Abstract
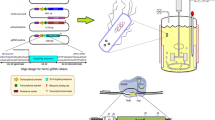
Escherichia coli strains W3110 and BL21 were engineered for the production of plasmid DNA (pDNA) under aerobic and transitions to microaerobic conditions. The gene coding for recombinase A (recA) was deleted in both strains. In addition, the Vitreoscilla hemoglobin (VHb) gene (vgb) was chromosomally inserted and constitutively expressed in each E. coli recA mutant and wild type. The recA inactivation increased the supercoiled pDNA fraction (SCF) in both strains, while VHb expression improved the pDNA production in W3110, but not in BL21. Therefore, a codon-optimized version of vgb was inserted in strain BL21recA−, which, together with W3110recA−vgb+, was tested in cultures with shifts from aerobic to oxygen-limited regimes. VHb expression lowered the accumulation of fermentative by-products in both strains. VHb-expressing cells displayed higher oxidative activity as indicated by the Redox Sensor Green fluorescence, which was more intense in BL21 than in W3110. Furthermore, VHb expression did not change pDNA production in W3110, but decreased it in BL21. These results are useful for understanding the physiological effects of VHb expression in two industrially relevant E. coli strains, and for the selection of a host for pDNA production.





Similar content being viewed by others
References
Heins AL, Weuster-Botz D (2018) Population heterogeneity in microbial bioprocesses: origin, analysis, mechanisms, and future perspectives. Bioprocess Biosyst Eng 41(7):889–916
Jaén KE, Olivares R, Sigala JC, Lara AR (2017) Effects of oxygen availability on plasmid DNA production by Escherichia coli. BMC Biotechnol 17:60
Passarinha LA, Diogo MM, Queiroz JA, Monteiro GA, Fonseca JP, Prazeres DMF (2006) Production of ColE1 type plasmid by Escherichia coli DH5α cultured under nonselective conditions. J Microbiol Biotecnol 16:20–24
Veeravalli K, Schindler T, Dong E, Yamada M, Hamilton R, Laird MW (2018) Strain engineering to reduce acetate accumulation during microaerobic growth conditions in Escherichia coli. Biotechnol Prog 34(2):303–314
Stark BC, Pagilla KR, Dikshit KL (2015) Recent applications of Vitreoscilla hemoglobin technology in bioproduct synthesis and bioremediation. Appl Microbiol Biotechnol 99:1627–1636
Pablos TE, Meza E, Le Borgne S, Gosset G, Ramírez OT, Lara AR (2011) Vitreoscilla hemoglobin expression in engineered Escherichia coli: improved performance in high cell-density batch cultivations. Biotechnol J 6(8):993–1002
Pablos TE, Sigala JC, Le Borgne S, Lara AR (2014) Aerobic expression of Vitreoscilla hemoglobin efficiently reduces overflow metabolism in Escherichia coli. Biotechnol J 9(6):791–799
Pablos TE, Olivares R, Sigala JC, Ramírez OT, Lara AR (2016) Toward efficient microaerobic processes using engineered Escherichia coli W3110 strains. Eng Life Sci 16(7):588–597
Hobernik D, Bros M (2018) DNA Vaccines—how far from clinical use? Int J Mol Sci 19:3605
Carnes AE, Williams JA (2014) Plasmid fermentation process for DNA immunization applications. Methods Molecular Biol 1143:197–217
Rozkov A, Larsson B, Gillström S, Björnestedt R, Schmidt SR (2008) Large-scale production of endotoxin-free plasmids for transient expression in mammalian cell culture. Biotechnol Bioeng 99:557–566
Brand E, Ralla K, Neubauer P (2012) Strategies for Plasmid DNA Production in Escherichia coli. In: Subramanian G (ed) Biopharmaceutical Production Technology, 1st edn. Wiley, Berlin, pp 1–41
Grunzel P, Pilarek M, Steinbrück D, Neubauer A, Brand E, Kumke MU, Neubauer P, Krause M (2014) Mini-scale cultivation method enables expeditious plasmid production in Escherichia coli. Biotechnol J 9(1):128–136
Galindo JM, Barrón BL, Lara AR (2016) Plasmid DNA production in shake flasks is improved by enzyme-controlled glucose release. Ann Microbiol 66(3):1337–1342
Ramírez EA, Velázquez D, Lara AR (2016) Enhancing plasmid DNA production in shake flask by enzyme-mediated glucose release and engineered E. c oli. Biotechnol Lett 38(4):651–657
Pilarek M, Brand E, Hillig F, Krause M, Neubauer P (2013) Enhanced plasmid production in miniaturized high-cell-density cultures of Escherichia coli supported with perfluorinated oxygen carrier. Bioprocess Biosyst Eng 36(8):1079–1086
Soto R, Caspeta L, Barrón BL, Gosset G, Ramírez OT, Lara AR (2011) High cell-density cultivation in batch mode for plasmid DNA vaccine production by a metabolically engineered E. c oli strain with minimized overflow metabolism. Biochem Eng J 56(3):165–171
Borja MG, Meza E, Gosset G, Ramírez OT, Lara AR (2012) Engineering E. c oli to increase plasmid DNA production in high cell-density cultivations in batch mode. Microb Cell Fact 11:132
Mairhofer J, Lara AR (2014) Advances in host and vector development for plasmid DNA vaccines production. In: Lawman MJP, Lawman PD (eds) Cancer vaccines-methods and protocols methods in molecular biology, 1139th edn. Springer, New York, pp 505–542
