidence totals for chronic or infectious disease. Such data are subject to stochastic variations, and the underlying area specific incidence risks are often the focus in data mining studies. In such studies, the objects include extraction of underlying spatial and spatiotemporal patterns, including detection of elevated risk (hotspots) and spatial outliers (Shekhar et al. 2015). The particular focus of this paper is on ecological epidemiology, in the sense of focusing on population aggregates (Morgenstern 1995), namely geographic areas, and on environmental and socio-economic risk factors for infectious disease (Ploubidis et al. 2012). The applications are to human infectious disease, namely TB incidence.
Different forms of spatial correlation analysis or model have been proposed in disease applications (human and veterinary), environmental science, ecology, crime and other settings. For example, Wikle (2003) reviews hierarchical spatial models applied in environmental science, including irregular lattice data (such as geographic areas) and regular lattice data (such as air pollution grids). Beale et al. (2010) consider how regression findings for spatial ecology data are affected by the method used (if at all) to reflect spatial dependence. To exemplify hierarchical models for veterinary data, Pioz et al. (2012) apply simultaneous autoregressive (SAR) models to investigate bluetongue spread in French municipalities, while Farnsworth and Ward (2009) apply Bayesian conditional autoregressive (CAR) models to avian influenza H5N1 outbreak data. In such applications, identifying elevated risk in particular areas, detecting elevated risk clusters, or assessing significant predictors of risk, are emphasized, in methods recognizing the explicitly spatial structure of the data. However, the underlying assumptions of such techniques should be assessed, and subject to modification when indicated.
Hierarchical models involving spatial random effects, both CAR and SAR forms, can be estimated by classical methods (Alam et al. 2015; Horabik and Nahorski 2010) or Bayesian methods (Waller and Carlin 2010; Lesage 1997). CAR spatial priors imply local smoothing of outcome rates, that is smoothing towards the local rather than global average (Gelman 1996; Waller and Carlin 2010). Such local discontinuity is demonstrated in the England TB application considered below. Marked variability in risks has been detected in other area studies of infectious disease (Duarte-Cunha 2015; Varga et al. 2015; Ploubidis et al. 2012), whereas spatial variability in relative risks of chronic diseases (cancer, diabetes, etc.) is generally less pronounced. When there are spatial discontinuities in risk, it is preferable to allow differing strengths of association between neighbouring areas, as opposed to uniform local smoothing under CAR priors (Gelman 1996; Smith et al. 2015).
Bayesian applications in disease mapping and ecological epidemiology commonly employ a CAR prior (Lee 2011) to express spatial clustering in underlying risks (Besag et al. 1991; Best 1999), including human TB incidence (Nunes 2007; Maciel et al. 2010). Most applications of CAR priors assume a normal density for the underlying risks combined with uniform local smoothing. However, normality assumptions may be vitiated by heteroscedasticity linked to spatial outliers or to marked discrepancies in risk between neighbouring areas. It is also desirable that spatial disease models represent variation in area disease risks that is not attributable to spatial dependence (i.e. heterogeneity as against clustering). Some spatial priors may represent this feature by using more than one set of random effects, but at the cost of identifiability.
This paper considers modification of the local smoothing principle when there are spatial discontinuities, namely discrepant levels of outcome rates (e.g. disease or crime incidence) between neighbouring areas. In particular, we consider modifications of the normality assumption for area random effects based on a scale mixture version of the Leroux et al. (1999) model, allowing for heterogeneity and clustering in a single set of random effects, but with the scale mixture providing adaptivity to local discontinuity and spatial outliers. The relevance of such an approach is illustrated with simulated data on TB incidence in 49 mainland US states, and an application to observed TB incidence in 326 English local authorities.
2 Defining conditional spatial priors
As discussed by Besag and Kooperberg (1995), one may use properties of the multivariate normal to obtain the univariate conditional autoregressive prior from a joint spatial prior and vice versa. Thus consider a joint multivariate normal density for the spatial risk effects \( {\text{s}} = ({\text{s}}_{1} , \ldots ,{\text{s}}_{{\text{n}}} ) \) for \( \text{n} \) areas, with mean zero and covariance \( \Sigma _{{\text{s}}} \),
Denote
as the precision matrix, and \( {\text{s}}_{{{\text{[i]}}}} = ({\text{s}}_{1} , \ldots ,{\text{s}}_{{{\text{i}} - 1}} ,{\text{s}}_{{{\text{i}} + 1}} , \ldots ,{\text{s}}_{{\text{n}}} ) \) as the totality of effects omitting the ith effect. The conditional distributions for each \( {\text{s}}_{\text{i}} \) take a univariate normal form (Rue and Held 2005, p. 22), namely
Following Besag and Kooperberg (1995, p 734) define \( {\text{h}}_{\text{ii}} { = 0,} \) and set
Also set
with variance parameter \( \updelta \), so that
The density (2) is then in the conditional autoregressive form specified by Besag (1974), namely
To obtain the joint density from the conditional one, symmetry of \( \text{Q} \) means \( - {\text{q}}_{\text{ij}}\,=\,- {\text{q}}_{\text{ji}} \), so that from (3) the constraint
applies.
