Abstract
Background
Personalized medicine requires the integration and analysis of vast amounts of patient data to realize individualized care. With Surgomics, we aim to facilitate personalized therapy recommendations in surgery by integration of intraoperative surgical data and their analysis with machine learning methods to leverage the potential of this data in analogy to Radiomics and Genomics.
Methods
We defined Surgomics as the entirety of surgomic features that are process characteristics of a surgical procedure automatically derived from multimodal intraoperative data to quantify processes in the operating room. In a multidisciplinary team we discussed potential data sources like endoscopic videos, vital sign monitoring, medical devices and instruments and respective surgomic features. Subsequently, an online questionnaire was sent to experts from surgery and (computer) science at multiple centers for rating the features’ clinical relevance and technical feasibility.
Results
In total, 52 surgomic features were identified and assigned to eight feature categories. Based on the expert survey (n = 66 participants) the feature category with the highest clinical relevance as rated by surgeons was “surgical skill and quality of performance” for morbidity and mortality (9.0 ± 1.3 on a numerical rating scale from 1 to 10) as well as for long-term (oncological) outcome (8.2 ± 1.8). The feature category with the highest feasibility to be automatically extracted as rated by (computer) scientists was “Instrument” (8.5 ± 1.7). Among the surgomic features ranked as most relevant in their respective category were “intraoperative adverse events”, “action performed with instruments”, “vital sign monitoring”, and “difficulty of surgery”.
Conclusion
Surgomics is a promising concept for the analysis of intraoperative data. Surgomics may be used together with preoperative features from clinical data and Radiomics to predict postoperative morbidity, mortality and long-term outcome, as well as to provide tailored feedback for surgeons.
Graphical abstract

Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Surgery is an important building block of the treatment for a multitude of diseases, for example in multivessel coronary artery disease [1] or solid cancer [2]. At the same time, surgery is dangerous with postoperative death being the third leading cause of death worldwide [3]. The complexity of surgical treatment results from a multitude of interactions between patients, clinical staff members and medical devices during pre-, intra- and postoperative phases. Optimal surgical results that can theoretically be achieved are not always reached in daily practice. Especially during the postoperative treatment pathway, complications like erosive bleeding from a pancreatic fistula [4] or mediastinitis after an esophageal anastomotic leakage [5] can occur. These complications result in a significant deterioration of the patient’s outcome or even in the death of the patient. Patient-associated factors such as age, sex, body mass index (BMI) or comorbidities such as cardiac disease or diabetes can have a large impact on the postoperative outcome and may therefore be used for outcome prediction [6, 7]. But also surgeon and hospital experience [8], as well as an adequate treatment of major postoperative surgical complications are influencing the mortality rates in different perioperative care systems [9]. With regard to the intraoperative setting, increasing evidence suggests that specific parameters and events are strongly associated with postoperative complications and outcome [10]. For example, intraoperative factors like a staple line bleeding in laparoscopic distal pancreatectomy can be a potential precursor of a later pancreatic fistula, as demonstrated using video review-based analysis [11]. Furthermore, it has been proven that surgical skill and technical performance are strongly associated with postoperative clinical outcome [12,13,14]. Nevertheless, the predictive accuracy of surgeons for postoperative complications appears limited, e.g. in case of anastomotic leakage in gastrointestinal surgery [15].
In non-surgical areas of multimodal treatment especially for cancer, personalized therapy recommendations along the treatment pathway have already been pursued with the approaches of Genomics, Epigenomics, Transcriptomics, and Microbiomics in (tumor) biology [16], and with Radiomics in clinical medicine [17]. However, an omics-approach in surgery that uses intraoperative data comprehensively, i.e. combines surgical data specific to the operating room to create a quantitative description of a surgical procedure is not yet established. This deficiency may result from the large scale and velocity of multimodal data in the operating room produced by heterogeneous sources of raw data, or might also be related to the high level of variability of surgical processes [18]. To pave the way for a personalized therapy, which makes use of intraoperative data, relevant surgical process characteristics need to be extracted from data sources like endoscopic videos, intraoperative imaging methods or vital sign monitoring using machine learning (ML) methods [18]. First steps in this endeavor have been made in the preoperative setting using ML to automatically identify high risk surgical patients [19]. Intraoperatively, ML has been widely studied for the recognition of surgical phases [20, 21], for the automatic assessment of the surgeon’s skill level [22], or for predicting postoperative adverse events and distractions [23]. With the OR black box, Jung et al. presented a first comprehensive approach of quantitatively identifying intraoperative events and distractions by collecting intraoperative data [24]. Based on this work, they developed an index to measure the severity of intraoperative events and thus identified patients at high risk of developing postoperative complications [25]. However, their analysis was based on human expert ratings and not on a computer-based approach, thus requiring a huge effort and time limiting its scalability. Up until now ML has not been applied within a comprehensive and concrete approach to use intraoperative surgical data to support surgical decision-making leading to a lack of success stories in the field of surgical data science [26]. We argue that the absence of a useful and classified representation of recorded intraoperative data, which would allow for the analysis of vast amounts of heterogeneous data is one major reason in this regard.
The aim of this article is to approach this shortcoming by defining the concept of Surgomics as a surgical answer to Genomics and Radiomics. After identifying and categorizing intraoperative surgomic features, this article investigates which surgomic features are perceived as clinically most relevant and technically most feasible by surgeons and (computer) scientists with a focus on the example of minimally invasive oncological surgery. In contrast to the general concept of surgical data science, Surgomics will focus on the intraoperative setting to use and integrate the rapidly increasing amounts of intraoperative sensor data. Surgomics may be used together with pre- and postoperative data to improve surgical treatment with surgical data science methods. Furthermore, Surgomics could enable digital surgery applications such as decision support systems, computer-assisted training, and robotics [27].
Materials and methods
We started with the idea to systematically explore intraoperative data and use methods of feature extraction from those vast amounts of data similar to Radiomics. In order to elaborate this raw idea we brought together experts from multiple surgical and scientific disciplines and came up with a definition “Surgomics”, i.e. its data sources, methods and applications. Then, we discussed ideas for surgomic features, categorized them, and used what we found to eliminate uncertainties in the definition of Surgomics. Finally, we validated the ideas for features and their relevance in a multicenter expert survey.
Definition of Surgomics
We define Surgomics as follows: Surgomics is the entirety of surgomic features that are process characteristics of a surgical procedure automatically derived from multimodal intraoperative data to quantify processes in the operating room. Surgomics is thus the surgical answer to Genomics and Radiomics and may be used together with preoperative features from clinical data science and Radiomics to predict postoperative morbidity, mortality and long-term outcome as well as to provide tailored feedback for surgeons.
Based on previous work in surgical data science [18], surgical process models [28], surgical workflow analysis [29], and health information networks [30], we aim to establish Surgomics as a concept to collect, process and structure multimodal data in the intraoperative setting (Fig. 1). To reach this goal, surgical expert knowledge needs to be combined with ML and data science methods to make effective and scalable use of the vast amounts of intraoperative data.
Concept of Surgomics. a In surgical data science, pre-, intra- and postoperative data are integrated to predict morbidity, mortality and long-term outcome. b Surgomics focuses on the intraoperative setting that comprises data sources like the surgical video or anesthesiological vital sign monitoring. c Surgomic features can be extracted from suitable data sources in an automated fashion, for example using machine learning or other data science methods
Categorization of surgomic features
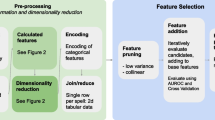
The aforementioned definition of Surgomics and surgomic features resulted as a first step in development by a multidisciplinary team (n = 19) consisting of experts from the fields of colorectal, pancreatic, upper gastrointestinal, robot-assisted and minimally invasive surgery on one hand, as well as translational surgical oncology, surgical data science, connected medical devices, medical informatics, medical data models, clinical decision support, radiomics, and endoscopic vision on the other hand. In a next step, the multidisciplinary team discussed the variety of data available in the intraoperative setting and their potential relevance for the patient's short-term surgical outcome (morbidity and mortality), as well as long-term outcome (survival). In the course of the discussion, examples of surgomic features were collected and categorized. Here, we aimed for a broad spectrum of features using different intraoperative data sources to foster future discussion on new features. Later on, additional information for each feature was added, i.e. unit of measurement, source of raw data, specificity for a certain surgical procedure and literature references if applicable (Table 1). Regarding the categorization of the surgomic features, the multidisciplinary team agreed on the following eight feature categories (Fig. 2):
-
1.
Surgical field
The surgical field category contains surgomic features which can be derived from the surgical video (for example in laparoscopy or thoracoscopy) and directly apply to the surgical procedure, for example the level of blood and smoke in the surgical field or imaging methods like the accumulation of indocyanine green (ICG) as a measure for the perfusion in an anastomosis.
-
2.
Instrument
The instrument category comprises features related to surgical instruments, e.g. the instrument location in the surgical video, the action performed with an instrument or medical device, device settings such as voltage in electrosurgery, or vacuum and pressure settings for suction and irrigation. The source of raw data can be the (endoscopic) video, but data can also be derived directly from the medical device sensor connected to an instrument.
-
3.
Temporal course of procedure
The category relating to the temporal course of procedure contains characteristics, which describe the surgery with respect to its workflow, e.g. the duration and order of performed surgical steps.
-
4.
Patient
The patient category represents patient-specific characteristics such as the visual texture of lung or liver derived from surgical video, or the pancreatic duct diameter. These features are derived directly from the patient and may also be available preoperatively, e.g. from computed tomography.
-
5.
Anesthesiological data
Examples for features in the anesthesiological data category are data of the blood gas analysis or the vital sign monitoring. The features can be derived from the (electronic or digitized) anesthesia protocol or from anesthesiological devices.
-
6.
Surgical skill and quality of performance
The category referring to the surgical skill and quality of performance contains features, which are specific to each surgeon. For instance, surgical skill can be objectified with the GOALS score [31] including features such as bimanual dexterity or tissue handling. Other features occurring in the course of surgery, directly depending on the operating surgeon, are the quality of a suture or the extent of dissection of lymphatic tissue in oncological surgery [32].
-
7.
Surgical team
These features depend on the constellation and interaction of the surgical team like the position of the different team members around the patient or the quality of team communication. Additionally, characteristics, which are individual for each team member, like their experience in the respective surgical procedure or their current stress level as measured by cortisol level or heart rate, are included in this category. Furthermore, team aspects such as the number of procedures that two or more members of the surgical and/or anaesthesiological team have performed together prior to the operation, can be considered within this category. However, data sources to collect intraoperative information on the team like a microphone or a room camera in the OR have to be used with caution considering data privacy concerns.
-
8.
Environment
Features regarding the environmental setting of the OR containing metadata of the surgery are summarized in this category. Examples are the emotional atmosphere, the noises, or the temperature in the operating room. Again, a microphone or a room camera could be used for data acquisition.
Rating of surgomic feature categories. Ratings are displayed in each subplot per feature category for clinical relevance regarding morbidity and mortality, clinical relevance regarding long-term (oncological) outcome and technical feasibility. Colors depict ratings of surgeons and scientists, respectively. The only significant difference between surgeons and scientists was in the category “surgical skill and quality of performance” regarding the relevance for morbidity and mortality (p = 0.002)
Multicenter expert survey
In the next step a questionnaire presenting all surgomic feature categories and their respective features was developed to validate and rate them. The questionnaire was implemented and analyzed with the online tool LimeSurvey (LimeSurvey GmbH, Hamburg, Germany) hosted by Heidelberg University (Heidelberg, Germany). We sent out the questionnaire to the departments of general and visceral surgery at three German university hospitals, six German research groups from computer science, the section for computer and telematic assisted surgery of the German Society of Surgery (DGCH), and participants of the endovis-challenges HeiChole 2019 and HeiSurf 2021 (https://endovis.grand-challenge.org/).
The opening page of the questionnaire explained the concept of Surgomics and mentioned the different partners involved in the project.
The participants were then asked for their gender, professional background (general or visceral surgeon, other surgical specialty, computer scientist, scientist from other discipline, other) and years of professional work experience since graduation (for surgeons since beginning of residency).
In the next step the participants were asked to rate the eight feature categories on a numeric rating scale from 1 to 10 (1 = not relevant/feasible; 10 = extremely relevant/feasible) regarding their:
-
(1)
Clinical relevance for morbidity ("patient suffers postoperative complications") and mortality ("patient dies within 90 days after surgery"),
-
(2)
Clinical relevance for the long-term oncological outcome ("survival, e.g. postoperative survival 10 years after surgery"),
-
(3)
Technical feasibility of automatic recognition with machine learning ("Artificial Intelligence").
Each feature category was displayed with at least one example. If the participant needed more information about a category, there was the option to hover over the category and get a more detailed description displayed.
