Abstract
Key message
The investigation provides initial knowledge on the distribution and evolution of the high and low methyl salicylate-producing trait in the Betula genus. Mislabelled birch species could be identified and removed.
Abstract
The genus Betula is the largest group of ecologically and economically dominant perennial woody plants in subalpine forests. The taxonomy of Betula is complex due to an extensive history of hybridization and periodic introgression events among the species. Although almost all land plants including birches produce methyl salicylate (MeSA) as a signaling molecule and in response to stress (“low MeSA producer”), some birch species produce high amounts of MeSA in the leaves and bark (“high MeSA producer”). Unfortunately, the evolution of high levels of MeSA production in the genus Betula remains unclear. The salicylic acid-binding protein 2 (SABP2) and salicylic acid methyltransferase (SAMT) genes involved in MeSA biosynthesis were incorporated into this study to examine the interspecific relationship of high and low MeSA-producing birches. Additionally, eight chloroplast and three nuclear regions were included to evaluate their potential application in species determination. The analysis resulted in 25 and 61 nucleotide variations, respectively, which allowed for a visualization of the genetic architecture in the 18 Betula species investigated. The high MeSA-producing B. lenta, B. grossa, and B. alleghaniensis formed the basal clade in the phylogenetic analysis, thus revealing their ancestral status, and the network analysis postulates that the diploid B. lenta is one of the ancestors of the genus Betula. The results indicate that the ability to produce high levels of MeSA that were initially present in the genus has been lost several times during its evolution. Placing species of the subgenus Acuminata alongside the subgenus Betula, together with a fragrance analysis, questions their ability to produce high levels of MeSA.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Birches (Betula L., Betulaceae) are ecologically important trees and shrubs that are widely distributed throughout the Northern Hemisphere (Furlow 1990). They diverged from other genera of the Betulaceae family around 75 million years ago (Bina et al. 2016). The genus Betula plays a vital role in landscape structure, forestry, breeding for biomass production and horticulture (Repola 2008; Ashburner and McAllister 2013; Smith et al. 2014), and some Betula species are listed as endangered in the International Union for Conservation of Nature’s (IUCN) red list of threatened species (Shaw et al. 2014).
Plant taxonomy helps in the monitoring and conservation of biodiversity (Hinchliff et al. 2015). However, hybridization, introgression, and misidentification of Betula species have led to difficulties in the systematics of this genus (Winkler 1904; Furlow 1990). Although birch species contain different biochemical compounds in their bark and buds (Williams and Arnold 2001) and vary in leaf shape and bark color (Li et al. 2005), these characteristics have not provided reliable interspecies relationships (Li et al. 2005; Schenk et al. 2008), and have resulted in several taxonomical classifications (Regel 1865; Winkler 1904; De Jong 1993; Skvortsov 2002). Regel (1865) acknowledged two subgenera: Eubetula consisting of six sections and Alnaster consisting of a single section, Acuminatae. Winkler (1904) classified the Betula genus extensively and further divided the genus into two main sections: Betulaster and Eubetula, with three subsections of Eubetula (Costatae, Nanae, and Albae). The same author suggested that the subsection Costatae required reconsideration (Winkler 1904). Ashburner and McAllister (2013) proposed the latest classification with four subgenera and eight sections, thereby substantially supporting the previous categorization (Skvortsov 2002).
Several studies have variously investigated the amplified fragment length polymorphism (AFLP) markers (Schenk et al. 2008), single genes (Li et al. 2007) and a combination of chloroplast barcoding and nuclear genes (Järvinen et al. 2004; Bina et al. 2016) to resolve the systematics of Betula. The official DNA barcoding regions rbcL and matK have been used to recognize different plant species due to their discrimination power, easy amplification, and good sequence quality (CBOL Plant Working Group 2009; Hollingsworth et al. 2011). However, the interspecific relationships of birch species remains unclear (Järvinen et al. 2004). The first (Li et al. 2005) and most recent (Wang et al. 2016) molecular phylogenetic analyses of the genus Betula, based on internal transcribed spacers (ITS), did not resolve all the ambiguities in the interspecific relationships of birches. These studies also advocated the inclusion of multiple nuclear and/or chloroplast regions to resolve these classification issues. The phylogenetic analysis by Wang et al. (2016), however, is congruent with the conclusions of Ashburner and McAllister (2013).
Apart from the classification debate, birches have an extensive history of medicinal use in the treatment of skin diseases, infections, rheumatism and urinary disorders (Menković et al. 2011; Shikov et al. 2011; Angmo et al. 2012; Rastogi et al. 2015). Analytical evidence suggests that terpenoids and methyl salicylate (MeSA) are the major components of birch essential oil (Can Baser et al. 2007; Pal et al. 2015). Terpenoids play a key role in plant defense (Singh and Sharma 2015) and their medicinal value is the subject of current clinical trials (Yin et al. 2017).
On the other hand, MeSA is a volatile compound produced by almost all land plants, including birches, as a signaling molecule and in response to stress (Forouhar et al. 2005) (termed here “low MeSA producers”). Some birch species produce high amounts of MeSA (“high MeSA producers”) in the leaves and bark and are used in many drugs for the treatment of muscle and joint pain (Williams and Arnold 2001; Dadákova et al. 2010). However, information about the content of the pharmacologically important MeSA in birch species is almost absent in the literature. Therefore, improved knowledge of the genetic framework of birches is required for selection and breeding purposes (Baum 2008).
Salicylic acid methyltransferase (SAMT) is a member of the O-methyltransferase gene family that catalyzes the methylation of salicylic acid (SA), while salicylic acid-binding protein 2 (SABP2) reverts MeSA to SA (Ross et al. 1999; Forouhar et al. 2005) by demethylation (Du and Klessig 1997). Therefore, SAMT and SABP2 are the putative vital genes for the variation in MeSA production. Thus, they are included in the study together with the internal transcribed spacer (ITS) (White et al. 1990), nitrate reductase (NIA) (Li et al. 2007) and alcohol dehydrogenase (ADH) (Järvinen et al. 2004) gene regions. Additionally, eight chloroplast coding and intergenic spacers, including rbcL, matK, trnH-psbA, psbK-psbI, trnQ-trnS, matK-trnK, trnC-petN, and rpoC2-rpoC1 (Schroeder et al. 2012; Schroeder and Fladung 2015), were also used to investigate their discrimination potential.