Xia XX, Qian ZG, Lee SY (2011) Comparative proteomic and genetic analyses reveal unidentified mutations in Escherichia coli XL1-Blue and DH5α. FEMS Microbiol Lett 314(2):119–124
Yau SY, Keshavarz-Moore E, Ward J (2008) Host strain influences on supercoiled plasmid DNA production in Escherichia coli: Implications for efficient design of large scale processes. Biotechnol Bioeng 101:529–544
Gonçalves GA, Prather KL, Monteiro GA, Prazeres DM (2014) Engineering of Escherichia coli strains for plasmid biopharmaceutical production: scale-up challenges. Vaccine 32(24):2847–2850
Monk JM, Koza A, Campodonico MA, Machado D, Seoane JM, Palsson BO, Herrgård MJ, Feist AM (2016) Multi-omics quantification of species variation of Escherichia coli links molecular features with strain phenotypes. Cell Syst 3(3):238–251
Marisch K, Bayer K, Scharl T, Mairhofer J, Krempl PM, Hummel K, Razzazi-Fazeli E, Striedner GA (2013) Comparative analysis of industrial Escherichia coli K–12 and B strains in high-glucose batch cultivations on process-, transcriptome- and proteome level. PLoS ONE 8(8):e70516
Noronha SB, Yeh HJC, Spande TF, Shiloach J (2000) Investigation of the TCA cycle and the glyoxylate shunt in Escherichia coli BL21 and JM109 using 13C-NMR/MS. Biotechnol Bioeng 68(3):316–327
Li Z, Nimtz M, Rinas U (2014) The metabolic potential of Escherichia coli BL21 in defined and rich medium. Microb Cell Fact 13:45
Roca AI, Cox MM (1997) RecA protein: structure, function, and role in recombinational DNA repair. Prog Nucleic Acid Res Mol Biol 56:129–223
Phue JN, Lee SJ, Trinh L, Shiloach J (2008) Modified Escherichia coli B (BL21), a superior producer of plasmid DNA compared with Escherichia coli K (DH5α). Biotechnol Bioeng 101:831–836
Konopka MC, Strovas TJ, Ojala DS, Chistoserdova L, Lidstrom ME, Kalyuzhnaya MG (2011) Respiration response imaging for real-time detection of microbial function at the single-cell level. Appl Environ Microbiol 77:67–72
Baert J, Delepierre A, Telek S, Toye D, Delamotte A, Lara AR, Jaén KE, Gosset G, Jensen P, Delvigne F (2016) Microbial population heterogeneity versus bioreactor heterogeneity: evaluation of Redox Sensor Green as an exogenous metabolic biosensor. Eng Life Sci 16(7):643–651
Fernandes RL, Nierychlo M, Lundin L, Pedersen AE, Puentes Tellez PE et al (2011) Experimental methods and modeling techniques for description of cell population heterogeneity. Biotechnol Adv 29(6):575–599
Datsenko KA, Wanner BL (2000) One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci USA 97:6640–6645
Sabido A, Martínez LM, de Anda R, Martínez A, Bolívar F, Gosset G (2013) A novel plasmid vector designed for chromosomal gene integration and expression: Use for developing a genetically stable Escherichia coli melanin production strain. Plasmid 69:16–23
FDA (2007) Guidance for Industry: considerations for plasmid DNA vaccines for infectious disease indications. Docket Number: 2005D-0047. Issued by: Center for Biologics Evaluation and Research
Wang JC (2002) Cellular ROLES of DNA topoisomerases: a molecular perspective. Nature 3:430–441
Kuzminov A (1999) Recombinational repair of DNA damage in Escherichia coli and bacteriophage lambda. Microbiol Mol Biol Rev 63:751–813
Dorman CJ (2008) Regulation of transcription in bacteria by DNA supercoiling. In: EI-Sharoud E (ed), Bacterial physiology: a molecular approach, Springer, Berlin, p 155–178
Tsai PS, Hatzimanikatis V, Bailey JE (1996) Effect of Vitreoscilla hemoglobin dosage on microaerobic Escherichia coli carbon and energy metabolism. Biotechnol Bioeng 49(2):139–150
Kim TS, Jung HM, Kim SY, Zhang L, Li J, Sigdel S, Park JH, Haw JR, Lee JK (2015) Reduction of acetate and lactate contributed to enhancement of a recombinant protein production in E. c oli BL21. J Microbiol Biotechnol 25(7):1093–1100
Hecker M, Schroeter A, Mach F (1983) Replication of pBR322 DNA in stringent and relaxed strains of Escherichia coli. Mol Gen Genet 190(2):355–357
Hofmann KH, Neubauer P, Riethdorf S, Hecker M (1990) Amplification of pBR322 plasmid DNA in Escherichia coli relA strains during batch and fed-batch fermentation. J Basic Microbiol 30:37–41
Reckinger AR, Jeong KS, Khodursky AB, Hiasa H (2007) RecA can stimulate the relaxation activity of topoisomerase I: molecular basis of topoisomerase-mediated genome-wide transcriptional responses in Escherichia coli. Nucl Acids Res 35(1):79–86
Acknowledgements
Financial support from CONACyT Grant Number 256617 is acknowledged.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Jaén, K.E., Velazquez, D., Delvigne, F. et al. Engineering E. coli for improved microaerobic pDNA production. Bioprocess Biosyst Eng 42, 1457–1466 (2019). https://doi.org/10.1007/s00449-019-02142-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00449-019-02142-5