3 Conditional autoregressive spatial priors
Various schemes for defining the \( {\text{h}}_{\text{ij}} \) and \( {\text{a}}_{\text{i}} \) can be used. A measure of spatial dependence \( 0 \le \upomega \le 1 \) is included by setting
where \( \text{w}_{ij} \) represent spatial interactions between areas i and j. If the interactions are specified as symmetric with \( {\text{w}}_{\text{ij}}\,=\,{\text{w}}_{\text{ji}} \), and also with \( {\text{w}}_{\text{ii}}\,=\,{0} \), the symmetry constraint (4) is ensured, with \( {\text{h}}_{{\text{ij}}} {\text{a}}_{{\text{i}}} = \omega {\text{w}}_{{\text{ij}}} = {\text{h}}_{{\text{ji}}} {\text{a}}_{{\text{j}}} \).
A common approach sets \( {\text{w}}_{\text{ij}}\,=\,{1} \) for adjacent areas and \( {\text{w}}_{\text{ij}} \,=\,{0} \) otherwise, with
then equal to the number, \( {\text{d}}_{\text{i}} \), of areas adjacent to area i. Equivalently, \( {\text{d}}_{\text{i}} \) is the number of areas in the locality \( {\text{N}}_{\text{i}} \) of area i (the areas surrounding area i, and excluding area i itself). This provides the conditionally autoregressive \( CAR(\upomega ) \) prior, with
where \( {\bar{\text{A}}}_{\text{i}} \) is the average of the \( \text{s}_{j} \) in locality \( \text{N}_{i} \), i.e.
Lower values of \( \upomega \) imply lesser degrees of spatial dependence between the \( {\text{s}}_{\text{i}} \), though the limiting case when \( \upomega = 0 \) has the disadvantage that the variance is not constant but depends on the number of neighbours \( {\text{d}}_{\text{i}} \). The \( {\text{CAR(1)}} \) prior (Besag et al. 1991) specifies relative risks entirely determined by spatial dependence, with
In any set of area disease rates, some spatial correlation is typically detected, and this motivates spatial priors which imply borrowing of strength from nearby areas. However, there may also be particular local variations in illness risks unrelated to those in surrounding areas, namely unstructured variation without spatial dependence. In principle, the \( CAR(\upomega ) \) prior (also called the proper CAR prior) can represent various levels of spatial dependence through the \( \upomega \) parameter, but this parameter does not calibrate well with marginal measures of spatial correlation, such as Moran’s I (Banerjee et al. 2004; Rodrigues and Assunção 2012). Values of \( \upomega \) exceeding 0.99 are needed to achieve modest values of Moran’s I.
In practice, to represent a mix between spatial dependence and simple unstructured variation, called clustering and heterogeneity respectively by Clayton et al. (1993), a common strategy is the so-called convolution prior (Ugarte et al. 2005; Waller and Carlin 2010). This represents the unknown area relative risk as a sum of a pure spatial effect following a \( {\text{CAR(1)}} \) prior, combined with an iid (or unstructured) random effect. Thus denote observed disease counts as \( {\text{y}}_{\text{i}} \), expected counts as \( \text{E}_{i} \) (expected disease counts in the demographic sense) and known area risk variables (predictors) as \( {\text{X}}_{\text{i}} \). Then one might specify
where ρi denotes an area specific relative risk, and \( \upphi \) is a variance term for iid unstructured effects \( {\text{h}}_{\text{i}} \). A drawback with this scheme is that identifiability may be impeded by the presence of two sets for random effects representing one underlying aspect of the data, namely variation in area illness risks.
4 The Leroux et al. spatial prior
A scheme for area effects, incorporating both clustering and heterogeneity, involves scale adjustments
with the parameter \( 0 \le \uplambda \le 1 \) providing a measure of spatial dependence (Leroux et al. 1999). This scheme, which may be represented as the LLB prior by virtue of its authors, has the benefit that only one set of random effects is involved in representing the pattern of area illness risks. This provides improved identifiability as compared to the convolution prior (Lee 2011). The case \( \uplambda = 0 \) corresponds to a lack of spatial interdependence (and i.i.d or unstructured errors \( {\text{s}}_{\text{i}} \)), with the advantage that the conditional variance is then simply \( \updelta \), independent of \( \sum\limits_{{{\text{j}} \ne {\text{i}}}} {{\text{w}}_{\text{ij}} } \). By contrast, \( \uplambda = 1 \) leads to a \( CAR(1) \) model, with purely spatial interdependence. In typical datasets \( \uplambda \) will be intermediate between these extreme values.
