Abstract
Previous studies have observed relationships between pancreatitis and gut microbiota; however, specific changes in gut microbiota abundance and underlying mechanisms in pancreatitis remain unknown. Metabolites are important for gut microbiota to fulfil their biological functions, and changes in the metabolic and immune environments are closely linked to changes in microbiota abundance. We aimed to clarify the mechanisms of gut–pancreas interactions and explore the possible role of metabolites and the immune system. To this end, we conducted two-sample Mendelian randomisation (MR) analysis to evaluate the casual links between four different types of pancreatitis and gut microbiota, metabolites, and inflammatory cytokines. A two-step MR analysis was conducted to further evaluate the probable mediating pathways involving metabolites and inflammatory cytokines in the causal relationship between pancreatitis and gut microbiota. In total, six potential mediators were identified in the causal relationship between pancreatitis and gut microbiota. Nineteen species of gut microbiota and seven inflammatory cytokines were genetically associated with the four types of pancreatitis. Metabolites involved in glucose and amino acid metabolisms were genetically associated with chronic pancreatitis, and those involved in lipid metabolism were genetically associated with acute pancreatitis. Our study identified alterations in the gut microbiota, metabolites, and inflammatory cytokines in pancreatitis at the genetic level and found six potential mediators of the pancreas–gut axis, which may provide insights into the precise diagnosis of pancreatitis and treatment interventions for gut microbiota to prevent the exacerbation of pancreatitis. Future studies could elucidate the mechanism underlying the association between pancreatitis and the gut microbiota.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pancreatitis is the most prevalent exocrine pancreatic disease that has a major impact on the homeostasis of the digestive system. The destruction of pancreatic structures during the development of pancreatitis causes abnormal pancreatic secretion, which can disrupt intestinal homeostasis and disturb gut microbiota (Akshintala et al. 2019; Capurso et al. 2016). Several clinical studies have reported overall changes in the gut microbiota in patients with pancreatic disease (Tan et al. 2015) and attempted to clarify the specific role of the gut microbiota in pancreatic disease development (Mirji et al. 2022). However, current research on the interaction between gut microbiota and the pancreas is relatively limited. Disturbance of gut microbiota due to pancreatitis and the underlying mechanisms remain to be investigated.
Researchers have preliminarily identified three ways in which gut flora intervenes in the course of pancreatitis. Gut microbiota can migrate directly to the pancreas via the duodenal-pancreatic duct and invade the pancreatic tissue via the mesenteric vein and peri-intestinal lymph nodes (Thomas and Jobin 2020; Diehl et al. 2013), exacerbating chronic pancreatitis (CP). Moreover, disturbed gut microbiota diminishes the production of short-chain fatty acids, leading to inadequate inhibition of the pro-fibrotic function of histone deacetylase and exacerbating CP (Pang and Zhuang 2010; Bombardo et al. 2018). Additionally, short-chain fatty acids and other microbiota productions could influence the inflammatory process by regulating the production of inflammatory cytokines and the differentiation of immune cells (Li et al. 2018; Sun et al. 2018).
Moreover, gut microbiota has been associated with acute pancreatitis (AP). Disturbed gut microbiota causes impaired intestinal barrier and ectopic microbiota, which may be one of the crucial mechanisms by which gut microbiota promote AP (Sonika et al. 2017). Notably, gut microbiota metabolites indirectly influence the course of AP by interacting with immune molecules, such as bifidobacteria, through its metabolite lactate and interfering with the TLR4/MyD88 and NLRP3/Caspase1 pathways to alleviate AP symptoms (Li et al. 2022a). This indicates that metabolites and the immune system may play a pivotal role in the intricate mechanisms through which the gut microbiota intervene in pancreatitis.