According to the monograph by Ashburner and McAllister (2013), B. alleghaniensis, B. alnoides, B. globispica, B. grossa, B. lenta, B. luminifera, B. maximowicziana and B. medwediewii are classified as high MeSA producers, whereas B. costata, B. davurica, B. ermani, B. nana, B. papyrifera, B. pendula, B. platyphylla, B. pubenscens, B. tianchanica and B. utilis are considered as low MeSA producers. Considering the uneven distribution of high MeSA content in the genus Betula, the purpose of the study was to uncover the interspecific relationships of high and low MeSA-producing birch species. Since data of the MeSA content in birches is only available in Ashburner and McAllister (2013), we applied an olfactory analysis by scratching and then smelling the bark to obtain an indication of high and low MeSA producers among birch species.
This study aims to answer the following questions: (1) Is the olfactory analysis of the MeSA fragrance consistent with the previous monographic report (Ashburner and McAllister 2013)? (2) Are the sequences of multiple chloroplasts and nuclear genomic regions capable of distinguishing between and within Betula groups of high and low MeSA productivity? (3) Are the phylogenies based on chloroplast and nuclear sequences compatible?
To this end, each chloroplast and the nuclear genomic region has been critically analyzed to detect nucleotide variations and to examine the discrimination potential. Although the taxonomy of Betula is complex, this investigation has helped in the design of further state-of-the-art experiments.
Materials and methods
Sample collection
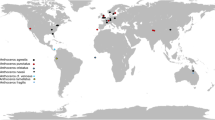
Seeds from eight Betula species (B. alleghaniensis, B. maximowicziana, B. medwediewii, B. grossa, B. costata, B. lenta, B. alnoides and B. globispica) were collected from different botanical gardens in Germany (Frankfurt, Giessen, Leipzig, Marburg and Tharandt), Great Britain (Liverpool) and Lithuania (Alytus). Samples of B. utilis were collected from a private supplier (Kiel, Germany). In addition, the Northwest German Forestry Research Institute provided frozen seeds of B. alleghaniensis (Table 1). Seeds were germinated in soil with pH 6.5 and 10–30% humidity in a natural environment without any fertilizer in a polyhouse at the Institute of Agricultural Process Engineering, Kiel University, Germany. Plantlet cultivation was carried out with the required dose of fertilizers in a glasshouse without any artificial light at the Thünen Institute of Forest Genetics, Grosshansdorf, Germany.
In addition, leaf or twig samples from nine more species (B. pubescens, B. pendula, B. papyrifera, B. luminifera, B. ermanii, B. platyphylla, B. nana, B. tianschanica, and B. davurica) were collected from botanical gardens in Canada (Montreal), Germany (Grosshansdorf, Göttingen and Munich) and Great Britain (Cambridge). The geographical distribution and place of origin are described in Table 1. Alnus incana and Carpinus betulus, as outgroup species, were sampled from BG Grosshansdorf.
Olfactory analysis of MeSA fragrance
To maintain consistency with previous studies, we conducted the olfactory analysis of MeSA fragrance by scratching the bark of young plants and twigs of the trees. Volunteers from the institute participated in the analysis. All volunteers were provided with bark from the young plants and twig samples from the different birches for scratching and sniffing. According to MeSA fragrance intensity, the samples were unanimously categorized as high, intermediate and low MeSA-producing Betula species (Supplementary Table 1).
Selection of chloroplast and nuclear regions and PCR primer design
Functions of the selected chloroplast and nuclear regions were defined according to previous reference studies (Supplementary Table 2). Previously described and newly designed primer combinations of all chloroplast and nuclear regions were used in the current study (Supplementary Table 2). Birch-specific primers were newly designed to amplify the SABP2, SAMT and NIA genes as well as the ITS regions with the help of the B. pendula genome (Salojärvi et al. 2017).
DNA extraction, PCR amplification and sequencing
Total DNA was extracted from the leaves of the plants (and the twigs of B. tianschanica, and B. davurica) according to the CTAB protocol of Dumolin et al. (1995). DNA extraction in B. nana, B. medwediewii, B. alleghaniensis, and B. lenta was difficult due to the presence of high levels of polysaccharides. Thus a pre-washing buffer (1.6 mL ice-cold TNE buffer: 200 mM Tris–HCl, 250 mM NaCl, 50 mM EDTA; Wang et al. 2013) was used to extract DNA of good quality.
The following PCR cycling conditions were used for all regions except ADH: 95 °C for 3 min, 40 cycles at 95 °C for 10 s, annealing with respective temperature for 30 s and 72 °C for 30 s. The amplification of ADH was carried out using the PCR reaction described by Järvinen et al. (2004). All PCR reactions were performed in a SensoQuest thermocycler (Göttingen, Germany). B. papyrifera, B. alnoides, B. globispica and B. pubescens individuals displayed two fragments after amplification with ADH primers. Therefore, gel purification of the ~ 1060 bp fragment of these species was carried out using a QIAquick Gel Extraction Kit. PCR products were confirmed on 1% agarose gel stained with Roti®-GelStain (Carl Roth, Karlsruhe, Germany). The StarSEQ (Mainz, Germany) service was used for sequencing.
Sequence examination and data validation
The electropherograms of each sequence were visually inspected. Upstream and downstream regions were trimmed. In addition, all sequences were aligned and screened for the presence of nucleotide substitutions using the SeqMan Pro15 program from the DNASTAR Lasergene bioinformatics software suite (Madison, Wisconsin USA). The sequences generated were validated with the NCBI gene database, if available. Verifying the reliability of the newly obtained sequences was an important step in avoiding any sequence misclassification.
Phylogenetic analysis
All sequences generated from the 29 birch individuals and the outgroup species were aligned with the bioinformatics computer program ClustalW and nexus files were created using MEGA7 (Kumar et al. 2016). Phylogenetic trees were inferred using ML (maximum likelihood) analysis with the substitution model JC69 + I for heterozygosity in the program PAUP* 4.0a (build 163) (https://paup.phylosolutions.com/). All characters, including indels and extended codes from the International Union of Pure and Applied Chemistry (IUPAC) (Johnson et al. 2010), were equally weighted and their status was kept unordered. Further, a bootstrap analysis of 100 replicates was performed to evaluate the individual clades with random and simple sequence additions (Maddison 1991). Generated trees were visualized using FigTree v1.4.3 (https://tree.bio.ed.ac.uk/software/figtree/) program. Furthermore, a phylogenetic network analysis was performed using the median-joining algorithm with the program package Network 5 (Bandelt et al. 1999).