Finally, participants were asked to rank the 52 available surgomic features within their respective category (Table 1) via drag and drop regarding their overall priority. Again, if the participant needed more information about a specific surgomic feature, there was the option to hover over the feature and display a more detailed description. Additionally, participants were invited to add missing surgomic features in a free text field.
Statistical analysis
The results of the survey were analyzed using descriptive statistics with Python 3.10 using pandas 1.4, scipy 1.8 and seaborn 0.11, (Python Software Foundation [33]). For the surgomic feature categories mean and standard deviation were calculated for clinical relevance and technical feasibility. For the priority ranking of the surgomic features in each category mean ± standard deviation of the rank were calculated. Results were calculated separately for surgeons and (computer) scientists. A test for normal distribution was performed with the Shapiro–Wilk-test [34] and results were compared with Welch's unequal variances t test [35].
Results
Overall, 52 intraoperative surgomic features have been identified in the multidisciplinary panel discussion. Table 1 gives an overview of the surgomic features and their properties. Most features were assigned to the “surgical skill and quality of performance” category (n = 11). The categories “temporal course of procedure”, “anesthesiological data” and “environment” had the least features (n = 3 each). The categories “surgical field” and the “surgical team” (n = 9 each) as well as “instrument” and “patient” (n = 7 each) had intermediate numbers of features. Overall, n = 9 (17%) features were procedure specific, meaning that they cannot be derived and analyzed in every surgery like the ICG-accumulation curve of an anastomosis.
The anonymous expert survey was then conducted with n = 66 participants from multiple centers. This included 34 surgeons, all working in general or visceral surgery, 8 (24%) were female and 26 (76%) were male. They had been working in their professional field since the beginning of residency for a mean of 11.9 ± 7.0 years. Furthermore, 28 scientists participated, 23 of them computer scientists (82%), 7 (25%) of them were female and 21 (75%) male. They had been working as scientists since graduation for a mean of 7.6 ± 7.8 years. Participants with “other” professional backgrounds were excluded from analysis (n = 4).
Clinical relevance for morbidity and mortality as rated by surgeons was highest for the feature category “surgical skill and quality of performance” (9.0 ± 1.3) with an average rating of 6.8 ± 1.1 over all categories and also for long-term outcome (8.2 ± 1.8) with an average rating of 5.8 ± 1.2 over all categories. Technical feasibility of automatic recognition with ML as rated by scientists was highest for the “Instrument” category (8.5 ± 1.7) with an average feasibility rating over all categories of 6.8 ± 1.3. Figure 3 gives an overview of the ratings for all categories.
Surgeons and scientists gave the highest ranking of overall relevance within their respective category to the same surgomic features “intraoperative adverse events” (surgical field category), “action performed with instrument” (instrument category), “vital sign monitoring data” (anesthesiological category), “difficulty of surgery” (surgical skill category), “familiar surgeries performed / experience with surgery” (surgical team category), and “emotional atmosphere in OR” (environment category). Table 2 gives an overview of the ranking for all surgomic features.
In the free text answers for new surgomic features a total of n = 34 new features were mentioned. Most ideas for new features were given in the “anesthesiological data” category and the “surgical field” category. Here, eight new feature ideas were added each, e.g. “degree of relaxation/depth of narcosis” and “anastomosis technique (in general which techniques were used)”, respectively. In the “anesthesiological data” category “ventilation parameters” were mentioned by n = 3 participants. The “instrument” category followed with five new feature ideas, e.g. “movement of instrument (trajectory)” (n = 2 participants). In this category one participant also suggested an intraoperative assistance based on the surgomic features, i.e. the “visual highlight of potentially dangerous instruments when inserted”. In category “temporal course of procedure” n = 6 participants suggested the duration of certain phases as a new feature, which was originally listed by us as part of the feature “duration of surgery”, but should be a separate feature. The full list of the free text answers for new surgomic feature ideas can be found in supplement 1.
Discussion
Conceptualizing Surgomics
With Surgomics we coined a neologism to comprehensively describe a surgical procedure based on automatic analysis of intraoperative data. This is in analogy to Radiomics, where the mere idea that “images are more than pictures, they are data” is far from new [61]. The specific challenge of surgical data is taking temporal data and the whole surgical team into account. Here, the representation of temporal data is far from trivial and actually an open research question. Data has been systematically collectedin the operating room to improve patient care [24], ML has been used for surgical skill assessment [62]. However, with Surgomics our aim is to go beyond assessment of an individual surgeon’s skill. The aim is to approach the massive amount of data that is generated daily in the operating rooms with a holistic perspective on all possible sources of data and provide a conceptual framework for future developments of automatic analysis of this data keeping the improvement of surgical patient care in focus. Thereby, Surgomics can also add a fine granular, objective analysis to previous approaches that proved the importance of intraoperative adverse event classification [44, 45].
We thus gathered a group of 19 specialists from both surgery and (computer) science to flesh out the concept of Surgomics. As a result we came up with a first collection of 52 surgomic features that may serve as surrogate parameters for the quality of patient care and defined eight categories to group them (Table 1).
When validating this conception with a broader audience of surgeons and (computer) scientists we found the rating of relevance for morbidity and mortality for the category “surgical skill and quality of performance” as the only one with a significant difference between surgeons and scientists (p = 0.002). In contrast to surgeons, scientists rated the category “anesthesiological data” as most relevant for morbidity and mortality and not “surgical skill and quality of performance”. We assume this surprising agreement to result from a selection bias, because the participating (computer) scientists are members of surgical-technical societies and/or have a long track record of scientific work in computer-assisted surgery. This highlights that a close scientific exchange between clinicians and computer scientists is beneficial or even mandatory to ensure optimal planning of the next steps to differentiate and develop individual surgomic features.
Surgeons rated the category “surgical skill and quality of performance” highest in relevance for morbidity and mortality as well as for long-term (oncological) outcome. The feasibility of this category, however, was rated penultimate by scientists. Interestingly, the surgomic feature rated with highest overall priority in this category was “difficulty of surgery” far beyond sub-dimensions of the established GOALS-score such as “tissue handling”, “tissue exposure” and “efficiency”.
A better relation between clinical relevance and technical feasibility was found for “anesthesiological data” with high ratings for morbidity and mortality, but not for long-term (oncological) outcome. At the same time this category had the second highest rating for feasibility, rendering it a good candidate for further investigation. When rating the features surgeons and scientists agreed on “vital sign monitoring” having the highest relevance. Here it would be interesting to investigate which parameter(s), such as heart rate, arterial blood pressure, central venous pressure or ventilation parameters are of highest relevance.
Another promising category is “patient” with the second highest rating in long-term (oncological) outcome and third in morbidity and mortality. Whereas the technical feasibility was rated lower, surgomic features that could be easily obtained with computer vision from laparoscopic video such as “texture of liver”, “texture of lungs” and “texture of pancreatic gland” had high relevance ratings.
Other feature categories that have been of high interest in the scientific literature on computer vision for laparoscopic surgery, namely “surgical field”, “instrument” and “temporal course of procedure” received only medium ratings for clinical relevance, but comparatively high technical feasibility. Because of that high technical feasibility these categories should also be a matter of future investigations, in order to create a success story of the Surgomics concept that would boost motivation to tackle more challenging features for example from the category of “surgical skill and quality of performance”. Good feature candidates would be the highly rated “intraoperative adverse events”, “bloodiness”, “action performed with anatomic structure”, “action performed with instrument”, and “instrument usage”.
The category “environment” was neither rated high in clinical relevance nor in technical feasibility and will thus have a low priority for future investigations.
Although many surgomic features have been identified and described, our list is far from complete. The free text questions for more surgomic feature ideas already revealed 34 new features. Furthermore, a relevant proportion of our 52 surgomic features is to be extracted from surgical video in minimally invasive surgery. More surgomic features from various sources of raw data will be identified in surgical and scientific discussions as well as during the development process of other features. Either way, the suitability of every surgomic feature for a certain application has to be validated thorougly before introduction into clinical practice.
A limitation of our study is that in our group only general and visceral surgeons participated, potentially missing features that would be important for other specialties such as cardiac or thoracic surgery. Also, other parts of the OR team such as anesthesiologists and scrub nurses have not been involved in our study, but probably will come up with other surgomic features in future discussions.
Furthermore, some definitions of features and their categories can be hard to distinguish. The instrument category could be perceived as part of the surgical field category, but we chose to define a different category to distinguish what happens in the surgical field and which instruments perform these activities integrating data that is not derived from the video, but from connected medical devices (e.g. electrocautery).Other surgomic features like the “emotional atmosphere in the OR”, which was assigned to the “environment” category, could as well be assigned to the “surgical team” category.
Pre-, intra- and postoperative use
Based on the idea of using intraoperative data for improvement of patient care, numerous potential applications for Surgomics along the whole treatment path arise. Preoperative information about the patient (comorbidities, previous surgeries) as well as their disease (tumor stage, lymph node involvement) is routinely collected. It is obvious that this information will influence the intraoperative course. For example, previous surgeries may result in more or less severe abdominal adhesions and some surgical centers may limit minimally invasive oncologic surgery to T2/T3 cancers. Antithrombotic medication may turn a straightforward procedure into a more demanding one. Whereas established metrics such as overall procedure time or intraoperative blood loss can hint towards those correlations, surgomic features such as bloodiness or length of certain key steps of a procedure can give deeper insights into correlations or even causality.
Intraoperatively, assistance-systems based on Surgomics may support learning surgeons with real time analysis and a recommendation of what other surgeons would do in the same situation. Furthermore, Surgomics may help with intraoperative decision-making for individual patients by correlating quantitative measurements such as gastric tube width or indocyanine green (ICG) perfusion quantification with other procedure or postoperative outcome parameters.
Postoperatively, the aim of Surgomics is to predict life-threatening complications. Even if the blood loss as estimated by the anesthesiologist is low, the “bloodiness” of an operation may be high in a specific part of the procedure leading to postoperative hematoma and infection. Even if the subjective impression of ICG perfusion of an anastomosis seems good, the accumulation of the perfusion in combination with an ruptured suture may result in anastomotic insufficiency. By applying the concept of Surgomics, clinical researchers may find correlations that can be used to develop clinical decision support systems based on those surgomic features. Here, the aim would be to predict complications of the individual patient before they become clinically apparent and to warn surgeons accordingly [56] to reduce the rate of “failure to rescue” [63].
On an individual surgeon level, Surgomics may be used as a foundation for an objective quantitative metric of surgeon performance. By incorporating not only audiovisual data from the OR but other sensors from medical devices, vital sign monitoring, etc. Surgomics is a more comprehensive concept than surgical sabermetrics. Here, Surgomics may be used for regular assessment of surgical residents to objectify their progress and tailor their training accordingly. For experienced surgeons, Surgomics may help to establish data-driven morbidity and mortality analysis and bring up hypotheses for performance improvement by providing deeper insights into correlation of intraoperative activities with postoperative outcomes. Also, in contrast to global scores of overall performance, Surgomics again may help surgeons by breaking down potential reasons for a case being exceptionally difficult by giving insights into factors of patient, surgeon, team, and environment. In this manner, Surgomics can be a methodological approach to use ML for automatic surgical quality assessment, or to better classify and compare procedures in a learning curve.
Furthermore, Surgomics may become a tool to automate the comprehensive description and classification of a surgical procedure, similar to TNM in pathology, going beyond already promising approaches to categorize intraoperative adverse events [44, 45]. Surgomics thus may serve to improve comparability of clinical performance and procedures in surgical trials.
Ethical considerations
When introducing a concept that provides comprehensive insights into the intraoperative course and might even produce objective data of inadequate surgeon performance, difficult ethical questions arise. Here, we are having the same situation as e.g. in aviation with flight recorders (black box) or with CIRS (critical incident reporting systems) in hospitals. If in analogy further approaches regarding the assessment of the surgeon’s skill and the quality of performance are to be made, ethical considerations are extremely important. There is no doubt that performance data needs to be collected objectively [14] with the surgical team members being fully aware of the recording, the further use of the data and the possible consequences of inadequate surgical performance. Moreover, concerns regarding privacy and litigation need to be addressed as investigated in a study on perception of the OR blackbox® [64]. However, nowadays some authors consider a routine video recording of surgeries an ethical duty and a standard of care with potential benefits for e.g. safety, fairness and candor [65]. Already in 2011, the American Board of Medical Specialties (ABMS) defined requirements that a systematic tracking of the surgeon’s performance is necessary [12]. Furthermore, in case there are legal implications, it might even be helpful to have proper documentation of the performed activities to protect surgeons. Apart from the legal aspects, video-based coaching sessions with senior surgeons focusing on the operative technique have already been successfully introduced [66]. Using Surgomics for an automated assessment of surgical skills with anonymous, constructive feedback based on expert knowledge may even be less intimidating than personal feedback from a surgical supervisor a surgeon is dependent on. Nevertheless, we are convinced that objective data of performance can open the room for discussion and improvement. This way, a positive culture regarding mistakes is to be established with surgeons being aware of their own performance level, strengths and room for improvement.