The symmetry condition \( {\text{h}}_{{{\text{ij}}}} {\text{a}}_{{\text{i}}} = {\text{h}}_{{{\text{ji}}}} {\text{a}}_{{\text{j}}} \) is maintained by setting
since \( {\text{h}}_{\text{ij}} {\text{a}}_{\text{i}} = \uplambda {\text{w}}_{\text{ij}} = \uplambda {\text{w}}_{\text{ji}} = {\text{h}}_{\text{ji}} {\text{a}}_{\text{j}} . \) So the conditional prior is
with \( \updelta \) a scale parameter. When \( \uplambda = 0 \) one obtains normal iid effects \( {\text{s}}_{\text{i}} \sim {\text{N}}(0,\,\updelta ). \) If the \( {\text{w}}_{\text{ij}} \) are defined by contiguity one obtains (Leroux et al. 1999, p 181)
5 Adaptiveness to non-normality and spatial discontinuities
Proposals to modify spatial priors to achieve greater robustness have been made, including the presence of heteroscedasticity and heavier tails (excess kurtosis) than under the normal. Thus Yan (2007), Brewer and Nolan (2007), and Reich and Hodges (2008) propose modified CAR priors to accommodate heteroscedasticity. Other forms of modified spatial prior are considered by Nathoo and Ghosh (2013) and Lawson and Clark (2002). These schemes are all modifications of the CAR prior, or of the convolution prior, as considered in Sect. 3. Modifications of the pure spatial \( CAR(1) \) prior, without allowance for spatially unstructured variation, may be appropriate for particular applications, such as dental decay as in Reich and Hodges (2008), but for area illness data an allowance for heterogeneity is generally needed. Modification of the proper \( CAR(\upomega ) \) prior are left with the problem that its \( \upomega \) parameter does not calibrate well with marginal measures of spatial correlation. Studies such as Yan (2007) and Lawson and Clark (2002) modify the convolution prior, with potential identifiability problems due to multiple sets of random effects. Thus Yan (2007) allows for heteroscedasticity in spatial effects via a double implementation of the \( CAR(1) \) prior, namely
Here we modify the constant scale assumption of the LLB prior in (7) and (8) using a scale mixture, with the benefit of providing an indicator of potential outlier status for each area. To implement a scale mixture, define \( \upkappa_{\text{i}} \sim {\text{Ga(0}} . 5\nu , 0. 5\nu ) \) where \( \nu \) is a hyperparameter. The proposed model reduces to the scale mixture version of the Student t when \( \uplambda = 0 \) (Boris Choy and Chan 2008). The \( \upkappa_{\text{i}} \) have average 1 with small values of \( \upkappa_{\text{i}} \) (under 1) acting as indicators of outlier status (West 1984). Under this scale mixture modification, the symmetry condition (4) is maintained by setting
since \( {\text{h}}_{\text{ij}} {\text{a}}_{\text{i}} = \uplambda {\text{w}}_{\text{ij}} \upkappa_{\text{j}} \upkappa_{\text{i}} = \uplambda {\text{w}}_{\text{ji}} \upkappa_{\text{i}} \upkappa_{\text{j}} = {\text{h}}_{\text{ji}} {\text{a}}_{\text{j}} \).Then the model for incidence counts becomes
where the conditional prior when the \( \text{w}_{\text{ij}} \) are binary indicators of adjacency is
This prior reduces to an unstructured i.i.d scale mixture Student-t density
when \( \uplambda = 0 \).
From (9) it can be seen that small \( \upkappa_{j} \) values indicate areas discrepant in risk from their neighbours (i.e. they indicate outliers in spatial terms), and reduce the amount of spatial borrowing of strength. Equivalently stated, a clustering of small \( \upkappa_{j} \) values can be taken as indicators of spatial volatility, namely discrepant illness risks in a set of adjacent areas. In regression applications, small \( \upkappa_{j} \) values will also indicate where the regression predictions in the neighbourhood of area \( \text{i} \), and their implied neighbourhood relative risk \( \sum\limits_{{{\text{j}} \in {\text{N}}_{{\text{i}}} }} {\upmu _{{\text{j}}} } /\sum\limits_{{{\text{j}} \in {\text{N}}_{{\text{i}}} }} {{\text{E}}_{{\text{j}}} } \), are discrepant from the modelled relative risk in area \( \text{i} \) itself \( \upmu_{\text{i}} /\text{E}_{\text{i}} \).
Let \( \uptau = 1/\updelta , \) and let \( I(i \sim j) = I(j \sim i) \) denote that areas \( i \) and \( j \) are neighbours under binary adjacency. Then the precision matrix in the multivariate normal (1) has diagonal terms
and off diagonal terms
A scale mixture approach to spatial dependence can be set within a broader literature on heavy tailed priors (e.g. student t, double exponential) that can be represented as two level hierarchical models (Yi and Xu 2008). One application of such priors is to predictor selection in high dimensional regression, with a likelihood penalty function that is a normal scale mixture (e.g. Polson et al. 2014). Besag et al. (1991) propose a double exponential prior for spatial effects as a robust alternative to the normal conditional autoregressive, with an application provided by Manda (2013).
Identification of random effects in spatial disease models is often problematic (e.g. MacNab 2014; Nathoo and Ghosh 2013), especially for models including multiple random effects, or when disease counts are relatively small. In the case of the model just discussed, identification of outliers (e.g. in terms of significantly low \( \upkappa_{i} \)), as well as identification of elevated risks \( \text{s}_{\text{i}} , \) will be improved for larger disease counts and/or longer observation periods. Identification of hyperparameters may also be problematic, especially with small samples. For example, in student t binary regression with data augmentation, Gelman et al. (2004, p 447) recommend a robust analysis with \( \nu \) not estimated but preset at 4.