Analyzed sequences of nuclear and chloroplast regions were treated as an individual-specific single-locus. Three different data sets (chloroplast, nuclear and a combined “league data set”) were created to evaluate the congruence between chloroplast and nuclear DNA. In addition, another phylogeny was constructed by deleting the possible ambiguous regions from the aligned league data set sequences (Supplementary Fig. 1). During alignment, sequences from fast-evolving genes and different species may produce potential gaps in the alignment defined as “ambiguous regions”. The presence of these regions may mean that the assumptions are inaccurate, while deleting them might result in the loss of vital information (Lutzoni et al. 2000).
The outgroup species, A. incana and C. betulus, were amplified with all the chloroplast primers. However, NIA and ADH primers were only able to amplify A. incana. Therefore, C. betulus was excluded from the nuclear phylogenetic analysis. Several heterozygous sites were observed in nuclear regions and treated according to the IUPAC nucleotide extended codes (https://www.bioinformatics.org/sms/iupac.html). The pairwise distance between Betula and the outgroup species was summarized using a function in MEGA7 based on the league data.
Results
Olfactory analysis of MeSA fragrance
To observe the consistency of high and low MeSA producers with monographic evidence (Ashburner and McAllister 2013), we conducted an olfactory analysis of MeSA fragrance by scratching the bark of young plants and twigs from the trees. The MeSA fragrance comparison analysis resulted in high (B. grossa, B. lenta and B. alleghaniensis), intermediate (B. maximowicziana, B. medwediewii) and low (all others) MeSA producers. Notably, the analysis showed that B. costata (subgenus Betula, section Costatae) produced the MeSA fragrance, while B. luminifera, B. alnoides (subgenus Acuminata, section Acuminatae) and B. globispica (subgenus Aspera, section Asperae) did not produce any MeSA that could unanimously be recognized by the human nose (Supplementary Table 1).
Sequence analysis
In the present study, 400 new Betula sequences were submitted to the NCBI GenBank database (Supplementary Table 3). Annotations and descriptions were assigned according to previous records available in the GenBank.
Polymorphic sites were found in chloroplast regions excluding the official barcode rbcL. The sequencing results indicated that the intergenic spacers trnH-psbA, psbK-psbI, trnC-petN and trnQ-trnS retained a higher degree of polymorphism than the coding regions matK, matK-trnK and rpoC2-rpoC1. The chloroplast intergenic spacer trnH-psbA had the highest number of substitutions, whereas the coding region rpoC2-rpoC1 showed an average number of nucleotide variations (Table 2).
The exon regions of the nuclear gene SABP2 (exon-3) and SAMT (exon-2) revealed a total number of 50 and 68 polymorphic sites, and 21 and 12 substitutions, respectively. The NIA and ADH sequences had a total number of 22 and 84 variable sites including six and 16 substitutions, respectively. In the ITS region, only 12 sites appeared to show variation (Table 3).
In addition, intraspecific sequence analysis of chloroplast regions was also conducted and this showed variations in B. maximowicziana, B. medwediewii, B. lenta and B. nana (Supplementary Table 4). Sequence alignments of B. alleghaniensis_10 sampled from the arboretum of the Thünen Institute, Grosshansdorf (BG), differed from the other B. alleghaniensis individuals. Therefore, we considered this individual as mislabelled and removed it from the study.
Phylogenetic analysis
Chloroplast regions
The aligned sequences of eight chloroplast regions of 29 individuals belonging to 18 birch species and two outgroup species contained 4587 characters, of which 204 were variable and 81 were parsimony-informative. A maximum likelihood phylogenetic tree was constructed (Fig. 1). According to our fragrance analysis (Supplementary Table 1), all species which produce low MeSA, except for B. maximowicziana and B. medwediewii, which produce an intermediate amount of MeSA, are clustered in clade I. Moreover, B. luminifera, B. alnoides and B. globispica, which are classified as high MeSA according to Ashburner and McAllister (2013) but which failed the olfactory analysis (Supplementary Table 1), are also clustered in clade I. On the other hand, the high MeSA producers are clustered together in clade II, except for B. costata, B. utilis and B. papyrifera.
Maximum likelihood (ML) phylogenetic tree based on eight cpDNA regions of different Betula species, using the substitution model JC69 + I for heterozygosity in the program PAUP* 4.0a. The Betula taxa with “h” produce a high amount of MeSA according to Ashburner and McAllister (2013), while those with “hh” produce a high amount of MeSA according to the olfactory fragrance analysis conducted in this study. The species with “hi” represent “h” high (according to Ashburner and McAllister (2013)) and “i” intermediate (according to the olfactory analysis, respectively)
Nuclear regions
The aligned sequences of five nuclear regions of 29 individuals belonging to 18 birch species and one outgroup species contained 2,818 characters, of which 377 were variable and 133 were parsimony-informative. B. papyrifera and B. utilis, which clustered with the high MeSA producers B. lenta, B. grossa and B. alleghaniensis in the chloroplast phylogeny (Fig. 1), congregated with most of the species from the subgenus Betula and Acuminata in clade I in the nuclear maximum likelihood phylogeny (Fig. 2). Therefore, clade II contains all the high MeSA producer species and only one low MeSA producer, B. costata.
Maximum likelihood (ML) phylogenetic tree based on five nuclear regions of different Betula species, using the substitution model JC69 + I for heterozygosity in the program PAUP* 4.0a. The Betula taxa with “h” produce a high amount of MeSA according to Ashburner and McAllister (2013), while those with “hh” produce a high amount of MeSA according to the olfactory fragrance analysis conducted in this study. The species with “hi” represent “h” high (according to Ashburner and McAllister (2013)) and “i” intermediate (according to the olfactory analysis, respectively)
League data set (combined chloroplast and nuclear sequence data)
The aligned chloroplast and nuclear sequences of all birch individuals were used to construct the concatenated data phylogeny (Fig. 3) which revealed a similar topology to the nuclear phylogeny (Fig. 2), with one exception: B. medwediewii displayed a discrete lineage (clade II). The pairwise distance between Betula and the outgroup species was summarized using a function in MEGA7 based on the league data (Supplementary Table 5).