Next steps
The next step to realize the concept of Surgomics would be the automatic recognition of features. Here, our work not only introduces a concept, but provides a conceptual framework for future research that can help to shape the process of intraoperative data analysis and provides guidance for surgeons and computer scientists alike. Surgeons with ideas for new surgomic features find the technical feasibility of feature categories rated by (computer) scientists and (computer) scientists that want to apply their technology to surgery find the clinical relevance of surgomic feature categories rated by surgeons. Based on the analysis of our survey with experienced professionals from surgery and (computer) science, surgomic features like “vital sign monitoring” (“anesthesiological data” category) as well as texture of liver, lung, and pancreas (“patient” category) have a suitable combination of expected clinical relevance and technical feasibility. Furthermore, the surgomic features “intraoperative adverse events”, “bloodiness”, “action performed with anatomy” (“surgical field” category), “action performed with instrument”, and “instrument usage” (“instrument” category), should be investigated. Because of their high technical feasibility and the already active scientific community in the field of laparoscopic computer vision, they represent “low hanging fruits” to create success stories for Surgomics.
This success also depends on open source availability of annotated surgical data and ML models. Annotated surgical data can be used in international ML challenges to validate algorithms for automatic extraction of surgomic features from surgical data. The openly available ML models could then be used by surgical researchers at different centers to investigate the use of surgomic features as new parameters in clinical research. This in turn would stimulate discussions within surgical societies to investigate more surgomic features and their application to clinical problems similar to current developments in Radiomics and Genomics for precision medicine.
Conclusion
The aim of Surgomics is to predict a patient's morbidity, mortality and long-term outcome using ML to derive personalized information from comprehensive intraoperative sensor data and combining it with pre- and postoperative information. With a multidisciplinary group of experts, we defined this concept, and came up with 52 features in eight categories that were rated by experienced professionals from surgery and (computer) science. This is a first step to establish Surgomics as the surgical answer to Radiomics and Genomics for improving surgical patient care.
References
Head SJ, Milojevic M, Daemen J, Ahn J-M, Boersma E, Christiansen EH, Domanski MJ, Farkouh ME, Flather M, Fuster V, Hlatky MA, Holm NR, Hueb WA, Kamalesh M, Kim Y-H, Mäkikallio T, Mohr FW, Papageorgiou G, Park S-J, Rodriguez AE, Sabik JF, Stables RH, Stone GW, Serruys PW, Kappetein AP (2018) Mortality after coronary artery bypass grafting versus percutaneous coronary intervention with stenting for coronary artery disease: a pooled analysis of individual patient data. Lancet Lond Engl 391:939–948. https://doi.org/10.1016/S0140-6736(18)30423-9
Sullivan R, Alatise OI, Anderson BO, Audisio R, Autier P, Aggarwal A, Balch C, Brennan MF, Dare A, D’Cruz A, Eggermont AMM, Fleming K, Gueye SM, Hagander L, Herrera CA, Holmer H, Ilbawi AM, Jarnheimer A, Ji J, Kingham TP, Liberman J, Leather AJM, Meara JG, Mukhopadhyay S, Murthy SS, Omar S, Parham GP, Pramesh CS, Riviello R, Rodin D, Santini L, Shrikhande SV, Shrime M, Thomas R, Tsunoda AT, van de Velde C, Veronesi U, Vijaykumar DK, Watters D, Wang S, Wu Y-L, Zeiton M, Purushotham A (2015) Global cancer surgery: delivering safe, affordable, and timely cancer surgery. Lancet Oncol 16:1193–1224. https://doi.org/10.1016/S1470-2045(15)00223-5
Nepogodiev D, Martin J, Biccard B, Makupe A, Bhangu A, National Institute for Health Research Global Health Research Unit on Global Surgery (2019) Global burden of postoperative death. Lancet Lond Engl 393:401. https://doi.org/10.1016/S0140-6736(18)33139-8
Wente MN, Veit JA, Bassi C, Dervenis C, Fingerhut A, Gouma DJ, Izbicki JR, Neoptolemos JP, Padbury RT, Sarr MG, Yeo CJ, Büchler MW (2007) Postpancreatectomy hemorrhage (PPH): an International Study Group of Pancreatic Surgery (ISGPS) definition. Surgery 142:20–25. https://doi.org/10.1016/j.surg.2007.02.001
Fabbi M, Hagens ERC, van Berge Henegouwen MI, Gisbertz SS (2020) Anastomotic leakage after esophagectomy for esophageal cancer: definitions, diagnostics, and treatment. Dis Esophagus. https://doi.org/10.1093/dote/doaa039
Wright CD, Kucharczuk JC, O’Brien SM, Grab JD, Allen MS (2009) Predictors of major morbidity and mortality after esophagectomy for esophageal cancer: A Society of Thoracic Surgeons General Thoracic Surgery Database risk adjustment model. J Thorac Cardiovasc Surg 137:587–596. https://doi.org/10.1016/j.jtcvs.2008.11.042
Bilimoria KY, Liu Y, Paruch JL, Zhou L, Kmiecik TE, Ko CY, Cohen ME (2013) Development and Evaluation of the Universal ACS NSQIP Surgical Risk Calculator: a decision aid and informed consent tool for patients and surgeons. J Am Coll Surg 217:833-842.e3. https://doi.org/10.1016/j.jamcollsurg.2013.07.385
Birkmeyer JD, Siewers AE, Finlayson EVA, Stukel TA, Lucas FL, Batista I, Welch HG, Wennberg DE (2002) Hospital volume and surgical mortality in the United States. N Engl J Med 346:1128–1137. https://doi.org/10.1056/NEJMsa012337
Knight SR, Shaw CA, Pius R, Drake TM, Norman L, Ademuyiwa AO, Adisa AO, Aguilera ML, Al-Saqqa SW, Al-Slaibi I, Bhangu A, Biccard BM, Brocklehurst P, Costas-Chavarri A, Chu K, Dare A, Elhadi M, Fairfield CJ, Fitzgerald JE, Ghosh D, Glasbey J, Henegouwen MI van B, Ingabire JCA, Kingham TP, Lapitan MC, Lawani I, Lieske B, Lilford R, Martin J, McLean KA, Moore R, Morton D, Nepogodiev D, Ntirenganya F, Pata F, Pinkney T, Qureshi AU, Medina AR-D la, Riad A, Salem HK, Simões J, Spence R, Smart N, Tabiri S, Thomas H, Weiser TG, West M, Whitaker J, Harrison EM, Gjata A, Modolo MM, King S, Chan E, Nahar SN, Waterman A, Vervoort D, Lawani I, Bedada AG, Azevedo BD, Figueiredo AG, Sokolov M, Barendegere V, Ekwen G, Agarwal A, Dare A, Liu Q, Correa JC, Malemo KL, Bake J, Mihanovic J, Kuncarová K, Orhalmi J, Salem H, Teras J, Kechagias A, Arnaud AP, Lindert J, Tabiri S, Kalles V, Aguilera-Arevalo M-L, Recinos G, Baranyai Z, Kumar B, Lakshmi HN, Zachariah SK, Alexander P, Venkatappa SK, Pramesh C, Amandito R, Fleming C, Ansaloni L, Pata F, Pellino G, Altibi AM, Nour I, Hamdun I, Elhadi M, Ghellai AM, Venskutonis D, Poskus T, Zilinskas J, Whitaker J, Malemia P, Tew YY, Borg E, Ellul S, Medina AR-D la, Wafqui fatima Z, Borowski DW, Dalen AS van, Wells C, Adamou H, Ademuyiwa A, Adisa A, Søreide K, Qureshi AU, Al-Slaibi I, Saqqa SA, Alser O, Tahboub H, Lohse HAS, Yip SS, Lapitan MC, Major P, Simões J, Soares AS, Bratu MR, Litvin A, Vardanyan A, Ingabire JA, Costas-Chavarri A, Gudal A, Albati N, Juloski J, Lieske B, Rems M, Rayne S, Straten SV, Moodley Y, Chu K, Moore R, Vázquez IO, Ruiz-Tovar J, Senanayake KJ, Thalgaspitiya SPB, Omer OA, Homeida A, Cengiz Y, Clerc D, Alshaar M, Bouaziz H, Altinel Y, Doe M, Freigofer M, Teasdale E, Kabariti R, Clements JM, Knight SR, Ashfaq A, Azodo I, Wagner G, Trostchansky I, Maimbo M, Linyama D, Nina H, Zeko A, Fermani CG, Modolo MM, Villalobos S, Carballo F, Farina P, Guckenheimer S, Dickfos M, Ajmera A, Chong C, Gourlay R, Hussaini S, Lee YJ, Majid A, Martin P, Miles R, Morris OJ, Phua J, Ridley W, Saluja T, Tan RR, Teh J, Wells A, Arora B, Dollie Q, Ho D, Ma Y, Perera OM, Truong A, Dawson AC, Lim B, Pahalawatta U, Phan J, Woon-Shoo-Tong X-MS, Yeoh A, Charman L, Drane A, Laura S, Lo CCW, Mozes A, Poon R, Tan HH, Wall E, Chopra P, Giovanni JD, Dhital B, Draganic B, Duller A, Gani J, Goh YK, Jeong JY, McManus B, Nagappan P, Pockney P, Rugendyke A, Sarrami M, Smith S, Wills V, Wong HV, Ye G, Zhang G, Brooker E, Feng D, Lau B, Ngai C, Birks S, Gyorki D, Pablos JO de, Abbosh A, Gillespie C, Mahmoud A, Kwan B, Lawson J, Warwick A, Bingham J, Cockbain AJ, Dudi-Venkata NN, Ellaby-Hall J, Finlay B, Humphries E, Pisaniello J, Pisaniello M, Salih S, Sammour T, Wahab HHA, Silva AD, Hayward N, Iyer K, Maddern G, Prevost GA, Annapureddy N, Settipalli KP, Yeo J, Hempenstall L, Pham L, Purcell S, Talavera C, Vaska AI, Chaggar G, Chrapko P, Cocco A, Coulter-Nile SMCJ, Ctercteko G, French J, Gong H, Gosselink M, Jegathees T, Jin I, Kalachov M, Kiefhaber K, Lee K, Luong J, Phan S, Pleass H, Veale K, Zeng Z, Au A, DeBiasio A, Deng I, Myooran J, Nair A, Stewart P, Stift A, Unger LW, Wimmer K, Ahmed N, Hasan S, Rahman S, O’Shea M, Padmore G, Peters A, Perduca P, Pulcina G, Tinton N, Buxant F, Dabin E, Garofalo G, Dossou F, Lawani I, Gnangnon FHR, Souaibou YI, Bedada AG, Motlaleselelo P, Tlhomelang O, Buarque IL, Gomes GMA, Barros AV, Batashki I, Damianov N, Stoyanov V, Dardanov D, Maslyankov S, Petkov P, Sokolov M, Todorov G, Zhivkov E, Akisheva A, Moreno MAC, Genov G, Ilieva I, Ivanov T, Karamanliev M, Khan A, Mitkov E, Yotsov T, Atanasov B, Belev N, Slavchev M, Nsengiyumva C, Jones E, Stock S, Ekwen G, Kyota S, Brown J, K TM, Samuel LN, Otuneme C, Prosper N, Umenze F, Boutros M, Caminsky N, Dumitra S, Garfinkle R, Morency D, Salama E, Banks A, Ferri L, He H, Katz A, Liberman AS, Meterissian S, Pang A, Parvez