6 Simulation example
A simulation example of the heteroscedastic LLB prior involves TB incidence with a spatial framework provided by the \( \text{n} = 49 \) mainland states (including the District of Columbia). Expected TB incidence counts \( \text{E}_{\text{i}} \) are obtained by applying actual US-wide age specific rates for TB in 2013 to state population estimates for 2013, taken from the US National Cancer Institute SEER site (http://seer.cancer.gov/popdata/). TB incidence rates are from the CDC National Tuberculosis Surveillance System, with just over 9500 incident cases in 2013, and an all ages rate of 3 per 100,000. Highest rates (over 6 per 100 thousand) are for the 75-84 and 85 + age groups.
We simulate TB incidence counts using these expected counts as offsets. The LLB hyperparameters (guide values) are set as \( \uplambda = 0.7, \) \( \uptau = 3 \), and with \( \nu \) taking values 3,10, and 25. Although the student t is defined for degrees of freedom of 2 or less, it has infinite variance, and Gelman et al. (2004) mention that “t’s with one or two degrees of freedom have infinite variance and are not usually realistic in the far tails”. One hundred sets of random effects are generated from the multivariate normal \( {\text{s}}_{{ 1\;\; :\;\;{\text{n}}}} \sim {\text{N(0,}}\,{\text{Q}}^{ - 1} ) \). Simulated TB incidence counts are then obtained via a Poisson simulation \( {\text{y}}_{\text{i}} \sim {\text{Po(E}}_{\text{i}} \uprho_{\text{i}} ) \), with \( \log (\uprho_{\text{i}} ) = \upbeta_{0} + \text{s}_{\text{i}} , \) where \( \upbeta_{0} = - 0.1 \), and \( \uprho_{i} \) is the simulated disease relative risk in state \( \text{i} \) (relative to that expected on the basis of US wide incidence levels). The R code used is set out in “Appendix”. Note that each of the 100 simulations involves a separate sample of \( \upkappa_{\text{i}} \sim Ga(0.5\nu ,\,0.5\nu ) \).
Analyses to estimate the parameters from the 100 sets of simulated data \( \{ y,\,E\} \) (with \( \text{E} \) as in the simulations) are carried out using the WINBUGS package (Lunn et al. 2009). An exponential prior with mean 10 is adopted for \( \nu \,\; \)(Fernandez and Steel 1998; Geweke 1993), a gamma prior with shape 1 and index 0.01 assumed for the inverse variance parameter \( \uptau \), a normal prior with mean zero and precision 0.001 assumed for the fixed effect \( \upbeta_{0} \), and a uniform \( U(0,\,1) \) prior assumed on \( \uplambda \). Estimates are based on the last 5000 iterations from two chain runs of 10,000 iterations, with convergence assessed using Brooks–Gelman–Rubin diagnostics (Brooks and Gelman 1998).
The focus is on the posterior means for the main parameters of the LLB prior and risk regression over the 100 samples, namely \( \nu ,\,\uplambda ,\,\upbeta_{0} \), and the variance of the spatial effects (which depends on both \( \uptau \) and the sampled \( \upkappa_{i} ) \). The posterior densities for \( \nu \) tend to be positively skew, so Table 1 also includes results for the posterior summary of log(\( \nu \)). Because each simulation involves a distinct set of \( \upkappa_{i} \), the actual variance of the \( \text{s}_{\text{i}} \) will vary between simulations . This variance \( \text{V}_{\text{t}} \) of spatial effects for simulation \( t \) is recorded in the vector var.s[] in the code in “Appendix”. Table 1 sets out the percentiles (20th, 50th, 80th) of the 100 posterior means for \( \nu ,\,\log (\nu ),\,\uplambda , \) and \( \upbeta_{0} \), and also the proportion of simulated datasets where the 95 % credible interval for a parameter includes the guide value. Thus for the setting \( \nu = 10 \), 50 out of the 100 samples have posterior means for \( \nu \) below 10.4, and 50 samples have posterior means above 10.4.
The expected \( \text{E}_{\text{i}} \) are relatively large, so the Poisson simulations may be subject to some excess dispersion, which to some extent attenuates the spatial structure present in the simulated data. Nevertheless, the recovered parameters effectively reproduce those used in generating the data. This feature is also apparent in a correlation between the actual and estimated \( \text{V}_{\text{t}} \) over the 100 samples of 0.97. Figure 1 plots the two series of \( \text{V}_{\text{t}} \) for the \( \nu = 10 \) option, including 95 % credible intervals for the estimated \( \text{V}_{\text{t}} \). Of substantive relevance in interpreting the parameters of the LLB model, there is a 0.72 correlation between the 100 posterior means for \( \uplambda , \) and the corresponding posterior means for Moran’s I, which are estimated from the \( \text{s}_{\text{i}} \) in each dataset. To further illustrate variation over the simulations, Fig. 2 shows, for each simulated dataset, the posterior mean (and 95 % interval) of log(\( \nu ) \) under the \( \nu = 3 \) option.