The ‘pancreas-gut axis’ concept was first proposed in 2016 (Perry et al. 2016). The authors demonstrated that acetate produced by gut microbiota metabolism could modulate pancreatic β-cell function and increase insulin secretion. A vicious cycle of ‘pancreatitis-gut microbiota disruption-pancreatitis exacerbation’ may exist in patients with pancreatitis. Capurso et al. (2016) reported that small intestinal bacterial overgrowth (SIBO) was present in 36% of CP patients and treating SIBO in pancreatitis patients could improve their exocrine insufficiency symptoms (Bashir et al. 2018). Exploring how pancreatitis alters the abundance of gut microbiota and how these changes, along with the disruption of metabolic homeostasis, influence the progression of pancreatitis is critical. However, most previous studies focussed exclusively on the impact of gut microbiota and associated metabolites on pancreatic diseases. Furthermore, although some observational studies have explored how gut microbiota abundance is altered in pancreatitis patients, these clinical studies often had small sample sizes, high heterogeneity, and inconsistent results across different regions.
Mendelian randomisation (MR), using genetic variants indexing of exposure to establish inter-causality relationships with outcomes (Davey Smith and Hemani 2014; Liu et al. 2022), can overcome the confounding biases inherent in observational studies. Hence, this study aimed to use a two-sample MR analysis based on the publicly available genome-wide association studies (GWAS) database from a large population to illustrate the effect of pancreatitis on the gut microbiota and perform two-step MR analysis to explore the role that metabolites and inflammatory cytokines play in this process. Understanding the specific alteration of gut microbiota abundance due to pancreatic inflammation and the associated mechanisms holds the potential for a breakthrough in alleviating the symptoms of pancreatitis and preventing its exacerbation.
Materials and methods
Study design
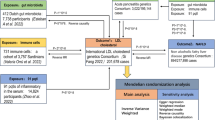
The MR study comprised two phases. In phase 1, we examined the causal effects of all four types of pancreatitis provided by FinnGen GWAS Database (CP, alcohol-induced chronic pancreatitis [AICP], AP, and alcohol-induced acute pancreatitis [AIAP]) on the gut microbiota abundance and metabolites and inflammatory cytokines levels. In phase 2, we assessed the mediating role of metabolites and inflammatory cytokines in the causal relationships between pancreatitis and gut microbiota; in this phase, positive metabolites and inflammatory cytokines in phase 1 were considered as exposures and positive microbiota as outcomes. Generally, we first examined the causal effects of pancreatitis on potential mediators and then assessed the causal effects of mediators on gut microbiota (Fig. 1).
The following criteria should be met in the two-step MR analysis: (1) the exposure should be causally associated with the outcome; (2) the mediator should be causally associated with the outcome independent of the exposure; (3) congruent directions for direct and indirect effects of the exposure on the outcome; and (4) the exposure should be causally associated with the mediator, but not vice versa. In each phase, a simple reverse MR analysis was performed to confirm the absence of reverse causality in the final positive results.
Data sources for pancreatitis, gut microbiota, inflammatory cytokines, and metabolites
Genetic variables of the gut microbiota were obtained from a GWAS dataset of the International Consortium MiBioGen (Kurilshikov et al. 2021); a large international research programme aimed at studying the impact of human genes on gut microbiota at a genome-wide level. This study included genomic and gut microbiota data from 24 cohorts of 18,340 people from multiple countries and ethnicities in Europe, the USA, the Middle East, and East Asia and is the largest GWAS on gut microbiota to date. Overall, 211 taxa (131 genera, 35 families, 20 orders, 16 classes, and 9 phyla) were included.
GWAS summary statistics for pancreatitis were obtained from the FinnGen Consortium R9 release data (Kurki et al. 2023). The current study incorporated the following phenotypes: AIAP (containing 931 cases and 376,346 controls), AICP (containing 1794 cases and 375,483 controls), AP (containing 6223 cases and 330,903 controls), and CP (containing 3320 cases and 330,903 controls). Patients with these four types of pancreatitis were included according to the diagnostic criteria of the International Classification of Diseases Tenth Edition, and the codes were K85.000 (AP), K85.200 (AIAP), K86.100 (CP), and K86.000 (AICP).
