Abstract
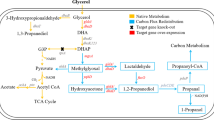
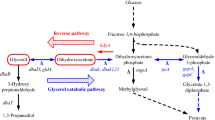
To investigate the relationship between the yield of 1,3-propanediol (1,3-PD) and the flux variation in metabolic pathways of Klebsiella pneumoniae, an optimized calculation method was constructed on basis of dynamic flux balance analysis by combining genome-scale flux balance analysis with a kinetic model of extracellular metabolites. Through optimizing calculations, a more completely expanded metabolic pathway was obtained, which includes the previously reported metabolic pathway and additional three pathways or site: a pentose phosphate pathway (PPP) elicited at the dihydroxyacetone (DHA) node to provide more reducing equivalents; a branch of synthetic amino acids at the 3-phosphoglycerate (3PG) node; and the α-ketoglutarate site in the tricarboxylic acid (TCA) cycle leading to anabolic pathways for glutamate and other amino acids. On this basis, the relationships between the dynamic flux distribution of the important nodes in the metabolic pathway and the yield of 1,3-propanediol were analyzed. First, dynamic flux change from DHA to the PPP is positively correlated with the yield. Second, variation in flux in the TCA cycle is also positively correlated with the yield of 1,3-propanediol. In addition, the influence of the feedback loop formed by the cofactor tetrahydrofolate on the flux change of TCA in the amino acid anabolic pathway was examined. These results are of important reference value and have guiding significance for the extension of the glycerol metabolism pathway in K. pneumoniae, the rational transformation of genetic engineering in bacteria, and the optimization of metabolic pathways for industrial production.








Similar content being viewed by others
Abbreviations
- 1,3-PD:
-
1,3-Propanediol
- 10fThf:
-
10-Formyltetrahydrofolate
- 3HPA:
-
3-Hydroxypropanal
- 3PG:
-
3-Phosphoglycerate
- 3PHP:
-
3-Phosphohydroxypyruvate
- Ac:
-
Acetic acid
- AKG:
-
α-Ketoglutarate
- Cit:
-
Citrate
- DFBA:
-
Dynamic flux balance analysis
- DHA:
-
Dihydroxyacetone
- dha :
-
Dha regulon
- DHAK:
-
Dihydroxyacetone phosphate kinase
- DHAP:
-
Dihydroxyacetone phosphate
- DHAPT:
-
Dihydroxyacetone phosphotransferase
- DOA:
-
Dynamic optimization method
- EtOH:
-
Ethanol
- F6P:
-
6-Phosphofructose
- FBA:
-
Flux balance analysis
- Fum:
-
Fumarate
- G3P:
-
3-Phosphoglyceraldehyde
- GDH:
-
Glycerol dehydrogenase
- GDHt:
-
Glycerol dehydratase
- Glu:
-
Glutamate
- Gly:
-
Glycine
- Glyc:
-
Glycerol
- LP:
-
Linear programming
- Mal:
-
Malate
- mlThf:
-
Methylenetetrahydrofolate
- Oaa:
-
Oxaloacetate
- ODE:
-
Ordinary differential equation
- PDH:
-
Pyruvate dehydrogenase
- PDOR:
-
1,3-Propanediol oxidoreductase
- PEP:
-
Phosphoenolpyruvate
- PFL:
-
Pyruvate formate lyase
- PGCD:
-
Phosphoglycerate dehydrogenase
- PGM:
-
Phosphoglycerate mutase
- PPP:
-
Pentose phosphate pathway
- PYR:
-
Pyruvate
- QP:
-
Quadratic programming
- Ser:
-
Serine
- SOA:
-
Static optimization method
- Succ:
-
Succinate
- SuccCoA:
-
Succinyl-coA
- TCA:
-
Tricarboxylic acid
- Thf:
-
Tetrahydrofolate
References
Gungormusler-Yilmaz M, Cicek N, Levin DB, Azbar N (2016) Cell immobilization for microbial production of 1,3-propanediol. Crit Rev Biotechnol 36:482–494