One also wishes to reproduce the patterns of outlier status (areas with significantly low \( \upkappa_{i} \)). This involves, for the setting \( \nu = 10 \) (and other hyperparameters as above), simulating 100 sets of \( \text{y} \) based on a single set of \( \upkappa_{\text{i}} \) values (the “actual” \( \upkappa_{\text{i}} ) \), sampled from a gamma density, \( \upkappa_{\text{i}} \sim Ga(5,\,5). \) The expected incidence counts are multiplied by 10 to increase the amount of information provided by the data. Re-estimation of \( \upkappa_{\text{i}} \) from the simulated datasets shows a shrinkage effect, with posterior mean re-estimated \( \upkappa_{\text{i}} \) closer to 1 than the actual \( \upkappa_{\text{i}} \) (see Fig. 3). However, the re-estimation does identify as outliers the states with unusually low actual \( \upkappa_{\text{i}} . \) For the five states with the lowest actual \( \upkappa_{\text{i}} , \) four have 95 % credible intervals on the re-estimated \( \upkappa_{\text{i}} \) that are entirely below 1, and no other states have re-estimated \( \upkappa_{\text{i}} \) with credible intervals entirely below 1.
7 Application: TB incidence for England local authorities
An application involves TB incidence data \( \text{y} \) for 326 English local authorities between 2011 and 2013. Two analyses are undertaken, one without predictors and one with two predictors: an index of multiple socio-economic deprivation (\( \text{X}_{1} \)) and population density (\( \text{X}_{2} \)). The impact of poverty on TB incidence is well documented (Lopez de Fede et al. 2008) and population density is associated with infectious disease risk as “the likelihood that a susceptible person will be exposed to an infectious tuberculosis patient increases with population density” (Rieder 1999). The two predictors are centred and divided by 100. Thus with predictors \( \text{X}_{\text{i}} = (1,\,\text{X}_{1\text{i}} ,\,\text{X}_{2\text{i}} ) \), under the scale mixture model we have
For the original Leroux et al. (1999) scheme, the conditional prior for \( \text{s}_{\text{i}} \) is as in (8).
Prior settings are as in Sect. 6, and inferences are from the last 5000 iterations from two chain runs of 10,000 iterations, with convergence assessed using Brooks–Gelman–Rubin diagnostics. Table 2 contains parameter summaries and comparison of measures of fit between the original LLB model (Sect. 4) and the heteroscedastic Leroux (Sect. 5). Fit is assessed using the posterior predictive loss (PPL) criterion (Gelfand and Ghosh 1998). Consider replicate observations \( \text{y}_{\text{rep}} \) sampled from the posterior predictive density \( \text{p}(\text{y}_{\text{rep}} |\text{y}) \). The PPL involves defining \( \text{t(z)} = \text{z}\log \text{z} - \text{z} \), and \( \xi_{\text{i}} = \text{t(y}_{\text{i},\,\text{rep}} ). \) Letting \( \eta_{i} \) and \( \phi_{i} \) denote posterior means for \( \text{y}_{\text{i},\,\text{rep}} \) and \( \xi_{\text{i}} , \) the PPL is
where the left term is a penalty complexity, and different \( \text{k} \) values put different stress on fit and parsimony. In Table 2, two values of \( \text{k} \) are used, \( \text{k} = 0.5 \) and \( \text{k} = 5 \), with the latter putting more stress on goodness of fit.
Also presented are predictive checks based on replicate observations. Posterior predictive probabilities \( \Pr (\text{y}_{\text{i},\,\text{rep}} > \text{y}_{\text{i}} |\text{y}) \) in extreme tails (e.g. values under 0.1 or over 0.9) indicate poorly fitted cases. The mixed predictive scheme (Marshall and Spiegelhalter 2003), providing checks that are close to leave-one-out cross validation (Green et al. 2009), was also applied. This involves sampling new random effects \( \text{s}_{\text{i},\,\text{rep}} \), and then sampling replicate data \( \text{y}_{\text{i},\,\text{rep},\,\text{mixed}} \) conditional on these new effects.
Table 2 shows that fit is generally improved under the heteroscedastic option, and predictive checks are also improved. The estimates for \( \uplambda \) suggest that spatial dependence is not overly pronounced, and hence illustrate the broader principle that a spatial prior represent unstructured as well as structured variation: estimates of \( \uplambda \) are all under 0.8. Figure 4 demonstrates disjunction between high risk and adjacent low risk areas. Table 2 also shows positive effects for both predictors but less precise effects under the scale mixture approach, in line with a general principle that neglecting heteroscedasticity may lead to mis-stated regression coefficient standard errors.