Maximum likelihood (ML) phylogenetic tree based on the league data set of different Betula species, using the substitution model JC69 + I for heterozygosity in the program PAUP* 4.0a. The Betula taxa with “h” produce a high amount of MeSA according to Ashburner and McAllister (2013), while those with “hh” produce a high amount of MeSA according to the olfactory fragrance analysis conducted in this study. The species with “hi” represents, “h” high (according to Ashburner and McAllister (2013)) and “i” intermediate (according to the olfactory analysis, respectively)
Silencing of ambiguous regions
A phylogenetic tree was reconstructed by deleting the possible ambiguous regions in the league data set to examine the reliability of the generated data (Supplementary Fig. 1) and then compared with the league data set (Fig. 3). Constructing a phylogeny excluding any ambiguous regions did not result in any marked differences (Supplementary Fig. 1). Therefore, it can be concluded that alignment artifacts do not have any substantial effect on the existing phylogenetic analyses.
Network analysis
According to the requirements, only the variable sites of the league data set were used to create a FASTA file and perform the network analysis (Bandelt et al. 1999). The network revealed four clades (Fig. 4) similar to the nuclear phylogeny (Fig. 2). Clade I, II, III, and IV included both high and low MeSA-producing birches with complete heterozygosity between the birches. The network analysis clearly shows the ancestral node formed by B. lenta that carries a high MeSA-producing ability.
The evolutionary network analysis of the Betula individuals based on variable sites of eight cpDNA and five nuclear loci. Red points (mv) and numbers on the lineages refer to ancestral nodes and mutation events, respectively. The Betula taxa with “h” produce a high amount of MeSA according to Ashburner and McAllister (2013), while those with “hh” produce a high amount of MeSA according to the olfactory fragrance analysis conducted in this study. The species with “hi” represent “h” high (according to Ashburner and McAllister (2013)) and “i” intermediate (according to the olfactory analysis, respectively)
Discussion
The aim of the present study was to examine the differences in the ability of the multiple chloroplast and nuclear regions to determine the phylogenetic relationship between high and low MeSA-producing birches.
The relevance of the investigated markers
A total number of thirteen regions, including the chloroplast and nuclear regions were investigated for nucleotide variation in selected birches. Among the eight chloroplast regions, the intergenic spacer trnaH-psbA delivered the highest number of variable sites, which is consistent with previous studies (Bina et al. 2016; Schroeder et al. 2017). The often-recommended official barcoding region matK is more variable in genera like Lycium (Fukuda et al. 2001) and indicated low variation in the genus Betula, which is in agreement with previous studies (Järvinen et al. 2004; Cräutlein et al. 2011).
Although the official barcoding region rbcL varies in Populus (Schroeder et al. 2012; Schroeder and Fladung 2015) and complements the trnH-psbA sequences (Kress and Erickson 2007), no variants were found in the present study. This observation agrees with the previous analysis that the gene content of the chloroplast genomes is preserved in comparison with the nuclear genome, resulting in low variation (Kato et al. 2000; Järvinen et al. 2004; Palme et al. 2004; Tang et al. 2004). The current study suggests using the multi-locus chloroplast combinations trnH-psbA, matK-trnK, and trnC-petN in agreement with previous results (Fazekas et al. 2008; Hollingsworth et al. 2009; Levin et al. 2009) since the official barcoding and the coding regions of the chloroplast are highly conserved with less polymorphic sites.
In agreement with Järvinen et al. (2004), the low-copy nuclear ADH gene region displayed the highest number of variables in this study, most probably since the amplified ADH sequences include few exons and few introns. The universal ITS primers are also able to amplify fungal DNA and additional steps are needed to confirm the resulting sequences (Jobes et al. 1997). To bypass this step, Betula-specific ITS primers were designed (Supplementary Table 1) using B. pendula as the reference genome (Salojärvi et al. 2017). The new primers allowed for an easy amplification of the nuclear ribosomal ITS regions (ITS1 and ITS2). According to previous analysis (Li et al. 2007), NIA is a useful marker and helps in the inference of Betula phylogeny. However, NIA demonstrates low genetic variation. The reason could be that the new NIA PCR primers, designed on the first exon of the gene in the current study, might reduce the degree of polymorphism.
The SABP2 and SAMT gene regions are involved in the biogenesis of MeSA and serve as a signaling molecule in the systemic acquired resistance (SAR) mechanism of plants (Shah et al. 2014). Considering their organized interaction with numerous molecules, SABP2 and SAMT genes displayed a higher percentage of polymorphism (17.75% and 12.50%, respectively) in comparison with all analyzed regions. The current analysis indicates that slow-evolving uniparental chloroplast and fast-evolving bi-parental nuclear regions are more efficient in discrimination of the low and high MeSA birch producers than those gene sets studied independently.
Phylogenetic relationship of birches
The phylogenetic analysis of the genus Betula showed that the MeSA producing ability is not monophyletic but distributed among species from different subgenera. The phylogeny based on chloroplast regions (Fig. 1) revealed the relationship between three North American species, B. alleghaniensis, B. lenta, the high MeSA-producers and B. papyrifera located in one clade. Although we used eight different chloroplast regions, the results based on the birch species considered in Järvinen et al. (2004) and Bina et al. (2016) are consistent based on just the chloroplast either matK or trnH-psbA regions. However, B. grossa, B. costata and B. utilis were not studied in Järvinen et al. (2004) and B. grossa not in Bina et al. (2016). In our study, B. utilis is closely grouped with B. grossa, while it formed a separate lineage in the phylogeny based on trnH-psbA region (Bina et al. 2016).
In addition to the chloroplast regions, five fast-evolving nuclear regions were also used to generate robust data for phylogenetic analysis. In the nuclear phylogeny (Fig. 2), B. papyrifera and B. utilis formed a branch with the species from the subgenus Betula, which agrees with previous ITS phylogenies (Bina et al. 2016; Wang et al. 2016). Individuals in the nuclear phylogeny (Fig. 2) were more revealing when compared with the chloroplast region (Fig. 1), which might be due to a high number of variations in the nuclear regions.
The combined analysis (Fig. 3) was influenced by phylogenetic signals from nuclear sequences and chloroplast regions which contributed enough information to improve resolution within the clades. It could also explain why the league data set (Fig. 3) resembles that of the nuclear phylogeny (Fig. 2). It is possible that most variable sequences dominate the moderate data set in a concatenated analysis, thus resulting in the topology (Gontcharov et al. 2004). Species from the subgenus Aspera (B. alleghaniensis, B. grossa and B. lenta) formed a separate clade, whereas species from the subgenus Betula were congregated in another clade (Figs. 1, 2 and 3). Section-wise distribution of species from the subgenus Betula remains complex in all the data sets (Figs. 1, 2 and 3), which supports previous analysis (Li et al. 2005; Bina et al. 2016; Wang et al. 2016).