Lee CS, Aroua MK, Daud WMAW, Cognet P, Pérès-Lucchese Y, Fabre PL, Reynes O, Latapie L (2015) A review: conversion of bioglycerol into 1,3-propanediol via biological and chemical method. Renew Sust Energ Rev 42:963–972
Vivek N, Sindhu R, Madhavan A, Anju AJ, Castro E, Faraco V, Pandey A, Binod P (2017) Recent advances in the production of value added chemicals and lipids utilizing biodiesel industry generated crude glycerol as a substrate–metabolic aspects, challenges and possibilities: an overview. Bioresour Technol 239:507–517
Celinska E (2010) Debottlenecking the 1,3-propanediol pathway by metabolic engineering. Biotechnol Adv 28:519–530
Lee JH, Jung M-Y, Oh M-K (2018) High-yield production of 1,3-propanediol from glycerol by metabolically engineered Klebsiella pneumoniae. Biotechnol Biofuels 11:104
Zeng AP, Ross A, Biebl H, Tag C, Günzel B, Deckwer WD (1994) Multiple product inhibition and growth modeling of clostridium butyricum and Klebsiella pneumoniae in glycerol fermentation. Biotechnol Bioeng 44:902–911
Zeng AP (1995) A kinetic model for product formation of microbial and mammalian cells. Biotechnol Bioeng 46:314–324
Zeng AP, Deckwer WD (1995) A kinetic model for substrate and energy consumption of microbial growth under substrate-sufficient conditions. Biotechnol Prog 11:71–79
Xiu ZL, Zeng AP, An LJ (2000) Mathematical modeling of kinetics and research on multiplicity of glycerol bioconversion to 1,3-propanediol. J Dalian Univ Technol 40:428–433
Sun YQ, Qi WT, Teng H, Xiu ZL, Zeng AP (2008) Mathematical modeling of glycerol fermentation by Klebsiella pneumoniae: concerning enzyme-catalytic reductive pathway and transport of glycerol and 1,3-propanediol across cell membrane. Biochem Eng J 38:22–32
Yuan JL, Zhu X, Zhang X, Yin HC, Feng EM, Xiu ZL (2014) Robust identification of enzymatic nonlinear dynamical systems for 1, 3-propanediol transport mechanisms in microbial batch culture. Appl Math Comput 232:150–163
Guo YJ, Feng EM, Wang L, Xiu ZL (2014) Complex metabolic network of 1,3-propanediol transport mechanisms and its system identification via biological robustness. Bioprocess Biosyst Eng 37:677–686
Zhang QR, Teng H, Sun YQ, Xiu ZL (2007) Metabolic flux analysis of bioconversion of glycerol into 1,3-propanediol by Klebsiella pneumoniae. In: 2007 1st international conference on bioinformatics and biomedical engineering, July ICBBE 6, 2007 July 8, 2007, IEEE, Wuhan, pp 1269–1272
Zhang QR, Teng H, Sun YQ, Xiu ZL, Zeng AP (2008) Metabolic flux and robustness analysis of glycerol metabolism in Klebsiella pneumoniae. Bioprocess Biosyst Eng 31:127–135
Zhang QR, Xiu ZL (2009) Metabolic pathway analysis of glycerol metabolism in Klebsiella pneumoniae incorporating oxygen regulatory system. Biotechnol Prog 25:103–115
Xu GX, Li CX (2017) Identifying the shared metabolic objectives of glycerol bioconversion in Klebsiella pneumoniae under different culture conditions. J Biotechnol 248:59–68
Feist AM, Herrgard MJ, Thiele I, Reed JL, Palsson BO (2009) Reconstruction of biochemical networks in microorganisms. Nat Rev Microbiol 7:129–143
Yilmaz LS, Walhout AJM (2017) Metabolic network modeling with model organisms. Curr Opin Chem Biol 36:32–39
Dai Z, Locasale JW (2017) Understanding metabolism with flux analysis: From theory to application. Metab Eng 43:94–102