Table 3 contains a more detailed assessment of predictive discrepancies between the two approaches for the regression without predictors. As mentioned above, the \( \upkappa_{\text{i}} \) effects will act to identify spatial outliers, with illness levels discrepant from their neighbours, and so Table 3 contains the 20 areas with the lowest posterior mean \( \upkappa_{\text{i}} \) under the scale mixture approach. One may assess spatial outlier status to some extent from the observed data. The first two columns of Table 3 contain maximum likelihood (ML) relative risks in each area \( \text{R}_{\text{i}} = \text{y}_{\text{i}} /\text{E}_{\text{i}} \), and relative risks in the neighbourhoods \( \text{N}_{\text{i}} \) of each area, with ML estimates \( \text{L}_{\text{i}} = \) \( \sum\limits_{{\text{j} \in \text{N}_{\text{i}} }} {\text{y}_{\text{j}} } /\sum\limits_{{\text{j} \in \text{N}_{\text{i}} }} {\text{E}_{\text{j}} }. \)
Table 3 shows two types of outlier. One consists of major urban centres with high risk themselves, but a low risk hinterland (e.g. areas 1,2,3, and 8 in the Table). For example, Fig. 5 shows estimated relative risk patterns around area 2 (Leicester). These areas are underpredicted under the constant scale model, with mixed predictive \( \Pr (\text{y}_{\text{i},\,\text{rep},\,\text{mixed}} > \text{y}_{\text{i}} |\text{y}) \) p-values under 0.025. Under the scale mixture model they have higher means \( \upmu_{\text{i}} \), closer to the observed \( \text{y}_{\text{i}} \), as there is less local borrowing of strength. The other type of outlier (e.g. areas 5 and 6 in the Table) are low risk areas with much higher risk neighbourhoods. These are overpredicted under the constant scale model, with \( \Pr (\text{y}_{{\text{i}},{\text{rep}}} > \text{y}|\text{y}) = 0.91 \), and \( \Pr (\text{y}_{\text{i},\,\text{rep},\,\text{mixed}} > \text{y}_{\text{i}} |\text{y}) = 1 \) for area 6. Under the scale mixture model, modelled means are reduced closer to the observed \( \text{y}_{\text{i}} \). For all 20 areas, 19 have mixed predictive p-values under 0.05 or over 0.95 under a constant scale model, whereas under the scale mixture, this is reduced to 12 out of 20.
Table 4 contains the 10 areas with the lowest posterior mean \( \upkappa_{\text{i}} \) under the scale mixture approach when the two covariates are included. These areas illustrate when modelled relative risk in area \( \text{i} \) itself \( \upmu_{\text{i}} /\text{E}_{\text{i}} \) are discrepant from implied relative risk \( \sum\limits_{{\text{j} \in \text{N}_{\text{i}} }} {\upmu_{\text{j}} } /\sum\limits_{{\text{j} \in \text{N}_{\text{i}} }} {\text{E}_{\text{j}} } \) in the locality of area \( \text{i} \). These discrepancies may be related to covariate patterns. Under a scale mixture approach, local borrowing of strength is lessened, and Table 4 shows that the predicted TB counts \( \upmu_{\text{i}}\) are closer to the actual counts than under the constant scale LLB.
8 Conclusion
Different forms of spatial correlation analysis or modelling have been proposed in disease applications, ecological epidemiology, environmental science and other settings. Both Bayesian and frequentist estimation have been used. Common themes include identifying elevated risk areas or clusters of areas, and finding predictors of risk, while recognizing the explicitly spatial structure of the observations. For example, in a review of regression findings from spatial species abundance data, Dorfmann (2007) shows that ignoring spatial dependence (e.g. in regression residuals) leads to possible bias in parameter estimates and optimistic standard errors. However, while it is important to incorporate spatial dependence in models for area data, the assumptions of such techniques should be assessed, and subject to modification when the data so indicate. In particular, spatial discontinuities suggest a modification to the principle of uniform local smoothing.
In particular, Bayesian analyses of spatially arranged data often employ a conditionally autoregressive prior, which can express spatial clustering commonly present in the underlying risks, but typically assume a normal density for risks and uniform conditional association. However, a more sensitive parameterisation with utility in detecting outliers and locally irregular risk patterns may be obtained by allowing for non-normality. Commonly applied conditionally autoregressive priors, such as the proper CAR prior and the convolution prior, also have potential deficits when the observations contain a mixture of spatial dependence and unstructured heterogeneity. The present paper has proposed a scale mixture version of the Leroux et al. (1999) spatial prior, combining the benefit of adaptability when risks are discrepant in adjacent areas, and also a less problematic approach to representing a mixture of clustering and heterogeneity.
The analyses here show improved fit to infectious disease data, which may often show pronounced risk variability between areas. In England, high risk areas are often major urban centres, whereas the neighbouring suburban or rural hinterlands of such centres may be low risk. In such situations some modification of the uniform local borrowing of strength principle may be beneficial.