Due to the morphological similarities with B. lenta and B. alleghaniensis, the hexaploid B. grossa is placed in the Lentae section by Ashburner and McAllister (2013), agreeing with our phylogenetic analysis (Figs. 1, 2 and 3). However, this result disagrees with previous results (Nagamitsu et al. 2006; Schenk et al. 2008; Wang et al. 2016), where it is clustered with species from the subgenus Betula. The reason for this incongruity might be the multiple loci used and may also indicate that one of the parents of B. grossa belongs to the subgenus Betula (Wang et al. 2016). The possible hybridization with a morphologically similar species of section Lentae lead to the formation of an allopolyploid, resulting in it being thus placed (Ashburner and McAllister 2013; Wang et al. 2016).
Ashburner and McAllister (2013) placed B. globispica in a list of high MeSA producer Betula species that is incongruous with our olfactory analysis (Supplementary Table 1) where it produced no MeSA fragrance. Also, in the phylogenetic analysis (Figs. 1, 2, 3 and Supplementary Fig. 1), it clustered with species from the subgenus Betula, which is consistent with previous results (Bina et al. 2016). The reason for this might be that B. globispica hybridized with a species from the Betula section and thus appeared with the species from the Betula section (Wang et al. 2016).
According to the olfactory analysis (Supplementary Table 1), species from the subgenus Acuminata are either intermediate or low MeSA producers and always clustered with species from the subgenus Betula. They did not form a distinct clade in our analysis (Figs. 1, 2 and 3 and Supplementary Fig. 1). The close relationship between the diploid B. maximowicziana and subgenus Betula clearly supports the previous AFLP and ITS studies (Schenk et al. 2008; Wang et al. 2016). The olfactory analysis partially agrees with the morphological distribution (Ashburner and McAllister 2013) since the fragrance analysis could not detect MeSA in the diploid B. luminifera and tetraploid B. alnoides, suggesting further analytical examination. Additionally, it is experimentally proven that no post-zygotic barriers exist between B. maximowicziana and B. pendula subspecies mandshurica. Thus, fertile hybrids can be formed in crosses between two species (Johnsson 1945). Such evidence of hybridization could explain the intermediate or low production of MeSA in the subgenus Acuminata.
The decaploid B. medwediewii always clustered with different species of the subgenus Betula (Figs. 1, 2, 3 and Supplementary Fig. 1), in contrast to previous results where it clustered with B. lenta and B. alleghaniensis (Li et al. 2005; Bina et al. 2016; Wang et al. 2016). All three B. medwediewii accessions were from different botanical gardens but each displayed similar clustering, making misidentification unlikely. The unexpected placement of B. medwediewii in a clade with species of the subgenus Betula may indicate that one of the progenitors of this polyploid belongs to the subgenus Betula. This possibility is likely since in the league data set (Fig. 3), B. medwediewii occupied its own clade. In addition, it is an intermediate MeSA producer species (Supplementary Table 1) suggesting its authenticity as well as providing a valid reason for Ashburner and McAllister (2013) placing it in the subgenus Aspera.
The current analysis led to the identification of a misclassified B. alleghaniensis_10 individual from the arboretum of the Thünen Institute in Grosshansdorf, Germany (Supplementary Fig. 2). Both phylogenies (Fig. 3 and Supplementary Fig. 2) were constructed using the league data set. However, considerable differences were found, suggesting that the misclassification of a single individual could affect the complete phylogenetic tree. Such conflicting results and differing opinions are common scenarios in the genus Betula, and could be due to the continuous process of hybridization and introgression (Furlow 1990; Järvinen et al. 2004).
In addition, grouping of the diploid B. costata (subgenus, Betula; section, Costatae) with species from the section Lentae (Figs. 1, 2 and 3 and Supplementary Fig. 1) is consistent with former studies (Li et al. 2005; Bina et al. 2016). However, B. costata individuals used in the present study passed the olfactory analysis (Supplementary Table 1) producing considerable MeSA fragrance, thus contradicting the earlier report (Ashburner and McAllister 2013). However, the possibility of hybridization with one of the high MeSA-producing Betula species from East Asia and the problem of misclassification needs to be considered. Nevertheless, an analytical examination of more individuals will be required for stronger conclusions regarding the taxonomical and high MeSA-producing status of B. costata.
The use of SAMT and SABP2 was considered worthwhile for the phylogenetic analysis since these genes are involved in the biosynthesis of MeSA (Shah et al. 2014). As far as we know, this is the first study where SABP2 and SAMT are used in the phylogenetic analysis of Betula. The SAMT gene was previously investigated for the Solanaceae family (Talline et al. 2005) and provided advantages and disadvantages in the analysis. In the current analysis, although SAMT and SABP2 gene regions displayed higher variations (Table 3), they did not confirm a profound phylogenetic relationship between the birches.
An assessment of the sequencing data set generated is a vital operation in systematics for characterizing intrinsic features. The alignment of sequences generated from diverse genomic regions and species may produce ambiguous regions resulting in incongruent assumptions. In addition, if these regions are ignored, they may threaten the correct distribution of the species in the phylogeny (Lutzoni et al. 2000). Nevertheless, differences between the chloroplast and nuclear phylogenies have been reported in previous studies (Semerikov et al. 2003; Järvinen et al. 2004). This also occurred, to a certain extent, in the current analysis, suggesting that multigene analyses may not fully resolve all internal branches (Karol et al. 2001). Considering the disagreement between the data sets, a phylogenetic tree was constructed which ignored ambiguous regions in the alignment (Supplementary Fig. 1). In this case, the phylogeny displayed similar topologies to the nuclear phylogeny (Fig. 3) with better resolution through the branch length. It was concluded that alignment artifacts do not have any substantial effect on the phylogenetic analyses.
In the chloroplast (Fig. 1), nuclear (Fig. 2) and league data (Fig. 3) phylogenies, an exclusive clustering was observed between B. grossa, B. alleghaniensis and B. lenta, which produce high MeSA levels in the bark (Supplementary Table 1; Ashburner and McAllister 2013). The constant phylogenetic positioning of these species suggests their evolutionary relationships. It is possible that one of the species could be among the ancestors of selected Betula species. Although, the biogeographic reconstruction analysis by Bina et al. (2016) suggests that the genus Betula originated in East Asia. Bina et al. (2016) and De Jong (1993) proposed that the two subgenera of Betulenta (including B. grossa, B. alleghaniensis, and B. lenta) and Betulaster contain the oldest species, followed by the ancestral reconstruction that advocated the migration of the species from North America to Europe (Bina et al. 2016). Even the occurrence of disjunction events recognized within the genus Betula has been reported between eastern Asia and North America (B. costata and B. alleghaniensis) (Li et al. 2005). Following this disjunction, Wang et al. (2016) also hypothesized that the common ancestor of the Betula species might have been constantly scattered over the Northern Hemisphere. Bina et al. (2016) also support the occurrence of vicariance events in different regions, including the East Asian, North American and European lineages.