E, Agarwal A, Dare A, Hameed U, Osman F, Sequeira S, Coburn N, Dare A, Jaffer A, Karanicolas P, Mosseler M, Musselman R, Liu X, Yip CW, Garces-Otero JS, Guzman C, Sierra S, Valencia AU, Rivera PAC, Camelo S, Gonzalez A, González-Orozco A, Paz MSM, Rivera CJ-P, Gonzalez F, Isaza-Restrepo A, Torres LN-, Madrid NA, Arango MCM, Sierra S, Bake J, Tsandiraki J, Jemendžic D, Kocman B, Šuman O, Canic R, Jurišic D, Karakas I, Rupcic AK, Pitlovic V, Samardžic J, Kopljar M, Bacic I, Domini E, Karlo R, Mihanovic J, Miljanic D, Simic A, Ahmed M, Nassrallah MA, Altaf R, Amjad T, Eltoum R, Haidar H, Hassan A, Khalil O, Qasem M, Ramesh R, Sajith G, Wisal M, Žatecký J, Bujda M, Jirankova K, Paclik A, Abdallah A, Almogy MA, El-sawy EA, ElFayoumy AM, Elghareeb N, Esmat NA, Fadel A, Habater A, Hamdy H, Hefni A, Kamal M, Abobakr NM, Sayed A, Shaker N, Taha E, Tharwat H, Zakaria O, Abdelmotaleb I, Al-Dhufri A, Al-Himyari HS, Sheikh EE, Eldmaty A, Elkhalawy A, M.Elkhashen A, Magdy K, Mostafa S, Sadia HD, Saleh M mahmoud, Samir D, Ali MYM, Nassar MA, Abdelhady S, Abdelrazek A, Abdelsalam I, El-Sawy A, Essam E, Gadelkarim M, Ghaly K, Hassabalnaby M, Masarani R, Shaaban NM, Sabry A, Salem M, Soliman NA, Zahran D, El.soud MRA, Badr ET, Borham H, Elmeslemany N, Elsayed M, Elsherif F, Eslam S, Gaber G, Ibrahim S, Kamh Y, Mahmoud A, Mohamed S gamal, Morshedy E, Omar C, Soliman FS, Abdelkawy S, Abdelmohsen N, Abdelshakour M, Dahy A, Gamal N, Gamal M, Hasan A, Hetta H, Mousa N, Omar M, Rabie S, Saad M, Saleh B, Mohamed MS, Shawqi M, Mousa HA, Alnoury M, Elbealawy M, Elshafey A, Ahmed MEIEDM, Ghonaim M, Hgag F, Ibrahim M, Morsy M, Loaloa MR, Refaat A, Samir H, Shahien F, Sobhy M, Sroor F, Abdellatif E, Adel M, Afifi AA, Afifi E, Antaky M, Dawoud A, Zoghby NE, El-remaily A, Elzanfaly AA, Gadallah A, Gamal FA, Hashem O, Youssef SM, Attyah AM, Munir M, Shazly O, Taha E, Wilson K, Adel S, Ali A, Eid E, Elhelow E, Elmahdy M, Elshatby B, Zakaria AH el-din, Hossny A, Ibrahim E, M.Yonis A, Metwalli M, Yousry B, Zid E, Yacoub MA, Abdelhakim A, Abouelsoad N, Alkhatib M, Ashraf A, Ashraf A, Elazab Y, Elfanty M, Elkabir O, Elsayed M, Elshimy A, Elsobky H, Eskander J, Gad A, Hamsho W, Abdelwahed NK, Magdy M, Moharam D, Osama A, Ramadan S, Roum R, Sayed T, Shehada T, Zidan AM, Abbas K, Ali A, Attia M, Balata M, Nakeeb AE, Elewaily MIE, Elfallal A, Elfeki H, Elkhadragy A, Emile S, Ezzat H, Hosni H, Mansour I, Omar W, Othman G, Sadek K, Shalaby M, Shehab-Eldeen N, Khalifa RA, Badr H, Eldeep M, Eldeep A, Mohammed AE, Khallaf S, Hegazy EM, Mahmoud R, Mikhail P, Morsi M, Mowafy S, Raafat D, Safy A, Sera M, Sera A shible, AbdAllah MSM, Abdelkader M, Abdou AO, Ahmed A, Gaafar S, Negm FI, Lapic M, Maher A, Mahmoud H, Mostafa A, Samir M, Samy F, Semeda N, Shalaby HI, El-taweel A, Elnagar AG, Hemidan AG, Hussein M, Kandil AA, Moawad M, Hamamah AAN, Soliman M, Abdelkhalek M, Tawakel NA, Abdelwahed AM, Abdou A, Atallah K, Elsherbeny MY, Emara E, Hamdy M, Hamdy O, Haron A, Ismail S, Metwally IH, Elgaml NMH, Nassar A, Refky B, Sadek M, Saleh M, Yunes A, Zakaria M, Zuhdy M, Fayed N, Mohammed MMH, Kütner S, Melnik P, Seire I, Teras J, Ümarik T, Ainoa E, Eerola V, Koppatz H, Koskenvuo L, Sallinen V, Takala S, Katunin J, Kechagias A, Turunen A, Christou N, Mathonnet M, Lavoue V, Timoh KN, Soulabaille L, Lesourd R, Merdrignac A, Sulpice L, André B, Chantalat E, Vaysse C, Dousset B, Gaujoux S, Martin G, Clonda O, Juodis D, Kienle K, Mravik A, Palmer S, Szabadhegyi G, Agbeko AE, Gyabaah S, Gyamfi FE, Naabo N, Senior AO, Yorke J, Owusu F, Abantanga F, Anyomih TTK, Muntaka A-JM, Abem EO, Sheriff M, Tabiri S, Wondoh PM, Balalis D, Korkolis D, Gkiokas G, Pantiora E, Theodosopoulos T, Ioannidis A, Konstantinidis K, Konstantinidou S, Machairas N, Paspala A, Prodromidou A, Chouliaras C, Papadopoulos K, Baloyiannis I, Mamaloudis I, Tzovaras G, Akrida I, Argentou M-I, Germanos S, Iliopoulos E, Maroulis I, Skroubis G, Theofanis G, Chatzakis C, Ioannidis O, Loutzidou L, Kalles V, Karathanasis P, Michalopoulos N, Theodoropoulos C, Theodorou D, Triantafyllou T, Garoufalia Z, Hasemaki N, Kontos M, Kouraklis G, Kykalos S, Liakakos T, Mpaili E, Papalampros A, Schizas D, Syllaios A, Tampaki EC, Tsimpoukelis A, Antonopoulou MI, Deskou E, Manatakis DK, Papageorgiou D, Zoulamoglou M, Anthoulakis C, Margaritis M, Nikoloudis N, Campo V, Ceballos A, Flores M-A, Giron W, Ko D, Martinez G, Recinos G, Lara VR, Rueda N, Sanchez A, Garrido JCGT, Aguilera-Arevalo M-L, Rivera AEA, Ixcajoc EBB, Zelaya LEB, Chacòn-Herrera P, Ruiz LMC, Echeverria-Davila G, Garcia M, García D, Mayen EFG, José N, Mazariegos N, Méndez D, Espinoza MP, Baranyai Z, Bardos D, Benke M, Illes K, Kokas BA, Szabó R, Appukuttan A, Asok A, D.k V, Malik K, Ravishankaran P, Tapkire R, Moorthy G, Abraham J, Muthuvel R, Alapatt J, Kattepur A, Pareekutty N, Garod M, Harris C, Wanniang C, Gupta A, Nehra D, Parshad S, Acharya R, Badwe R, Bhandare M, Jain U, Kirti K, Nair N, Shrikhande S, Thakkar P, Anandan P, S AC, Narasannaiah AH, Jagarlamudi T, Venkatappa SK, R RM, Manangi M, Raghavendra A, Rao KS, S V, Sajjan V, Shenoy A, Chikkanayakanahalli SS, Tharanath K, V S, Adidharma P, Agarwal R, Amandito R, Gultom PA, Arifin GR, Billy M, Elfizri Z, Fahira A, Felicia D, Gunardi TH, Johanna N, Nugrahadi NR, Panigoro SS, Rahmayanti S, Sihotang RC, Brata SY, Winoto H, Barati N, Karami M, Khorshidi H, Naderifar H, Abdulla MA, Coleman M, Doherty RJ, Hannon R, Murphy B, Stakelum A, Winter D, Aljohmani L, Farnan R, Seldon Y, Tan T, Varghese S, Alherz M, Ather M, Bajilan M, Graziadei V, Pilkington I, Quidwai O, Ridgway P, Shiwani H, Tahir A al-Rahman, Blunnie E, Burke D, Kennedy N, Macdonagh K, O’Neill M, Rooney S, Falco G, Ferrari G, Mele S, Nita GE, Ugoletti L, Zizzo M, Confalonieri G, Pesenti G, Tagliabue F, Baronio G, Ongaro D, Pata G, Compagnoni B, Salvadori R, Taglietti L, D’Alessandro N, Lascio PD, Pascale G, Bortolasi L, Campagnaro T, Carlini M, Lisi G, Lombardi D, Pedrazzani C, Spoletini D, Turri G, Violi P, Altomare DF, Aquilino F, Musa N, Papagni V, Picciariello A, Vincenti L, Andreotti D, Occhionorelli S, Tondo M, Basso SMM, Cirelli R, Maino MEM, Piozzi GN, Picone E, Scaramuzzo R, Sinibaldi G, Amendola A, Anastasio L, Bucci L, Caruso E, Castaldi A, Maso SD, Dinuzzi VP, Esposito G, Gaudiello M, Giglio MC, Greco PA, Luglio G, Manfreda A, Marra E, Mastella F, Pagano G, Peltrini R, Pepe V, Sacco M, Sollazzo V, Spiezio G, Cianchetti E, Menduni N, Carvello MM, Candido FD, Spinelli A, Corsi F, Sorrentino L, Marino F, Asti ELG, Bonavina L, Rausa E, Asta M, Belli A, Bianco F, Cervone C, Delrio P, Falato A, Bucci AF, Guarino R, Pace U, Rega D, Luca ED, Gallo G, Sammarco G, Sena G, Vescio G, Santandrea L, Ugolini G, Zattoni D, Chetta N, Logrieco G, Vanella S, Garulli G, Zanini N, Bondurri A, Cammarata F, Colombo F, Foschi D, Lamperti GMB, Maffioli A, Sampietro GM, Yakushkina A, Zaffaroni G, Ansaloni L, Cicuttin E, Sibilla MG, Impellizzeri H, Inama M, Moretto G, Mochet S, Ponte E, Usai A, Mancini S, Sagnotta A, Solinas L, Bolzonaro E, Tamini N, Curletti G, Galleano R, Malerba M, Campanella S, Cocorullo G, Colli F, Marco PD, Falco N, Fontana T, Mambou L jospin K, Brocca AL, Licari L, Randisi B, Rizzo G, Rotolo G, Salamone G, Tutino R, Venturelli P, Malabarba S, Sgrò A, Vella I, Cirillo B, Crocetti D, Toma GD, Lapolla P, Mingoli A, Sapienza P, Belvedere A, Bianchini S, Binetti M, Birindelli A, Tonini V, Podda M, Pulighe F, Rosa MD, Bono L, Borghi F, Geretto P, Giuffrida MC, Lauro C, Marano A, Pellegrino L, Salusso P, Sasia D, Campanelli M, Luc AR, Trompetto M, Cardia R, Cillara N, Giordano AN, Costanzo A, Giovilli MA, Turati L, Canonico S, Pellino G, Sciaudone G, Selvaggi F, Selvaggi L, Albsoul N, AlBsoul A, Alkhatib AA, Alsallaq O, Amarin JZ, Ayoub R, Bsisu I, Muhtaseb MSE, Jabaiti M, Melhem J, Nour I, Qwaider YZ, Salameh MH, Suleihat A, Suradi HH, Alammarin M, Aljaafreh A, Hani MB, Hani ZB, Hani FB, Fahmawee T, Hamouri S, Katanani C, Tawalbeh R, Tawalbeh T, Zawahrah H, Chaar MKA, Abusalem L, Al-Masri M, Al-Najjar H, Barghuthi L, Ahmed Z, Maulana A, Ngotho O, Kamau C, Mwenda AS, Bosire F, Mwachiro E, Parker R, Simel I, Sylvester K, Althini AAM, Elbarouni S, Elbeshina AE, Gwea A, Malek A, Farag WAM, Abdalei A, Malik ABA, Abo-khammash A, Abuhlaiga M, Adnan N, Albaggar M, Alfitory A, Aljanfi A, Almuzghi F, Altumei Z, Alzabti F, Ashoushan H, Assalhi M, Azzubia J, Bnhameida S, Delhen M, Elshafei H, Elteir H, Esbaga F, Gobbi AA, Hamouda F, Hilan H, Ismail R, Jebran F, Kasbour M, Maderi G, Mohammad S, Mohammed B, Murtadi H, Mustafa H, Rajab M, Trenba S, Wafaa M, Sagheir EA, Almigheerbi A, Alzahaf A, Bahroun SG, Dallah NB, Elshaibani M, Eswaye H, Karar M, Omar S, Younes E, Younes M, Zreeg D, Abujamra S, Ashour F, Elgammudi M, Aljadidi WOF, Saddouh E, Sharif R, Alabuzidi A, Alwerfally A, Aribi S, Bibas F, Elfaituri T, Elhajjaji Y, Khaled A, Khalil W, Layas T, Soula E, Tarek A, Hallalah M fathi khalleefah A, Abujamra S, Ahmed HA, Alsharef T, Saoud AAB, Gharmoul TE, Elhadi A, Elrais S, Shebani A, Zarti H, Zeiton A, Ambrazevicius M, Kaselis N, Stakyte M, Aliosin O, Cizauskaite A, Dailidenas S, Eismontas V, Kybransiene M, Nutautiene V, Samalavicius N, Simcikas D, Slepavicius A, Tamosiunas A, Ubartas N, Zeromskas P, Bradulskis S, Dainius E, Juocas J, Kubiliute E, Kutkevicius J, Opolskis A, Parseliunas A, Subocius A, Venskutonis D, Virbickaite E, Zuikyte D, Bogusevicius