References
Alam M, Rönnegård L, Shen X (2015) Fitting conditional and simultaneous autoregressive spatial models in hglm. R J 7(2):5–18
Banerjee S, Carlin B, Gelfand A (2004) Hierarchical modeling and analysis for spatial data. Chapman & Hall/CRC, Boca Raton
Beale L, Abellan J, Hodgson S, Jarup L (2008) Methodologic issues and approaches to spatial epidemiology. Environ Health Perspect 116(8):1105–1110
Beale C, Lennon J, Yearsley J et al (2010) Regression analysis of spatial data. Ecol Lett 13(2):246–264
Berkhof J, van Mechelen I, Hoijtink H (2000) Posterior predictive checks: principles and discussion. Comput Stat 15(3):337–354
Besag J (1974) Spatial interaction and the statistical analysis of lattice systems. J R Statist Soc B 36:192–236
Besag J, Kooperberg C (1995) On conditional and intrinsic autoregressions. Biometrika 82:733–746
Besag J, York J, Mollié A (1991) Bayesian image restoration, with two applications in spatial statistics. Ann Inst Stat Math 43:1–21
Best N (1999) Bayesian ecological modelling. In: Lawson A, Biggeri A, Böhning D, Lesaffre E, Viel J-F, Bertollini R (eds) Disease mapping and risk assessment for public health. Wiley, New York, pp 194–201
Boris Choy S, Chan J (2008) Scale mixtures distributions in statistical modelling. Aust N Z J Stat 50(2):135–146
Brewer M, Nolan A (2007) Variable smoothing in Bayesian intrinsic autoregressions. Envirometrics 18:841–857
Brooks S, Gelman A (1998) General methods for monitoring convergence of iterative simulations. J Comput Gr Stat 7:434–445
Clayton D, Bernardinelli L, Montomoli C (1993) Spatial correlation in ecological analysis. Int J Epidemiol 22(6):1193–1202
Dormann C (2007) Effects of incorporating spatial autocorrelation into the analysis of species distribution data. Glob Ecol Biogeogr 16(2):129–138
Duarte-Cunha M (2015) Marcelo da Cunha G, Souza-Santos R (2015) Geographical heterogeneity in the analysis of factors associated with leprosy in an endemic area of Brazil: are we eliminating the disease? BMC Infect Dis 15:196
Farnsworth M, Ward M (2009) Identifying spatio-temporal patterns of transboundary disease spread: examples using avian influenza H5N1 outbreaks. Vet Res 40(3):1–14
Fernandez C, Steel M (1998) On Bayesian modeling of fat tails and skewness. J Am Statist Assoc 93:359–371
Gelfand A, Ghosh S (1998) Model choice: a minimum posterior predictive loss approach. Biometrika 85(1):1–11
Gelman A (1996) Bayesian model-building by pure thought: some principles and examples. Stat Sin 6(1):215–232
Gelman A, Carlin J, Stern H, Rubin D (2004) Bayesian Data Analysis, 2nd edn. Chapman & Hall/CRC Press, Boca Raton
Geweke K (1993) Bayesian treatment of the independent student linear model. J Appl Econom 8:19–40
Green M, Medley G, Browne W (2009) Use of posterior predictive assessments to evaluate model fit in multilevel logistic regression. Vet Res 40(4):1–10
Horabik J, Nahorski Z (2010) A statistical model for spatial inventory data: a case study of N2O emissions in municipalities of southern Norway. Clim Change 103(1–2):263–276
Lawson A, Clark A (2002) Spatial mixture relative risk models applied to disease mapping. Stat Med 21:359–370
Lee D (2011) A comparison of conditional autoregressive models used in Bayesian disease mapping. Spat Spat temp Epidemiol 2:79–89
Leroux B, Lei X, Breslow N (1999) Estimation of disease rates in small areas: a new mixed model for spatial dependence. In: Halloran M, Berry D (eds) Statistical models in epidemiology: the environment and clinical trials. Springer, New York, pp 135–178
LeSage J (1997) Bayesian estimation of spatial autoregressive models. Int Reg Sci Rev 20(1–2):113–129
Lopez De Fede A, Stewart J et al (2008) Tuberculosis in socio-economically deprived neighborhoods: missed opportunities for prevention. Int J Tuberc Lung Dis 12(12):1425–1430
Lunn D, Spiegelhalter D, Thomas A, Best N (2009) The BUGS project: evolution, critique and future directions. Stat Med 28(25):3049–3067
Maciel E, Pan W, Dietze R et al (2010) Spatial patterns of pulmonary tuberculosis incidence and their relationship to socio-economic status in Vitoria, Brazil. Int J Tuberc Lung Dis 14(11):1395–1402
MacNab Y (2014) On identification in Bayesian disease mapping and ecological-spatial regression models. Stat Method Med Res 23(2):134–155
Manda S (2013) Macro determinants of geographical variation in childhood survival in South Africa using flexible spatial mixture models. In: Kandala N-B, Ghilagaber G (eds) Advanced techniques for modelling maternal and child health in Africa. Springer, Dordrecht, pp 147–168