Considering parallel migration, hybridization and introgression events in the genus Betula and the network analysis (Fig. 4), we speculate that the diploid B. lenta, (subgenera; Betulenta) which has the ability to produce high MeSA, is one of the ancestors of the genus Betula. To construct the network analysis (Fig. 4), we utilized a substantial number of nuclear and chloroplast regions. Our network analysis (Fig. 4) results are consistent with previous studies where B. lenta also formed an ancestor node (Bina et al. 2016), along with the high MeSA producer B. uber (Ashburner and McAllister 2013).
We also speculate that in the process of evolution, the ancestrally high MeSA-producing trait was passed on to some younger Betula species, whereas some species lost their MeSA-producing ability, suggesting divergence in the evolutionary events among the species (Hughes 2012; Bina et al. 2016). Our analysis supports some of the previous studies; traits which are not maintained through natural selection may become less functional over time and follow a relaxed evolutionary selection (Reich et al. 2003). If some traits have no active function in the genome, they could disappear in future generations (Lahti et al. 2009).
Conclusions
The present study provides initial knowledge on the distribution of the high and low MeSA-producing trait in the Betula genera. The network analysis suggests that B. lenta is one of the ancestors and that the MeSA-producing trait has been lost several times in the evolution of younger Betula species. The chloroplast and nuclear phylogenies reveal homogenous topologies suggesting compatibility between organelle and nuclear results. This is noteworthy, considering that even a small segment of a nuclear DNA sequence produces high-resolution levels and provides more parsimony information to the chloroplast DNA. The data generated could be important for the Barcode of Life project and will be used to develop rapid molecular test methods to reduce the cost of sequencing for the breeding and selection of birches.
Comprehensive sampling together with genome-wide studies could resolve the questionable phylogenetic position of some Betula species. Additionally, an analytical examination can help to make precise decisions about the taxonomical positions of high and low MeSA-producing Betula species.
Future perspectives
The study successfully provides strong evidence about the evolutionary state of the ancestral species to establish a visible and reliable basis for further next-generation sequencing experiments. The SABP2 and SAMT genes serve as candidate genes for MeSA biosynthesis. The original idea to include these genes in the phylogenetic analysis was to correlate putative nucleotide substitutions with MeSA production. On the basis of the current study, different low and high MeSA-producing Betula species will be selected for bark and leaf RNA sequencing analyses.
References
Angmo K, Adhikari BS, Rawat GS (2012) Changing aspects of traditional healthcare system in western Ladakh, India. J Ethnopharmacol. https://doi.org/10.1016/j.jep.2012.07.017
Ashburner K, McAllister HA (2013) The genus Betula: a taxonomic revision of birches. Kew Publishing, London
Bandelt HJ, Peter F, Arne R (1999) Median-joining networks for inferring intraspecific phylogenies. Mol Biol Evol. https://doi.org/10.1093/oxfordjournals.molbev.a026036
Baser KHC, Demirci B (2007) Studies on Betula essential oils. ARKIVOC. https://doi.org/10.3998/ark.5550190.0008.730
Baum D (2008) Trait evolution on a phylogenetic tree: relatedness, similarity, and the myth of evolutionary advancement; Nature Education. https://www.nature.com/scitable/topicpage/trait-evolution-on-a-phylogenetic-tree-relatedness-41936/. Accessed 15 Jun 2019
Bina H, Yousefzadeh H, Ali SS et al (2016) Phylogenetic relationships, molecular taxonomy, biogeography of Betula, with emphasis on phylogenetic position of Iranian populations. Tree Genet Genomes. https://doi.org/10.1007/s11295-016-1037-4
CBOL Plant Working Group, Hollingsworth PM, Forrest L et al (2009) DNA barcode for land plants. Proc Natl Acad Sci. https://doi.org/10.1073/pnas.0905845106
Cräutlein M, Korpelainen H, Pietiläinen M (2011) DNA barcoding: a tool for improved taxon identification and detection of species diversity. Biodivers Conserv. https://doi.org/10.1007/s10531-010-9964-0
Dadákova E, Vrchotová N, Tříska J (2010) Content of selected biologically active compounds in tea infusions of widely used European medicinal plants. J Agrobiol. https://doi.org/10.2478/s10146-009-0003-2
De Jong PC (1993) An introduction to Betula: Its morphology, evolution, classification and distribution with a survey of recent work, Proceedings of the IDS Betula symposium, 2–4 October 1992. International Dendrology Society, Richmond
Du H, Klessig DF (1997) Identification of a soluble, high-affinity salicylic acid-binding protein in Tobacco. Plant Physiol. https://doi.org/10.1104/pp.113.4.1319
Dumolin S, Demesure B, Petit RJ (1995) Inheritance of chloroplast and mitochondrial genomes in pedunculate oak investigated with an efficient PCR method. Theor Appl Genet. https://doi.org/10.1007/BF00220937
Fazekas AJ, Burgess KS, Kesanakurti PR, Graham SW et al (2008) Multiple multilocus DNA barcodes from the plastid genome discriminate plant species equally well. PLoS ONE. https://doi.org/10.1371/journal.pone.0002802
Ferris C, King RA, Gray AJ (1997) Molecular evidence for the maternal parentage in the hybrid origin of Spartina anglica C.E. Hubbard. Mol Ecol. https://doi.org/10.1046/j.1365-294X.1997.00165.x
Forouhar F, Yang Y, Kumar D, Chen Y et al (2005) Structural and biochemical studies identify tobacco SABP2 as a methyl salicylate esterase and implicate it in plant innate immunity. PNAS. https://doi.org/10.1073/pnas.0409227102