A, Buzaite K, Cepuliene D, Cesleviciene I, Cesna V, Gribauskaite J, Ignatavicius P, Jokubauskas M, Liugailaite M, Margelis E, Mazelyte R, Pankratjevaite L, Pažusis M, Rackeviciute A, Saladyte J, Škimelyte M, Šlenfuktas V, Sudeikyte M, Tamelis A, Vanagas T, Žumbakys Ž, Atkociunas A, Dulskas A, Kuliavas J, Birutis J, Paškevicius S, Šatkauskas M, Danys D, Jakubauskas M, Jakubauskiene L, Kryzauskas M, Lipnickas V, Makunaite G, Rasoaherinomenjanahary F, Rasolofonarivo H, Samison LH, Banda B, Malemia P, Msosa V, Izzuddin AIA, Das A, Gan YY, Sheng TS, Siaw J yng, Rahim MFA, Jamari DZHA, Husin NC, Kamarulzaman MY, Lim YP, Kamil NAM, Hassan MRM, Sahid SM, Mustafa J, Ng EHB, Khazim WKW, Ern NC, Lingeshan P g, Sulaiman SE, Ang SE, Sithik MNBM, Cheong YJ, Tata MD, Xian LJ, Kadravello A, Koh I-E, Ng L-Y, Yong YJNW, Palayan K, Sam CX, Jin PS, Hwei JTE, Tang Y, Ter AZ, Wong MP-K, Zakaria AD, Zakaria Z, Henry F, Kalaiselvan T, Karim MFSA, Aziz MRA, Aziz NA, Khong TL, Lau PC, Lim HC, Roslani AC, Seak JCK, Wong S-W, Wong LF, Chin LY, Anyanwu MC, Borg E, Busuttil Z, Calleja T, Chircop KL, Cutajar R, Dimech AM, Ellul S, Galea J, Perai KG, Gatt R, Kelman L, Micallef E, Nwolu F, Sammut K, Thompson J, Warwicker S, Zammit M, Cordera F, González EC, Sánchez-García J, Camacho FJB, López FJB, Valle CJZF del, Acosta E, Espinoza IRG, Moreno P, Cortes-Flores AO, Orozco CF, Ojeda AG, González SCD, Martinez L, Medina AR-D la, Amador BM, Novoa A, Espejo DAO, Jimenez A, Rosales FL, Vanoye EG, Gonzalez LAG, Miranda-Ackerman RC, Solano-Genesta M, Alvarez-Cano A, Romero-Garza HH, Medina-Franco H, Mejía-Fernández L, Salgado-Nesme N, Vergara-Fernandez O, Gutiérrez-Mota GM, Vera FXH, Lopez AL, Villela GM, Padilla F de JR, Marin WT, Maldonado MM, Suárez RS, Troche JM, Benyaiche C, Outani O, Amine S, Benkabbou A, Majbar AM, Mohsine R, Rafik A, Oung T, Tin MM, Borowski DW, Plarre P, Borowski DW, Plarre P, Alberga A, Sluiter N, Tuynman J, Blok R, Cömert D, Hompes R, Kalff M, Stellingwerf ME, Tanis P, Henegouwen M van B, Praag EM van, Wisselink D, Gerhards M, Cardozo JL, Westerduin E, Jonge J de, Geloven A van, Schilt K van, Boer F den, Stoots S, Vlek S, Adams J, Al-Busaidi IS, Budd G, Choi S il, Chu MJJ, Ganugapati A, McKinstry L, Pascoe R, Richards S, Rosser K, Stevenson A, White R, Farik S, Kwun J, Murad A, Cowan S, Hall T, Hayton M, Sani LM, Garba SO, Adamou H, Magagi IA, Habou O, Aliyu H, Daniyan M, Sholadoye TT, Abdullahi L, Anyanwu L-J, Mohammad AM, Muhammad AB, Sheshe AA, Suleiman I, Adesina A, Awolowo A, Onuoha C, Salami O, Taiwo O, Taiwo A, Kache S, Makama JG, Sale D, Abiola O, Ajao A, Ajiboye A, Etonyeaku A, Olaogun J, Adebanjo A, Adesanya O, Afolayan MO, Balogun O, Makanjuola A, Nwokocha S, Ojewola RW, Olajide TO, Aderounmu A, Adesunkanmi A-R, Adisa A, Agbakwuru A, Aderogba AA, Alatise OI, Arowolo O, Lawal O, Mohammed T, Ndegbu C, Olasehinde O, Wuraola F, Akinkuolie A, Etonyeaku A, Mosanya A, Ayandipo O, Elemile P, Lawal TA, Sani SA, Garba S, Sani RH, Olori S, Onyebuashi H, Umoke I, Adenuga A, Adeyeye A, Habeeb O, Lawal B, Nasir A, Aahlin EK, Kjønås D, Myrseth E, Abbasy J, Alvi A, Saleem O, Afzal A, Nazir A, Farooq M, Liaqat A, Naqi SA, Raza A, Sarfraz M, Sarwar M, Banglani M, Munir A, Sehrish R, Ayub B, Sayyed R, Altaf A, Ayub S, Qureshi AU, Saeed K, Syed B, Akbar SA, Anwer AW, Khan RN, Khan AI, Khattak S, Mohtasham S, Parvaiz MA, Syed AA, Ansari AB, Shahzad N, Khaliq T, Rashid I, Waqar SH, Al-saleem HA, Alqumboz AA, Alqadi M, Amro A, Assa R, Awesat E, Ayyad R, Hammad M, Haymony A, Hijazi B, Hmeidat B, Lahaseh R, Qawasmi A, Rajabi A, Shehada M, Shkokani S, Yaghi Y, Yaghi N, AlZohour M, Farid M, Habes YM, Juba W, Nubani Y, Rabee A, Sa’deh M, Abed S, Basos IA, Alswerki M, Ashour D, Awad I, Diab S, Jamassi AE, El-Kahlout S, Elhout S, Hajjaj ANK, Hasanain D, Hajjaj BN, Obaid M, Saikaly E, Salhi A, Al-Tammam H, Almasri M, Baniowda M, Beshtawi D, Horoub A, Misk R, Mohammad B, Qasrawi R, Sholi T, Abu-Nimeh S, Abu-srour A, Abukhalaf SA, Adawi S, Alsalameh B, Ayesh K, Elqadi M, Hammouri A, Mustafa FK, Marzouqa N, Melhem S, Miqdad D, Mohamad B, Rawhi M, Ahammala ABA, Ataya AA, Jayyab IA, Al-Shwaikh S, Alagha O, Alasttal M, Awadallah H, Elblbessy M, Fares J, Jarbou A, Mahfouz I, Albahnasawi MA, Mahadi AA, Abuelhatal H, Abuelqomboz A, Almoqayyad A, Alwali A, Balaawi R, Hamouda M, Humeid M, Jedyan A, Hamam TMA, Matar G, Salem A, Samra T, Shaheen N, Shihada K, A.Nemer A, Amrain MAA, Alamrain AA, Jamie NA, Abu-Rous MR, Alfarra N, AlTaweel M, Alwhaidi N, Hamed R, Saqqa B, Shaheen A, Aljaber D, Aljaberi L, Alwaheidi M, Jawaada A, Khaldi H, Qahoush R, Qari J, Saadeh R, Salim A, Yacoub A, Abbas A, Shua`ib RA, Zainah BA, AbuSirrees M, Babaa B, Barhoush O, Qadomi AB, Daraghmeh L, Haji R, Khatatbeh A, Khatib L, Qarariah S, Quzmar Y, Safadi K, Salameh R, Hassan M, Herzallah S, Massad L, Nazzal A, Nazzal R, Escobar D, V GMM, Gonzalez AR, Mar JEC, Paredes NOC, Cuba V, Lopez W, Jimenez MMN, Bartra NAS, Ojeda OS, Sequeiros D, Pacheco AT, Vergara M, Abarca S, Alcorta R, Borda-Luque G, Zegarra IEE, López CL, Marrufo M, Mogrovejo C, Nomura A, Angeles YR, Meza MRV, Zavala G, Arrascue JNC, Cabrera JCH, Vera JJML, Osorio M, Díaz EAY, Fontanilla MA, Fuentes JR, Salazar AL, Dominguez G, Lopez MP, Macalindong S, Onglao MA, Ramirez A, Sacdalan MD, Tampo MM, Uy GL, Mangahas J, Yabut K, Cañete JP, Cansana BE, Castro EJ, Lipana MK, Roxas MF, Zara VJ, Chrol M, Franczak P, Orlowski M, Budzynski P, Budzynski A, Bury P, Czerwinska A, Dworak J, Dziedzic J, Kisielewski M, Kulawik J, Lasek A, Major P, Malczak P, Migaczewski M, Pedziwiatr M, Pisarska M, Radkowiak D, Rubinkiewicz M, Rzepa A, Skoczylas T, Stanek M, Truszkiewicz K, Wierdak M, Winiarski M, Zarzycki P, Zub-Pokrowiecka A, Kowalewski P, Roszkowski R, Waledziak M, Tomé M, Patrocinio S, Guerreiro I, Almeida F, Sousa X de, Monteiro N, Santos MTC, Oliveira D de, Serra ML, Morgado D, Neves C, Oliveira AC, Pimentel A, Silva S, Carvalho M, Carvalho L, Magalhães J, Matos L, Monteiro T, Ramos C, Santos V, Barbosa J, Costa-Maia J, Devezas V, Fareleira A, Fernandes C, Gonçalves D, Mora H, Morais M, Sousa FS de, Santos SC, Logrado A, Tojal A, Amorim E, Cunha MF, Fazenda A, Neves JPM, Miguel IIS da NG, Veiga D, Azevedo J, Louro HC, Leite M, Azevedo J, Menezes MB, Gama B, Brito D, Martins MCC, Magalhães AG e, Longras AC, Lourenço R, Matos D, Castro L, Policarpo F, Romano J, Leite M, Monteiro C, Pinto D, Duarte M, Martins SF, Oliveira M, Galvão D, Martins L, Silva A, Taranu V, Vieira B, Neves J, Oliveira S, Ribeiro H, Cinza M, Felix R, Machado A, Oliveira J, Patrício J, Lima RP de, Pereira M, Melo MR, Velez C, Silva AA da, Claro M, Santos DC, Ferreira A, Capote H, Rosado D, Taré F, Nogueira O, Ângelo M, Baiao JM, Guimarães A, Marques J, Albano MN, Silva M, Costa AV da, Caroço TV, Braga SA, Capunge I, Fragoso M, Guimarães J, Pinto B, Ribeiro J, Angel M, Fialho G, Guerrero M, Costa FC, Cardoso D, Cardoso V, Alves M, Estalagem I, Louro T, Marques C, Martelo R, Morgado M, Canotilho R, Correia AM, Martins P, Peyroteo M, Gomes J, Monteiro R, Romano M, Alves DM, Peixoto R, Quintela C, Jervis MJ, Melo D, Pacheco A, Paixão V, Pedro V, Pimenta J, Castro JP de, Rocha A, Beuran M, Bratu MR, Ciubotaru C, Diaconescu B, Hostiuc S, Negoi I, Stoica B, Na N, Anokhin E, Kuznetsov G, Oganezov G, Paramzin F, Romanova E, Rutkovskii V, Rutkovskii V, Shushval M, Zabiyaka M, Dzhumabaev K, Ivanov V, Mamedli Z, Achkasov S, Balkarov A, Nabiev E, Nagudov M, Rybakov E, Saifutdinova K, Sushkov O, Vardanyan A, Costas-Chavarri A, Joseph L, Ndayishimiye I, Ingabire JA, Faustin N, Mutabazi AZ, Mvukiyehe JP, Nsengimana VJP, Uwakunda C, Abbas MM, Akeel N, Aljiffry M, Awaji K, Farsi A, Jamjoum G, Khoja A, Maghrabi A, Malibary N, Nassif M, Saleem A, Sultan A, Tashkandi W, Tashkandi H, Trabulsi N, Ba MB, Diallo AC, Ndong A, Cuk V, Jankovic U, Juloski J, Koh SZ, Koh F, Lee KC, Lee KY, Lee S, Leong WQ, Lieske B, Lui SA, Prakash P, Grosek J, Norcic G, Tomazic A, Fitchat N, Jaich R, Wineberg D, Koto MZ, Baiocchi D, Clarke D, Steenkamp CJ, Straten SV, Bannister S, Boutall A, Chinnery G, Coccia A, Dell A, Karjiker P, Kloppers C, Loxton N, Mabogoane T, Malherbe F, Panieri E, Rayamajhi S, Spence R, Wyngaard T van, Warden C, Madiba TE, Moodley Y, Pillay N, Brooks S, Kruger C, Merwe LHVD, Gool F, Kariem M, Bougard H, Chu K, Kariem N, Noor F, Pillay R, Steynfaardt L, González LG, Santos JMM, Martín-Borregón P, Caballero JM, García CN, Fraga PR, Parga GDC, Veiga MPF, López LG, Pino HI, Cal IL, Otero ML, Sixto MN, Señorans MPG, Fernández LR, Poblador AR, Crespo ER, Sanchez-Santos R, Vigorita V, Batanero EA, Asnel D, Canales IC, Saiz EC, Alvarez IDS, Vico TD, Arias SF, Martínez DF, Bernardo CG, Flórez LJG, Gutierrez CG, Munar MG, Molina CAMZ, Merayo M, Campos JLM, Gijon MM, Otero-Diez JL, Miravalles JLR, Solar-Garcia L, Sánchez AS, Truan N, Villalobos CA, Díaz YC, Jimenez M, Montesdeoca D, Navarro-Sánchez A, Vega V, Heredia JB de, Gómez Z, Jezieniecki C, Morán APL, Montes-Manrique M, Rodriguez-Lopez M, Soriano MR, Díaz JT, Fernandez AV, Argudo N, Pera M, Jansà LT, Domínguez MG, Goded I, Golet MR, El-Abur IT, Fornals AU, Campos VZ, Martinez MDMA, Bosch M, García-Catalá L, Sánchez-Guillén L, Artigau E, Romeu NG, Bergkvist DJ, Perez BE, Morató O, Olona C, Diéguez B, Forero-Torres A, Losada M, Gomez-Abril S, Gonzálvez P, Martinez R, Martínez SN, Payá-Llorente C, Rubio ÁP, Martinez SS, Tomás JCS, Juan