Marshall E, Spiegelhalter D (2003) Approximate cross validatory predictive checks in disease mapping. Stat Med 22:1649–1660
Morgenstern H (1995) Ecologic studies in epidemiology: concepts, principles, and methods. Annu Rev Public Health 16:61–81
Nathoo F, Ghosh P (2013) Skew-elliptical spatial random effect modeling for areal data with application to mapping health utilization rates. Stat Med 32(2):290–306
Nunes C (2007) Tuberculosis incidence in Portugal: spatiotemporal clustering. Int J Health Geogr 6:30
Pioz M, Guis H, Crespin L et al (2012) Why did bluetongue spread the way it did? Environmental factors influencing the velocity of bluetongue virus serotype 8 epizootic wave in France. PLoS One 7(8):e43360
Ploubidis G, Palmer M, Blackmore C et al (2012) Social determinants of tuberculosis in Europe: a prospective ecological study. Eur Respir J 40(4):925–930
Polson N, Scott J, Windle J (2014) The Bayesian bridge. J R Stat Soc B 76(4):713–733
Reich B, Hodges J (2008) Modeling longitudinal spatial periodontal data: a spatially adaptive model with tools for specifying priors and checking fit. Biometrics 64(3):790–799
Rieder H (1999) Epidemiologic basis of tuberculosis control. International Union Against Tuberculosis and Lung Disease, Paris
Rodrigues E, Assunção R (2012) Bayesian spatial models with a mixture neighborhood structure. J Multivar Anal 109:88–102
Shekhar S, Jiang Z, Ali R et al (2015) Spatiotemporal data mining: a computational perspective. ISPRS Int J Geoinform 4(4):2306–2338
Smith T, Wakefield J, Dobra A (2015) Restricted Covariance Priors with Applications in Spatial Statistics. Bayesian Anal 10(4):965–990
Ugarte M, Ibáñez B, Militino A (2005) Detection of spatial variation in risk when using CAR models for smoothing relative risks. Stoch Environ Res Risk Assess 19(1):33–40
Varga C, Pearl D, McEwen S et al (2015) Area-level global and local clustering of human Salmonella enteritidis infection rates in the city of Toronto, Canada, 2007–2009. BMC Infect Dis 15:359
Waller L, Carlin B (2010) Disease mapping. In: Gelfand A, Diggle P, Guttorp P, Fuentes M (eds) Handbook of spatial statistics. Chapman and Hall/CRC, Boca Raton, pp 217–243
West M (1984) Outlier models and prior distributions in Bayesian linear regression. J R Stat Soc B 46:431–439
Wikle C (2003) Hierarchical models in environmental science. Int Stat Rev 71(2):181–199
Rue H, Held L (2005) Gaussian Markov random fields: theory and applications. Chapman and Hall/CRC, Boca Raton
Yan J (2007) Spatial stochastic volatility for lattice data. J Agric Biol Environ Stat 12(1):25–40
Yi N, Xu S (2008) Bayesian LASSO for quantitative trait loci mapping. Genetics 179(2):1045–1055
Author information
Authors and Affiliations
Corresponding author
Appendix
Appendix
The R code for simulating data for 49 mainland US states is as follows:
# 49 by 49 binary adjacency matrix
W <- read.table(“adj_state.txt”)
# numbers of neighbours
d = c(4,5,6,3,7,3,3,2,2,5,6,5,4,6,4,7,3,1,5,5,3,4,4,8,4,6,5,3,3,5,5,4,3,5,6,4,6,2,2,6,8,4,6,3,6,2,5,4,6)
# expected events (TB incidence)
E <- c(147,199.5,89.6,1144.7,157.6,111.2,28.5,20.2,621.5,293.7,47.4,389.3,
197.3,94.5,86.5,133.4,138.4,42.4,180.1,207.7,302.4,164.4,89.1,184.3,31.4,56,83.8,41.3,272.8,62.7,606.1,297.6,22,354.4,115.3,121.5,399.6,
32.8,145.3,25.5,197.4,765.3,79.8,19.7,250.2,211.6,58.4,175.8,17.5)
# parameter and data definitions
N <- 49; Tau <- 3; lam = 0.7; T <- 100; nu <- 10; nu.2 <- nu/2
kap <- Qdiag <- numeric(N); var.s <- numeric(T)
y <- matrix(,N,T); Q <- C <- matrix(,N,N)
library(mvtnorm)
# simulation
for (t in 1:T) {for(i in 1:N) {# scale mixture effects
kap[i] <- rgamma(1,nu.2,nu.2);
Qdiag[i] <- Tau*kap[i]*(1-lam + lam*d[i])}
for(i in 1:N) {for (j in 1:N) {
Q[i,j] <- (i ==j)*Qdiag[i]-(1-(i ==j))*Tau*lam*W[i,j]*kap[i]*kap[j]}}
C <- solve(Q)
s <- rmvnorm(1, mean = rep(0, nrow(C)), sigma = C, method = c(“svd”))
eta <- log(E)-0.1 + s
mu <- exp(eta)
var.s[t] <- var(s[1:N])
for (i in 1:N){y[i,t] <- rpois(1,mu[i])}}
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Congdon, P. Representing spatial dependence and spatial discontinuity in ecological epidemiology: a scale mixture approach. Stoch Environ Res Risk Assess 31, 291–304 (2017). https://doi.org/10.1007/s00477-016-1292-9
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00477-016-1292-9