Fukuda T, Yokoyama J, Ohashi H (2001) Phylogeny and biogeography of the genus Lycium (Solanaceae): inferences from chloroplast DNA sequences. Mol Phylogenet Evol. https://doi.org/10.1006/mpev.2001.0921
Furlow JJ (1990) The genera of Betulaceae in the southeastern United States. J Arnold Arboretum 71:1–67. https://biostor.org/reference/61852
Gontcharov AA, Marin B, Melkonian M (2004) Are combined analyses better than single gene phylogenies? A case study using SSU rDNA and rbcL sequence comparisons in the Zygnematophyceae (Streptophyta). Mol Biol Evol. https://doi.org/10.1093/molbev/msh052
Hinchliff CE, Smith SA, Allman JF, Burleigh JG et al (2015) Synthesis of phylogeny and taxonomy into a comprehensive tree of life. Proc Natl Acad Sci USA. https://doi.org/10.1073/pnas.1423041112
Hollingsworth ML, Andra CA, Forrest LL et al (2009) Selecting barcoding loci for plants: evaluation of seven candidate loci with species-level sampling in three divergent groups of land plants. Mol Ecol Resour. https://doi.org/10.1111/j.1755-0998.2008.02439.x
Hollingsworth PM, Graham SW, Little DP (2011) Choosing and using a plant DNA barcode. PLoS ONE. https://doi.org/10.1371/journal.pone.0019254
Hughes AL (2012) Evolution of adaptive phenotypic traits without positive Darwinian selection. Heredity. https://doi.org/10.1038/hdy.2011.97
Järvinen P, Palmé A, Morales LO et al (2004) Phylogenetic relationships of Betula species (Betulaceae) based on nuclear ADH and chloroplast matK sequences. Am J Bot. https://doi.org/10.3732/ajb.91.11.1834
Jobes DV, Thien LB (1997) A conserved Motif in the 5.8S Ribosomal RNA (rRNA) gene is a useful diagnostic marker for plant internal transcribed spacer (ITS) sequences. Mol Biol Report. https://doi.org/10.1023/A:1007462330699
Johnson AD (2010) An extended IUPAC nomenclature code for polymorphic nucleic acids. Bioinformatics. https://doi.org/10.1093/bioinformatics/btq098
Johnsson H (1945) Interspecific Hybridization within the genus Betula. Hereditas. https://doi.org/10.1111/j.1601-5223.1945.tb02752.x
Karol KG, McCourt RM, Cimino MT, Delwiche CF (2001) The closest living relatives of land plants. Science. https://doi.org/10.1126/science.1065156
Kato T, Kaneko T, Sato S, Nakamura Y, Tabata S (2000) Complete structure of the chloroplast genome of a Legume, Lotus japonicus. DNA Res 5:6. https://doi.org/10.1093/dnares/7.6.323
Kress WJ, Erickson DL (2007) A two-locus global DNA barcode for land plants: the coding rbcL gene complements the non-coding trnH-psbA spacer region. PLoS ONE. https://doi.org/10.1371/journal.pone.0000508
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for Bigger Datasets. Mol Biol Evol. https://doi.org/10.1093/molbev/msw054
Lahti DC, Johnson NA, Ajie BC, Otto SP et al (2009) Relaxed selection in the wild. Trends Ecol Evol. https://doi.org/10.1016/j.tree.2009.03.010
Levin RA, Blanton J, Miller JS (2009) Phylogenetic utility of nuclear nitrate reductase: a multi-locus comparison of nuclear and chloroplast sequence data for inference of relationships among American Lycieae (Solanaceae). Mol Phylogenet Evol. https://doi.org/10.1016/j.ympev.2008.12.005
Li JH, Shoup S, Chen ZD (2005) Phylogenetics of Betula (Betulaceae) inferred from sequences of nuclear ribosomal DNA. Rhodora. https://doi.org/10.3119/04-14.1
Li JH, Shoup S, Chen ZD (2007) Phylogenetic relationships of diploid species of Betula (Betulaceae) inferred from DNA sequences of nuclear nitrate reductase. Syst Bot. https://doi.org/10.1600/036364407781179699
Lutzoni F, Wagner P, Reeb V, Zoller S (2000) Integrating ambiguously aligned regions of DNA sequences in phylogenetic analyses without violating positional homology. Syst Biol. https://doi.org/10.1080/106351500750049743
Maddison D (1991) The discovery and importance of multiple Islands of most-parsimonious trees. Syst Biol. https://doi.org/10.1093/sysbio/40.3.315
Martins TR, Barkman TJ (2005) Reconstruction of solanaceae phylogeny using the nuclear gene SAMT. Syst Bot. https://doi.org/10.1600/0363644054223675
Menković N, Savikin K, Tasić S, Zdunić G, Stesević D et al (2011) Ethnobotanical study on traditional uses of wild medicinal plants in Prokletije Mountains (Montenegro). J Ethnopharmacol. https://doi.org/10.1016/j.jep.2010.09.008
Nagamitsu T, Kawahara T, Kanazashi A (2006) Endemic dwarf birch Betula apoiensis (Betulaceae) is a hybrid that originated from Betula ermanii and Betula ovalifolia. Plant Species Biol. https://doi.org/10.1111/j.1442-1984.2006.00147.x
Pal M, Mishra T, Kumar A et al (2015) Chemical constituents and antimicrobial potential of essential oil from Betula utilis growing in high altitude of Himalaya (India). J Essent Oil Bear Plants. https://doi.org/10.1080/0972060X.2015.1036569
Palme AE, Su Q, Palsson S, Lascoux M (2004) Extensive sharing of chloroplast haplotypes among European birches indicates hybridization among Betula pendula B. pubescens and B. nana. Mol Ecol. https://doi.org/10.1046/j.1365-294X.2003.02034.x
Rastogi S, Pandey MM, Kumar Singh Rawat A (2015) Medicinal plants of the genus Betula-traditional uses and a phytochemical-pharmacological review. J Ethnopharmacol. https://doi.org/10.1016/j.jep.2014.11.010
Regel E (1865) Bemerkungen uber die Gattungen Betula und Alnus nebst Beschreibung einiger neuer Arten. Bull Soc Nat (Moscou) 38:388–434
Reich PB, Wright IJ, Cavender-Bares JM, Craine J et al (2003) The evolution of plant functional variation: traits, spectra, and strategies. Int J Plant Sci. https://doi.org/10.1086/374368
Repola J (2008) Biomass equations for birch in Finland. Silva Fennica. https://doi.org/10.14214/sf.236
Ross JR, Nam KH, D'Auria JC, Pichersky E (1999) S-Adenosyl-l-methionine: salicylic acid carboxyl methyltransferase, an enzyme involved in floral scent production and plant defense, represents a new class of plant methyltransferases1. Arch Biochem Biophys. https://doi.org/10.1006/abbi.1999.1255