RT, Simón AG, Maté P, Prieto-Nieto MI, Rubio-Perez I, Urbieta A, Bravo MV, Abelló D, Frasson M, Garcia-Granero A, Gurumeta AA, Abad-Motos A, Pablo EL, Nozal B, Ripollés-Melchor J, Salvachúa R, Ferrero E, Tellez LG-S, Vázquez IO, Picardo AL, López JAR, Matilla LPZ, Fernandez CC, Bezanilla SC, Tesouro JE, Fernandez-Diaz MJ, Cardo JG, Ruiz MG, Gonzalez-Tolaretxipi E, Fraile JJ, Poch C, Rodriguez-Aguirre M, Pesqueira NT, Trugeda-Carrera MS, Torre J de la, Blanco-Colino R, Espin-Basany E, Espinosa-Bravo M, Comas CM, Afonso ER, Déniz JR, Raber CS, Tremolosa MV, Chandrasinghe P, Kumarage S, Arachchilage NW, Senanayake KJ, Elkamel AAA, Adam MA, Saleh M, Blomme N, Thorell A, Wogensen F, Älgå A, Ansarei D, Celebioglu F, Heinius G, Nigard L, Pieniowski E, Ahlqvist S, Björklund I, Cengiz Y, Frånberg A, Håkansson M, Adamo K, Franklin O, Sund M, Wiberg R, Andersson Y, Chabok A, Nikberg M, Kugelberg A, Canonica C, Christoforidis D, Fasolini F, Gaffuri P, Giuliani M, Meani F, Popeskou SG, Pozza S, Wandschneider W, Peterer L, Widmer LW, Zimmermann B, Bakoleas P, Chanousi I, Charalampidou L, Grochola LF, Heid F, Ntaoulas S, Outos M, Peros G, Podolska-Skoczek H, Reinisch KB, Zielasek C, Clerc D, Demartines N, Gilgien J, Kefleyesus A, St-Amour P, Toussaint A, Alhimyar M, Alsaid B, Alyafi A, Alkhaledi A, Kouz B, Omarain A, Al-Sabbagh Y, Alkhatib H, Sara S, Alhaj A, Danial A, Kadoura L, Albared SM, Monawar Y, Nahas L, Abd B, Saad A, Wakkaf H, Bouaziz H, Bouzaiene H, Ghalleb M, Akaydin E, Akbaba AC, Atakul O, Baltaci E, Besli S, Burgu G, Cenal U, Muijnck C de, Demirkaya HC, Dogruoz A, Gezer ZI, Gündogdu Y, Kara M, Korkmaz HK, Kurtoglu GK, Ozben V, Ozmen BB, Pektas AM, Sel EK, Yenidünya N, Bengur FB, Oral BM, Yozgatli TK, Abdullayev S, Gunes ME, Sahbaz NA, Banaz T, Kargici K, Kuyumcu OF, Yanikoglu E, Yesilsancak M, Yilmaz D, Aktas MK, Rencuzogullari A, Isik A, Leventoglu S, Yalçinkaya A, Yüksel O, Kalayci MU, Kara Y, Sarici IS, Akin A, Alemdag G nur, Arslan E, Baki BE, Bodur MS, Calik A, Altinbas BC, Cihanyurdu I, Erkul O, Gül B, Guner A, Köse B, Semiz A, Sevim S, Tayar S, Tomas K, Tüfek O yavuz, Türkyilmaz S, Ulusahin M, Usta A, Yildirim R, Güler SA, Tatar OC, Varol E, Kirimtay B, Uysal M, Yildiz A, Kose E, Ciftci AB, Çolak E, Eraslan H, Kucuk GO, Yemez K, Lule H, Bienfait M, Lule H, Bua E, Doe M, Okalany N, Birindelli A, Basarab M, Bielosludtsev O, Freigofer M, Kolhanova K, Perepelytsia K, Romanukha K, Savenkov D, Siryi S, Tereshchenko M, Viacheslav N, Volovetskyi A, Kebkalo A, Tryliskyy Y, Tyselskiy V, Bruce E, Chow BL, Iddles E, McGuckin S, Newall N, Ramsay G, Sharma P, Stewart C, Wong J, Badran A, Bath M, Belais F, Butt E, Joshi K, Kapur M, Shaw M, Townson A, Williams CYK, Gray T, Greig R, Husain M, Murray E, Mustafa A, Asif A, Gokul A, Shah M, Akitikori MT, Charalabopoulos A, Davidson S, McNally S, Rupani S, Juma F, Mills SC, Muirhead L, Sellars K, Walsh U, Warren O, Chambers A, Hunt R, Teasdale E, Boyce S, Cornwall H, Tol I, Argyriou EO, Eardley N, Povey M, Aithie JMS, Irfan A, McGuigan M-C, Starr R, Warren CR, Archibald J, Kirby G, Kisyov I, Khoo CK, Lee R, Photiou D, Davis R, Prasad U, Yang PZ, Bird J, Leung E, Summerour V, Currow C, Kiam J, Tan GJS, Muthusami A, Pegba-Otemolu I, Urbonas T, Nunoo-Mensah J, Smolskas E, Boddy A, Gravante G, Hunter D, Andrew D, Koh A, Thompson A, Adams L, Clements HA, Silva KD, Ekpete O, Haque S, Henderson S, Ibrahim B, Jayasinghe T, Livie J, Mailley K, Nair G, Tan D, Baggaley C, Dawidziuk A, Szyszka B, Barter C, Gandhi N, Hassell K, Hitchin S, Kelsall J, Nagy E, Nessa A, Whisker L, Yanni F, Ali M, Arora D, Hediwattege S, Kumarasinghe N, Rathore M, Tennakoon A, Ahmad SMA, Bajomo O, Nadira F, Celentano V, Bhangu A, Glasbey J, Griffiths E, Karri RS, Mak JKC, Nepogodiev D, Pipe M, Bhatti MI, Rabie M, Boyle C, Hamilton D, Mihuna A, Ng JCK, Nicholson G, Oliwa A, Pearson R, Rose A, Yong SQ, Boereboom C, Hanna M, Walter C, Greensmith TS, Mitchell R, Monaghan E, Crawford J, Moug S, Blackwell J, Boyd-Carson H, Herrod P, Al-Allaf O, Beattie M, Bullock C, Burman S, Clark G, Flamey N, Flannery O, Harding A, Kodiatt B, Lawday S, Mahapatra S, Nagesh NM, Ng M, Rye D, Yoong A, Clark L, Deans C, Edirisooriya M, Fairfield CJ, Harrison EM, Carrington EV, Wong TLE, Yusuf B, Chamberlain C, Duke K, Kmiotek E, Botes A, Condie N, Schrire T, Shah R, Thomas-Jones I, Yates C, Anthony N, Matthews E, Sahnan K, Tankel J, Tucker S, Beatty JW, Ziprin P, Duggan W, Kantartzi A, Sridhar S, Khaw RA, Srivastava P, Underwood C, Brum HA do C, Chopra S, Davis L, Hughes R, Tulley J, Alberts J, Athisayaraj T, Olugbemi M, Ahmad K, Chan C, Chapman G, Fleming H, Fox B, Grewar J, Hulse K, Rutherford D, Sinead M, Smith S, Speake D, Vaughan-Shaw PG, Christodoulides N, Kudhail S, Welch M, Husaini SM, Lambracos S, Anyanwu C, Suresh R, Thomas JS, Gleeson E, Platoff R, Saif A, Enumah Z, Etchill E, Gabre-Kidan A, Bernstein M, Carrano FM, Connors J, Lynn P, Melis M, Newman E, Foster DS, Perrone K, Titan A, Weiser TG, Ahmad S, Bafford ACMD, Molin MD, Hanna N, Zafar SN, Hemmila M, Napolitano L, Wong JJ, Chandler J, Wood L, Wren S, Ottesen T, You L, Yu K, Yañez M del pilar A, Fernandes MF, González D, Cubas S, González MC, Zubiaurre V, Demolin R, Giroff N, Sciuto P, Campos M, Cantera GR, Wagner G, Deepika G, Maimbo M, Simuchimba E, Bulaya A, Chibuye C, Chirengendure B, Kabale M-R, Kabongo K, Linyama D, Munthali J, Mweso O, Pikiti F, Otieno J, Chan E, Lai LT, Blackman B, Richards S, Subramaniam S, Karim R, Kok N, Lee YD, Ali S, Sinha A, Corrigan R, Barnes N, Wong F, Dennis G, Jedamzik J, Phillips E, Piette W, Hentenryck MV, Koco H, Lawani S, Kassa MW, Bezerra TS, Gribnev P, Dimitrov D, Krastev P, Oum S, Bonghaseh DT, Farsi MA, Alsharqawi N, Agarwal A, Acevedo V, Barbosa ACC, Giron F, Rodriguez JPL, Kucan D, Rosko D, Barsic N, Župan D, Hegazi A, Truncíková V, Fryba V, Mohamed M, Sultan A, Nagi A, Temerik AR, Elshawy ME, Mahmoud MI, Omar S, Anwar M, Rageh T, Elmokadem A, Gaballa K, Teppo S, Turunen A, Pengermä P, Ballouhey Q, Bergeat D, Weyl A, Hain E, Gyedu A, Yenli E, Osei-Poku D, Rompou V-A, Zoikas A, Gaitanidis A, Koukis G, Perivoliotis K, Tavlas P, Galanos-Demiris K, Zografos G, Karavokyros I, Xanthopoulou G, Iordanidou E, Ayau F, Garcia A, Damján P, Wason D, L AB, Rangganata E, Kamath P, O’Connor DB, Pinto M, Perrone F, Tropeano FP, Troilo F, Bossi D, Scala D, Pulitanò L, Carella M, Pietrabissa A, Gori A, Giraudo G, Simone VD, Russo AA, Braccio B, Al-Taher R, Athamneh S, Parker A, Sawiee A, Kattia A, Salem M, Tababa O, Shaeeb Z, Syminas V, Jurgaitis J, Damuleviciene G, Svagzdys S, Poskus T, Razafimanjato NNM, Loo LC, Tiong IC, Muhmad WFW, Vijeyan H, Ying TL, Grech G, Arrangoiz R, Ley VBJ, Arizpe D, Ley VBJ, Lara EL, López EVC, Eaazim J, Gouberville MG de, Bastiaenen V, Rottier S, Nahab F, Ji MY, Seyoji M, Nwachukwu C, Emeghara O, Muhammed SE, Idowu A, Sowemimo O, Ogundoyin O, Akande O, Lott A, Nadeem M, Laghari AA, Loya A, Mushtaq H, Abdullah MT, Abuhilal B, Atawneh M, Hamdan H, Alhabil B, Srour A, Mousa I, Medina LDS, Sacdalan MD, Lapitan MC, Sacdalan MD, Sacdalan MD, Bartosiak K, Ferreira P, Francisco V, Lemos R, Frutuoso L, Fernandes S, Fonseca T, Pereira J, Rachadell J, Torre A, Martins FM, Carvalho AC, Ferreira JR, Silva BR da, Devesa H, Vieira A, Mónica I, Amaro M, Sousa D, Reia M, Louro J, Martins A, Dominguez J, Santos I, Oliveira NMF, Pereira JC, Silva-Vaz P, Freire L, Escrevente R, Negoita VM, Shakhmatov D, Nezerwa Y, Radulovic R, Moore R, Obery G, Viljoen F, Mendes T, Suarez A, Moncada E, Fernandez-Hevia M, Martínez CC, Garcia JMG, Zunzarren MG, Idris T, Eklöv K, Grahn O, Amin L, Blomqvist M, Ajani C, Kraus R, Seeger N, Willemin M, Rayya F, Ayash M, Msouti R, Kannas I, Abazid E, Esper A, Slim S, Kavcar AS, Aytac E, Dural AC, Ilker A, Eray IC, Kurnaz E, Altiner S, Tepe MD, Sahin C, Savli E, Innocent A, Babirye L, Diachenko A, Hordoskiy V, Curry H, Chau CYC, Robertson H, Mahmoud A, Lennon H, Loi L, Kirkham E, McCann C, Watts D, Gurung B, Wilson M, Tribedi T, Garofalo E, Zahra B, MacDonald S, Daniels I, Ng N, Khosla S, Olivier J, Yue SYP, Suresh G, Wellington J, Lorejo E, Mossaad M, Tryliskyy Y, Crutcher M, Alimi M, Baiu I, Abdou H, Conway A, Peck C, Wagner G, Perez MAP, Trostchansky I, Zulu S, Nakazwe M, Knight SR, Drake TM, Nepogodiev D, Fitzgerald JE, Ademuyiwa A, Alexander P, Ingabire JCA, Al-Saqqa SW, Biccard BM, Borda-Luque G, Borowski DW, Burger S, Chu K, Clarke D, Costas-Chavarri A, Davies J, Donaldson R, Ede C, Garden OJ, Ghosh D, Glasbey J, Kingham TP, Salem HK, Anyomih TTK, Koto MZ, Lapitan MC, Lawani I, Lesetedi C, Aguilera-Arevalo M-L, Mabedi C, Maimbo M, Magill L, Alakaloko FM, Makupe A, Martin J, Medina AR-D la, Monahan M, Moore R, Msosa V, Mulira S, Mutabazi AZ, Muller E, Musowoyo J, Adisa AO, Olory-Togbe JL, Pius R, Qureshi AU, Rayne S, Roberts T, Sacdalan MD, Shaw CA, Smart N, Smith M, Spence R, Straten SV, Tabiri S, Tayler V, Weiser TG, Windsor J, Yorke J, Yepez R, Lilford R, Morton D, Bhangu A, Sundar S, Harrison EM, Runigamugabo E, Verjee A, Chen J, Daya L, Aroussi NE, Farina V, Olivier TG, Nacarino MG, Hammani A, Honjo S, Jacobs R, Kimura H, Litvin A, Nkoronko M, Nour I, Yepez JJO, Pagano G, Pata F, Hung WP, Raj A, Pozo AR, Rommaneh M, Fabiano SCS, Gago CMS, Yip SS, Srinivas A, Sung C-Y, Tai A, Aranda YCV, Venturini S, Vervoort D, Lartigue JW (2021) Global variation in postoperative mortality and complications after cancer surgery: a multicentre, prospective cohort study in 82 countries. The Lancet 397:387–397. https://doi.org/10.1016/S0140-6736(21)00001-5