Chen P-W, Theisen MK, Liao JC (2017) Metabolic systems modeling for cell factories improvement. Curr Opin Biotechnol 46:114–119
Palsson B (2015) Systems biology: constraint-based reconstruction and analysis. Cambridge University Press, Cambridge
Mahadevan R, Edwards JS, Doyle FJ (2002) Dynamic flux balance analysis of diauxic growth in Escherichia coli. Biophys J 83:1331–1340
Feng X, Xu Y, Chen Y, Tang YJ (2012) Integrating flux balance analysis into kinetic models to decipher the dynamic metabolism of Shewanella oneidensis MR-1. PLoS Comp Biol. https://doi.org/10.1371/journal.pcbi.1002376
Sánchez BJ, Pérez-Correa JR, Agosin E (2014) Construction of robust dynamic genome-scale metabolic model structures of Saccharomyces cerevisiae through iterative re-parameterization. Metab Eng 25:159–173
Nikdel A, Braatz RD, Budman HM (2018) A systematic approach for finding the objective function and active constraints for dynamic flux balance analysis. Bioprocess Biosyst Eng 41:641–655
Orth JD, Thiele I, Palsson BO (2010) What is flux balance analysis? Nat Biotechnol 28:245–248
Feist AM, Palsson BO (2010) The biomass objective function. Curr Opin Microbiol 13:344–349
Liao YC, Huang TW, Chen FC, Charusanti P, Hong JSJ, Chang HY, Tsai SF, Palsson BO, Hsiung CA (2011) An experimentally validated genome-scale metabolic reconstruction of Klebsiella pneumoniae MGH 78578, iYL1228. J Bacteriol 193:1710–1717
Saitua F, Torres P, Pérez-Correa JR, Agosin E (2017) Dynamic genome-scale metabolic modeling of the yeast Pichia pastoris. BMC Syst Biol 11:27
Sun J, van den Heuvel J, Soucaille P, Qu Y, Zeng AP (2003) Comparative genomic analysis of dha regulon and related genes for anaerobic glycerol metabolism in bacteria. Biotechnol Prog 19:263–272
Schirch V, Szebenyi DM (2005) Serine hydroxymethyltransferase revisited. Curr Opin Chem Biol 9:482–487
Lu XY, Ren SL, Lu JZ, Zong H, Song J, Zhuge B (2018) Enhanced 1,3-propanediol production in Klebsiella pneumoniae by a combined strategy of strengthening the TCA cycle and weakening the glucose effect. J Appl Microbiol 124:682–690
Amador-Noguez D, Feng X-J, Fan J, Roquet N, Rabitz H, Rabinowitz JD (2010) Systems-level metabolic flux profiling elucidates a complete, bifurcated tricarboxylic acid cycle in Clostridium acetobutylicum. J Bacteriol 192:4452–4461
Zhou Y, Pijuan M, Zeng RJ, Yuan Z (2009) Involvement of the tca cycle in the anaerobic metabolism of polyphosphate accumulating organisms (PAOs). Water Res 43:1330–1340
Kajiura H, Mori K, Tobimatsu T, Toraya T (2001) Characterization and mechanism of action of a reactivating factor for adenosylcobalamin-dependent glycerol dehydratase. J Biol Chem 276:36514–36519
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Grant no. 21476042).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Human and animal rights statement
This article does not contain any studies with human participants or animals performed by any of the authors.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Pan, DT., Wang, XD., Shi, HY. et al. Dynamic flux balance analysis for microbial conversion of glycerol into 1,3-propanediol by Klebsiella pneumoniae. Bioprocess Biosyst Eng 41, 1793–1805 (2018). https://doi.org/10.1007/s00449-018-2002-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00449-018-2002-4