Salojärvi J, Smolander O, Nieminen K, Rajaraman S, Safronov O et al (2017) Genome sequencing and population genomic analyses provide insights into the adaptive landscape of silver birch. Nat Genet. https://doi.org/10.1038/ng.3862
Schenk MF, Thienpont CN, Koopman WJM et al (2008) Phylogenetic relationships in Betula (Betulaceae) based on AFLP markers. Tree Genet Genom. https://doi.org/10.1007/s11295-008-0162-0
Schroeder H, Hoeltken AM, Fladung M (2012) Differentiation of Populus species using chloroplast single nucleotide polymorphism (SNP) markers-essential for comprehensible and reliable poplar breeding. Plant Biol. https://doi.org/10.1111/j.1438-8677.2011.00502.x
Schroeder H, Fladung M (2015) Differentiation of Populus species by chloroplast SNP markers for barcoding and breeding approaches. iForest Biogeosci Forestry. https://doi.org/10.3832/ifor1326-007
Schroeder H, Kersten B, Fladung M (2017) Development of multiplexed marker sets to identify the most relevant poplar species for breeding. Forests. https://doi.org/10.3390/f8120492
Semerikov VL, Yhang H, Sun M, Lascoux M (2003) Conflicting phylogenies of Larix (Pinaceae) based on cytoplasmic and nuclear DNA. Mol Phylogenet Evol. https://doi.org/10.1016/S1055-7903(02)00447-5
Shah J, Chaturvedi R, Chowdhury Z, Venables B, Petros RA (2014) Signaling by small metabolites in systemic acquired resistance. Plant J Cell Mol Biol. https://doi.org/10.1111/tpj.12464
Shaw K, Stritch L, Rivers M, Roy S, Becky W, Govaerts R (2014) The Red List of Betulaceae. BGCI. Richmond. https://www.bgci.org/resources/bgci-tools-and-resources/the-red-list-of-betulaceae/. Accessed 15 Oct 2019
Shikov AN, Djachuk GI, Sergeev DV et al (2011) Birch bark extract as therapy for chronic hepatitis C-a pilot study. Phytomedicine. https://doi.org/10.1016/j.phymed.2011.01.021
Singh B, Sharma RA (2015) Plant terpenes: defense responses, phylogenetic analysis, regulation and clinical applications. 3 Biotech. https://doi.org/10.1007/s13205-014-0220-2
Skvortsov AK (2002) A new system of the genus Betula L.-the birch. Bull Mosc Soc Nat 107:73–76
Small RL (2004) Phylogeny of Hibiscus sect. Muenchhusia (Malvaceae) Based on chloroplast rpL16 and ndhF, and nuclear ITS and GBSSI Sequences. Syst Bot. https://doi.org/10.1600/036364404774195575
Small RL, Cronn RC, Wendel JF (2004) Use of nuclear genes for phylogeny reconstruction in plants. Aust Syst Bot. https://doi.org/10.1071/sb03015
Smith A, Granhus A, Astrup R et al (2014) Functions for estimating above ground biomass of birch in Norway. Scand J For Res. https://doi.org/10.1080/02827581.2014.951389
Tang J, Xia H, Cao M et al (2004) A comparison of rice chloroplast genomes. Plant Physiol. https://doi.org/10.1104/pp.103.031245
Wang N, McAllister HA, Bartlett PR et al (2016) Molecular phylogeny and genome size evolution of the genus Betula (Betulaceae). Ann Bot. https://doi.org/10.1093/aob/mcw048
Wang N, Thomson M, Bodles WJ et al (2013) Genome sequence of dwarf birch (Betula nana) and cross-species RAD markers. Mol Ecol. https://doi.org/10.1111/mec.12131
White TJ, Bruns T, Lee S, Taylor T (1990) Amplification and direct sequencing of fungal ribosomal RNA Genes for phylogenetics. PCR protocols: a guide to methods and applications. Academic Press, New York, pp 315–322
Williams J Jr, Arnold M (2001) Sources of genetic structure in the woody perennial Betula occidentalis. Int J Plant Sci. https://doi.org/10.1086/321921
Winkler H (1904) Betulaceae. In: Engler A (ed) Das Pflanzenreich Heft, 19 (IV.61), 1–49. W. Engelmann, Leipzig
Yin J, Li X, Zhan Y et al (2017) Cloning and expression of BpMYC4 and BpbHLH9 genes and the role of BpbHLH9 in triterpenoid synthesis in birch. BMC Plant Biol. https://doi.org/10.1186/s12870-017-1150-z
Acknowledgements
Open Access funding provided by Projekt DEAL. We are highly grateful to the botanical gardens and Northwest German Forestry Research Institute (listed in Table 1) for providing Betula seed samples upon our request for scientific research purposes only without any conflict of interest. We thank PD Dr. Birgit Kersten for assisting with the submission of data to the NCBI GenBank, to Dr. Niels Müller for consultation in sequencing and the manuscript review (both Thünen Institute of Forest Genetics, Grosshansdorf, Germany), and to all colleagues at the Thünen Institute of Forest Genetics, Grosshansdorf, Germany and the Institut für Landwirtschaftliche Verfahrenstechnik Kiel for their support, including Katrin Groppe for laboratory assistance, and Dr. Hilke Schröder, Dr. Jutta Buschbom and Khira Deecke for helpful discussions on phylogenic analyses. Thanks are due to Dina Führmann (Thünen Institute, Braunschweig, Germany) and Nicola Wilton (Language Boutique®, Hamburg, Germany) for English language editing.
Funding
This project was funded by the “Energiewende und Umweltinnovationen” state program for the economy; the Ministry of Energy Transition, Agriculture, Environment and Rural Areas of Schleswig–Holstein, Germany.
Author information
Authors and Affiliations
Contributions
MF is the first supervisor of this work and contributed to the experimental outline, data validation and correcting of the manuscript. Project leader CRM is the second supervisor and contributed to sample collection and writing the project proposal. EH provided guidance throughout the experiments and was also responsible for English language editing. Lab work, writing of the paper, data collection and analysis was completed by KS as a part of his Ph.D. work.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by J. Hormaza.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Singewar, K., Moschner, C.R., Hartung, E. et al. Species determination and phylogenetic relationships of the genus Betula inferred from multiple chloroplast and nuclear regions reveal the high methyl salicylate-producing ability of the ancestor. Trees 34, 1131–1146 (2020). https://doi.org/10.1007/s00468-020-01984-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00468-020-01984-x