Bohnen JD, Mavros MN, Ramly EP, Chang Y, Yeh DD, Lee J, de Moya M, King DR, Fagenholz PJ, Butler K, Velmahos GC, Kaafarani HMA (2017) Intraoperative adverse events in abdominal surgery: what happens in the operating room does not stay in the operating room. Ann Surg 265:1119–1125. https://doi.org/10.1097/SLA.0000000000001906
Zimmitti G, La Mendola R, Manzoni A, Sega V, Malerba V, Treppiedi E, Codignola C, Monfardini L, Garatti M, Rosso E (2021) Investigation of intraoperative factors associated with postoperative pancreatic fistula following laparoscopic left pancreatectomy with stapled closure: a video review-based analysis : video-review for predictors of pancreatic leak. Surg Endosc 35:941–954. https://doi.org/10.1007/s00464-020-07912-x
Birkmeyer JD, Finks JF, O’Reilly A, Oerline M, Carlin AM, Nunn AR, Dimick J, Banerjee M, Birkmeyer NJO (2013) Surgical skill and complication rates after bariatric surgery. N Engl J Med 369:1434–1442. https://doi.org/10.1056/NEJMsa1300625
Goldenberg MG, Goldenberg L, Grantcharov TP (2017) Surgeon performance predicts early continence after robot-assisted radical prostatectomy. J Endourol 31:858–863. https://doi.org/10.1089/end.2017.0284
Curtis NJ, Foster JD, Miskovic D, Brown CSB, Hewett PJ, Abbott S, Hanna GB, Stevenson ARL, Francis NK (2020) Association of surgical skill assessment with clinical outcomes in cancer surgery. JAMA Surg 155:590. https://doi.org/10.1001/jamasurg.2020.1004
Karliczek A, Harlaar NJ, Zeebregts CJ, Wiggers T, Baas PC, van Dam GM (2009) Surgeons lack predictive accuracy for anastomotic leakage in gastrointestinal surgery. Int J Colorectal Dis 24:569–576. https://doi.org/10.1007/s00384-009-0658-6
Hasin Y, Seldin M, Lusis A (2017) Multi-omics approaches to disease. Genome Biol 18:83. https://doi.org/10.1186/s13059-017-1215-1
Mayerhoefer ME, Materka A, Langs G, Häggström I, Szczypiński P, Gibbs P, Cook G (2020) Introduction to radiomics. J Nucl Med 61:488–495. https://doi.org/10.2967/jnumed.118.222893
Maier-Hein L, Vedula SS, Speidel S, Navab N, Kikinis R, Park A, Eisenmann M, Feussner H, Forestier G, Giannarou S, Hashizume M, Katic D, Kenngott H, Kranzfelder M, Malpani A, März K, Neumuth T, Padoy N, Pugh C, Schoch N, Stoyanov D, Taylor R, Wagner M, Hager GD, Jannin P (2017) Surgical data science for next-generation interventions. Nat Biomed Eng 1:691–696. https://doi.org/10.1038/s41551-017-0132-7
Corey KM, Kashyap S, Lorenzi E, Lagoo-Deenadayalan SA, Heller K, Whalen K, Balu S, Heflin MT, McDonald SR, Swaminathan M, Sendak M (2018) Development and validation of machine learning models to identify high-risk surgical patients using automatically curated electronic health record data (Pythia): a retrospective, single-site study. PLoS Med. https://doi.org/10.1371/journal.pmed.1002701
Garrow CR, Kowalewski K-F, Li L, Wagner M, Schmidt MW, Engelhardt S, Hashimoto DA, Kenngott HG, Bodenstedt S, Speidel S, Müller-Stich BP, Nickel F (2021) Machine learning for surgical phase recognition: a systematic review. Ann Surg 273:684–693. https://doi.org/10.1097/SLA.0000000000004425
Junger D, Frommer SM, Burgert O (2022) State-of-the-art of situation recognition systems for intraoperative procedures. Med Biol Eng Comput 60:921–939. https://doi.org/10.1007/s11517-022-02520-4
Lavanchy JL, Zindel J, Kirtac K, Twick I, Hosgor E, Candinas D, Beldi G (2021) Automation of surgical skill assessment using a three-stage machine learning algorithm. Sci Rep 11:5197. https://doi.org/10.1038/s41598-021-84295-6
Bhandari M, Nallabasannagari AR, Reddiboina M, Porter JR, Jeong W, Mottrie A, Dasgupta P, Challacombe B, Abaza R, Rha KH, Parekh DJ, Ahlawat R, Capitanio U, Yuvaraja TB, Rawal S, Moon DA, Buffi NM, Sivaraman A, Maes KK, Porpiglia F, Gautam G, Turkeri L, Meyyazhgan KR, Patil P, Menon M, Rogers C (2020) Predicting intra-operative and postoperative consequential events using machine-learning techniques in patients undergoing robot-assisted partial nephrectomy: a Vattikuti Collective Quality Initiative database study. BJU Int 126:350–358. https://doi.org/10.1111/bju.15087
Jung JJ, Jüni P, Lebovic G, Grantcharov T (2020) First-year analysis of the operating room black box study. Ann Surg 271:122–127. https://doi.org/10.1097/SLA.0000000000002863
Jung JJ, Jüni P, Gee DW, Zak Y, Cheverie J, Yoo JS, Morton JM, Grantcharov T (2020) Development and evaluation of a novel instrument to measure severity of intraoperative events using video data. Ann Surg 272:220–226. https://doi.org/10.1097/SLA.0000000000003897
Maier-Hein L, Eisenmann M, Sarikaya D, März K, Collins T, Malpani A, Fallert J, Feussner H, Giannarou S, Mascagni P, Nakawala H, Park A, Pugh C, Stoyanov D, Vedula SS, Cleary K, Fichtinger G, Forestier G, Gibaud B, Grantcharov T, Hashizume M, Heckmann-Nötzel D, Kenngott HG, Kikinis R, Mündermann L, Navab N, Onogur S, Roß T, Sznitman R, Taylor RH, Tizabi MD, Wagner M, Hager GD, Neumuth T, Padoy N, Collins J, Gockel I, Goedeke J, Hashimoto DA, Joyeux L, Lam K, Leff DR, Madani A, Marcus HJ, Meireles O, Seitel A, Teber D, Ückert F, Müller-Stich BP, Jannin P, Speidel S (2022) Surgical data science - from concepts toward clinical translation. Med Image Anal 76:102306. https://doi.org/10.1016/j.media.2021.102306
Lam K, Abràmoff MD, Balibrea JM, Bishop SM, Brady RR, Callcut RA, Chand M, Collins JW, Diener MK, Eisenmann M, Fermont K, Neto MG, Hager GD, Hinchliffe RJ, Horgan A, Jannin P, Langerman A, Logishetty K, Mahadik A, Maier-Hein L, Antona EM, Mascagni P, Mathew RK, Müller-Stich BP, Neumuth T, Nickel F, Park A, Pellino G, Rudzicz F, Shah S, Slack M, Smith MJ, Soomro N, Speidel S, Stoyanov D, Tilney HS, Wagner M, Darzi A, Kinross JM, Purkayastha S (2022) A Delphi consensus statement for digital surgery. Npj Digit Med 5:100. https://doi.org/10.1038/s41746-022-00641-6
Gibaud B, Forestier G, Feldmann C, Ferrigno G, Gonçalves P, Haidegger T, Julliard C, Katić D, Kenngott H, Maier-Hein L, März K, de Momi E, Nagy DÁ, Nakawala H, Neumann J, Neumuth T, Rojas Balderrama J, Speidel S, Wagner M, Jannin P (2018) Toward a standard ontology of surgical process models. Int J Comput Assist Radiol Surg 13:1397–1408. https://doi.org/10.1007/s11548-018-1824-5
Bodenstedt S, Rivoir D, Jenke A, Wagner M, Breucha M, Müller-Stich B, Mees ST, Weitz J, Speidel S (2019) Active learning using deep Bayesian networks for surgical workflow analysis. Int J Comput Assist Radiol Surg 14:1079–1087. https://doi.org/10.1007/s11548-019-01963-9
Heinze O, Birkle M, Köster L, Bergh B (2011) Architecture of a consent management suite and integration into IHE-based regional health information networks. BMC Med Inform Decis Mak 11:58. https://doi.org/10.1186/1472-6947-11-58
Vassiliou MC, Feldman LS, Andrew CG, Bergman S, Leffondré K, Stanbridge D, Fried GM (2005) A global assessment tool for evaluation of intraoperative laparoscopic skills. Am J Surg 190:107–113. https://doi.org/10.1016/j.amjsurg.2005.04.004
Harris A, Butterworth J, Boshier PR, MacKenzie H, Tokunaga M, Sunagawa H, Mavroveli S, Ni M, Mikhail S, Yeh C-C, Blencowe NS, Avery KNL, Hardwick R, Hoelscher A, Pera M, Zaninotto G, Law S, Low DE, van Lanschot JJB, Berrisford R, Barham CP, Blazeby JM, Hanna GB (2020) Development of a Reliable Surgical Quality Assurance System for 2-stage Esophagectomy in Randomized Controlled Trials. Ann Surg Publish Ahead of Print: https://doi.org/10.1097/SLA.0000000000003850
Van Rossum G, Drake FL (2009) Python 3 Reference Manual. CreateSpace, Scotts Valley, CA
Shapiro SS, Wilk MB (1965) An analysis of variance test for normality (complete samples). Biometrika 52:591–611. https://doi.org/10.1093/biomet/52.3-4.591
Welch BL (1938) The significance of the difference between two means when the population variances are unequal. Biometrika 29:350–362. https://doi.org/10.2307/2332010
Forrest JA, Finlayson ND, Shearman DJ (1974) Endoscopy in gastrointestinal bleeding. Lancet Lond Engl 2:394–397. https://doi.org/10.1016/s0140-6736(74)91770-x
Takahashi H, Yamasaki M, Hirota M, Miyazaki Y, Moon JH, Souma Y, Mori M, Doki Y, Nakajima K (2013) Automatic smoke evacuation in laparoscopic surgery: a simplified method for objective evaluation. Surg Endosc 27:2980–2987. https://doi.org/10.1007/s00464-013-2821-y
Moccia S, Wirkert SJ, Kenngott H, Vemuri AS, Apitz M, Mayer B, De Momi E, Mattos LS, Maier-Hein L (2018) Uncertainty-aware organ classification for surgical data science applications in laparoscopy. IEEE Trans Biomed Eng 65:2649–2659. https://doi.org/10.1109/TBME.2018.2813015
Wagner M, Mayer BFB, Bodenstedt S, Stemmer K, Fereydooni A, Speidel S, Dillmann R, Nickel F, Fischer L, Kenngott HG (2018) Computer-assisted 3D bowel length measurement for quantitative laparoscopy. Surg Endosc 32:4052–4061. https://doi.org/10.1007/s00464-018-6135-y
Neumuth T, Jannin P, Strauss G, Meixensberger J, Burgert O (2009) Validation of knowledge acquisition for surgical process models. J Am Med Inform Assoc JAMIA 16:72–80. https://doi.org/10.1197/jamia.M2748
Nwoye CI, Yu T, Gonzalez C, Seeliger B, Mascagni P, Mutter D, Marescaux J, Padoy N (2022) Rendezvous: attention mechanisms for the recognition of surgical action triplets in endoscopic videos. Med Image Anal 78:102433. https://doi.org/10.1016/j.media.2022.102433
Diana M, Noll E, Diemunsch P, Dallemagne B, Benahmed MA, Agnus V, Soler L, Barry B, Namer IJ, Demartines N, Charles A-L, Geny B, Marescaux J (2014) Enhanced-reality video fluorescence: a real-time assessment of intestinal viability. Ann Surg 259:700–707. https://doi.org/10.1097/SLA.0b013e31828d4ab3
Lin J, Clancy NT, Qi J, Hu Y, Tatla T, Stoyanov D, Maier-Hein L, Elson DS (2018) Dual-modality endoscopic probe for tissue surface shape reconstruction and hyperspectral imaging enabled by deep neural networks. Med Image Anal 48:162–176. https://doi.org/10.1016/j.media.2018.06.004
Francis NK, Curtis NJ, Conti JA, Foster JD, Bonjer HJ, Hanna GB, EAES committees (2018) EAES classification of intraoperative adverse events in laparoscopic surgery. Surg Endosc 32:3822–3829. https://doi.org/10.1007/s00464-018-6108-1
Dell-Kuster S, Gomes NV, Gawria L, Aghlmandi S, Aduse-Poku M, Bissett I, Blanc C, Brandt C, ten Broek RB, Bruppacher HR, Clancy C, Delrio P, Espin E, Galanos-Demiris K, Gecim IE, Ghaffari S, Gié O, Goebel B, Hahnloser D, Herbst F, Orestis I, Joller S, Kang S, Martín R, Mayr J, Meier S, Murugesan J, Nally D, Ozcelik M, Pace U, Passeri M, Rabanser S, Ranter B, Rega D, Ridgway PF, Rosman C, Schmid R, Schumacher P, Solis-Pena A, Villarino L, Vrochides D, Engel A, O’Grady G, Loveday B, Steiner LA, Van Goor H, Bucher HC, Clavien P-A, Kirchhoff P, Rosenthal R (2020) Prospective validation of classification of intraoperative adverse events (ClassIntra): international, multicentre cohort study. BMJ m2917. https://doi.org/10.1136/bmj.m2917
Yamazaki Y, Kanaji S, Matsuda T, Oshikiri T, Nakamura T, Suzuki S, Hiasa Y, Otake Y, Sato Y, Kakeji Y (2020) Automated surgical instrument detection from laparoscopic gastrectomy video images using an open source convolutional neural network platform. J Am Coll Surg 230:725-732.e1. https://doi.org/10.1016/j.jamcollsurg.2020.01.037
Schwaner KL, Dall’Alba D, Jensen PT, Fiorini P, Savarimuthu TR (2021) Autonomous needle manipulation for robotic surgical suturing based on skills learned from demonstration. In: 2021 IEEE 17th international conference on automation science and engineering (CASE), pp 235–241
Yamazaki Y, Kanaji S, Kudo T, Takiguchi G, Urakawa N, Hasegawa H, Yamamoto M, Matsuda Y, Yamashita K, Matsuda T, Oshikiri T, Nakamura T, Suzuki S, Otake Y, Sato Y, Kakeji Y (2021) Quantitative comparison of surgical device usage in laparoscopic gastrectomy between surgeons’ skill levels: an automated analysis using a neural network. J Gastrointest Surg Off J Soc Surg Aliment Tract. https://doi.org/10.1007/s11605-021-05161-4
Maier-Hein L, Wagner M, Ross T, Reinke A, Bodenstedt S, Full PM, Hempe H, Mindroc-Filimon D, Scholz P, Tran TN, Bruno P, Kisilenko A, Müller B, Davitashvili T, Capek M, Tizabi MD, Eisenmann M, Adler TJ, Gröhl J, Schellenberg M, Seidlitz S, Lai TYE, Pekdemir B, Roethlingshoefer V, Both F, Bittel S, Mengler M, Mündermann L, Apitz M, Kopp-Schneider A, Speidel S, Nickel F, Probst P, Kenngott HG, Müller-Stich BP (2021) Heidelberg colorectal data set for surgical data science in the sensor operating room. Sci Data 8:101. https://doi.org/10.1038/s41597-021-00882-2
Hashimoto DA, Rosman G, Witkowski ER, Stafford C, Navarette-Welton AJ, Rattner DW, Lillemoe KD, Rus DL, Meireles OR (2019) Computer vision analysis of intraoperative video: automated recognition of operative steps in laparoscopic sleeve gastrectomy. Ann Surg 270:414–421. https://doi.org/10.1097/SLA.0000000000003460
Yang C-C, Chen C-Y, Kuo Y-T, Ko C-C, Wu W-J, Liang C-H, Yun C-H, Huang W-M (2022) Radiomics for the prediction of response to antifibrotic treatment in patients with idiopathic pulmonary fibrosis: a pilot study. Diagn Basel Switz 12:1002. https://doi.org/10.3390/diagnostics12041002
Wan S, Wei Y, Zhang X, Yang C, Hu F, Song B (2022) Computed tomography-based texture features for the risk stratification of portal hypertension and prediction of survival in patients with cirrhosis: a preliminary study. Front Med. https://doi.org/10.3389/fmed.2022.863596
Schuh F, Fink MA, Feisst M, Eckert C, Dörr-Harim C, Knebel P, Diener MK, Büchler MW, Mihaljevic AL, Probst P (2022) Protocol of a prospective study investigating the association of PAncreatic parenchymal RISk factors with postoperative pancreatic fistula after partial pancreaticoduodenectomy (PARIS trial). BMJ Open 12:e054138. https://doi.org/10.1136/bmjopen-2021-054138
Kolbinger FR, Lambrecht J, Leger S, Ittermann T, Speidel S, Weitz J, Hoffmann R-T, Distler M, Kühn J-P (2022) The image-based preoperative fistula risk score (preFRS) predicts postoperative pancreatic fistula in patients undergoing pancreatic head resection. Sci Rep 12:4064. https://doi.org/10.1038/s41598-022-07970-2
Brunner M, Kovacevic J, Krautz C, Merkel S, Hartmann A, Grützmann R, Haller F, Weber GF (2022) Analysis of intraoperative frozen pancreatic resection margin and prediction of postoperative pancreatic fistula risk during pancreatoduodenectomy. J Am Coll Surg 234:928–937. https://doi.org/10.1097/XCS.0000000000000142
Heller AR, Mees ST, Lauterwald B, Reeps C, Koch T, Weitz J (2020) Detection of deteriorating patients on surgical wards outside the ICU by an Automated MEWS-based early warning system with paging functionality. Ann Surg 271:100–105. https://doi.org/10.1097/SLA.0000000000002830
Chang L, Hogle NJ, Moore BB, Graham MJ, Sinanan MN, Bailey R, Fowler DL (2007) Reliable assessment of laparoscopic performance in the operating room using videotape analysis. Surg Innov 14:122–126. https://doi.org/10.1177/1553350607301742
Belagiannis V, Wang X, Shitrit HBB, Hashimoto K, Stauder R, Aoki Y, Kranzfelder M, Schneider A, Fua P, Ilic S, Feussner H, Navab N (2016) Parsing human skeletons in an operating room. Mach Vis Appl 27:1035–1046. https://doi.org/10.1007/s00138-016-0792-4
Guzmán-García C, Gómez-Tome M, Sánchez-González P, Oropesa I, Gómez EJ (2021) Speech-based surgical phase recognition for non-intrusive surgical skills’ assessment in educational contexts. Sensors. https://doi.org/10.3390/s21041330
Chen AB, Liang S, Nguyen JH, Liu Y, Hung AJ (2021) Machine learning analyses of automated performance metrics during granular sub-stitch phases predict surgeon experience. Surgery 169:1245–1249. https://doi.org/10.1016/j.surg.2020.09.020
Gillies RJ, Kinahan PE, Hricak H (2016) Radiomics: images are more than pictures, they are data. Radiology 278:563–577. https://doi.org/10.1148/radiol.2015151169
Vedula SS, Ishii M, Hager GD (2017) Objective assessment of surgical technical skill and competency in the operating room. Annu Rev Biomed Eng 19:301–325. https://doi.org/10.1146/annurev-bioeng-071516-044435
Suurmeijer JA, Henry AC, Bonsing BA, Bosscha K, van Dam RM, van Eijck CH, Gerhards MF, van der Harst E, de Hingh IH, Intven MP, Kazemier G, Wilmink JW, Lips DJ, Wit F, de Meijer VE, Molenaar IQ, Patijn GA, van der Schelling GP, Stommel MWJ, Busch OR, Koerkamp BG, van Santvoort HC, Besselink MG, Dutch Pancreatic Cancer Group (2022) Outcome of pancreatic surgery during the first six years of a mandatory audit within the Dutch Pancreatic Cancer Group. Ann Surg. https://doi.org/10.1097/SLA.0000000000005628
Gordon L, Reed C, Sorensen JL, Schulthess P, Strandbygaard J, Mcloone M, Grantcharov T, Shore EM (2022) Perceptions of safety culture and recording in the operating room: understanding barriers to video data capture. Surg Endosc 36:3789–3797. https://doi.org/10.1007/s00464-021-08695-5
Jesudason E (2022) Surgery should be routinely videoed. J Med Ethics. https://doi.org/10.1136/medethics-2022-108171
Hu Y-Y, Peyre SE, Arriaga AF, Osteen RT, Corso KA, Weiser TG, Swanson RS, Ashley SW, Raut CP, Zinner MJ, Gawande AA, Greenberg CC (2012) Postgame analysis: using video-based coaching for continuous professional development. J Am Coll Surg 214:115–124. https://doi.org/10.1016/j.jamcollsurg.2011.10.009
Funding
Open Access funding enabled and organized by Projekt DEAL. This work has been funded by the German Federal Ministry of Health (BMG) within the “Surgomics” project (Grant Number BMG 2520DAT82), by the German Research Foundation DFG within the Cluster of Excellence EXC 2050: “Center for Tactile Internet with Human-in-the-Loop (CeTI)” (Project Number 390696704) and by project funding within the Else Kröner Fresenius Center for Digital Health (EKFZ), Dresden, Germany.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Disclosures
A. Stern, H. Alwanni, L. Mündermann, and J. Fallert are employees of KARL STORZ SE & Co. KG. M. Wagner, S. Bodenstedt, F. Fritz-Kebede, M. Dugas, M. Distler, J. Weitz, B. Müller-Stich and S. Speidel are project leaders of the Surgomics-project, funded by the German Federal Ministry of Health (Grant Number 2520DAT82) with medical device manufacturer KARL STORZ SE & Co. KG being a project partner. J. M. Brandenburg, A. Schulze, A. C. Jenke, M.T.J. Daum, F. R. Kolbinger, N. Bhasker, G. Schneider, G. Krause-Jüttler, O. Burgert, D. Wilhelm, F. Nickel, and L. Maier-Hein have no conflicts of interest or financial ties to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wagner, M., Brandenburg, J.M., Bodenstedt, S. et al. Surgomics: personalized prediction of morbidity, mortality and long-term outcome in surgery using machine learning on multimodal data. Surg Endosc 36, 8568–8591 (2022). https://doi.org/10.1007/s00464-022-09611-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00464-022-09611-1