Data on genetic variables for 41 inflammatory cytokines were obtained from a study providing genome variant associations in 8293 Finnish individuals. This study combined the results of the Cardiovascular Risk in Young Finns Study and FINRISK surveys (Ahola-Olli et al. 2017). The metabolite GWAS included 24,925 European participants from 14 cohorts, whose blood metabolites were profiled using a quantitative nuclear magnetic resonance metabolomics platform. Notably, there was no overlap in the population or cohort selection between the exposure and outcome groups (Table 1).
Selection of instrumental variables
The following criteria were used to select the optimal instrumental variables (IVs) to improve the authenticity and accuracy of the study conclusions. (1) When we selected pancreatitis as the exposure, we used a genome-wide statistical significance threshold (P < 5 × 10−8) to select IVs. When we chose inflammatory cytokines as the exposure, we adjusted the threshold downwards to a locus-wide significance level (P < 5 × 10−6) if a few relevant single-nucleotide polymorphisms (SNPs) were identified. When the gut microbiome data were defined as the exposure, the threshold of P values was relaxed to 1 × 10–5 to ensure that a suitable number of SNPs were included in the analysis. (2) Because the presence of strong linkage disequilibrium (LD) might result in bias, we ensured that there was no LD among the selected IVs. Data from the 1000 Genomes Project European samples were used as the reference panel to calculate the LD between the SNPs, and the SNPs reaching the threshold (r2 < 0.01, kb = 10,000) were retained for subsequent analysis. (3) The F-statistics for the selected IVs reached a threshold of > 10, ensuring that the causal estimations had no weak instrument bias. Sufficient SNPs were used in our analysis, and their numbers are shown in the figures.
MR analysis
This study used a two-step MR method. The phylum, family, and genus levels were included in the analysis. Metabolites and inflammatory cytokines were considered mediators of the causal effects of pancreatitis on gut microbiota. First, we performed an MR pleiotropy residual sum and outlier (MR-PRESSO) test to remove all outlier SNPs (Verbanck et al. 2018). Second, pleiotropy and heterogeneity tests were conducted to ensure the robustness of the results. Cochran’s Q- and Q-derived P values were calculated to assess heterogeneity (Greco et al. 2015), and MR-Egger intercept P values were calculated to assess pleiotropy (Bowden et al. 2015). The MR-Egger results were accepted when the genetic variants had pleiotropic effects. Finally, five popular MR methods were used to analyse features containing multiple IVs: the inverse-variance weighted (IVW) test, weighted mode, MR-Egger regression, weighted median estimator, and simple mode. In our analyses, there were no cases in which the number of SNPs was less than three; hence, all the above methods were available (Fig. 2). In some cases, IVW was considered a method with relatively high statistical validity; therefore, our analysis was based primarily on IVW results (Greco et al. 2015), with the other four methods acting as complementarity methods. We used the coefficient method as our main method to estimate the mediation effect (VanderWeele 2016) and the delta method to calculate the value of mediation proportion (Burgess et al. 2015).
Ethics
This study used publicly available data from specific human experimentation studies approved by the Ethical Standards Committee. Accordingly, ethical approval was not required for this study.
Statistical analysis
All analyses were performed using the packages TwoSampleMR (version 0.5.6) and MR-PRESSO (version 1.0) in R (version 4.2.2) Studio. Indirect effect of the intermediates was estimated using the website of the RMediation (https://amplab.shinyapps.io/MEDCI/), and the package RMediation (version 1.2.2) could also function similarly. The false discovery rate q value was calculated using the Benjamini–Hochberg method to test multiple hypotheses.
Results
Causal effects of pancreatitis on gut microbiota
Causal effects of CP on gut microbiota
At the phylum level, CP occurrence was genetically associated with a low abundance of Lentisphaerae (β = − 1.013, standard error (SE) = 0.051, P value = 0.046). At the genus level, CP occurrence was genetically associated with a low abundance of Ruminococcus torques (β = − 0.071, SE = 0.023, P value = 0.0016). Contrastingly, CP occurrence was genetically associated with a high abundance of Eubacterium brachy (β = 0.152, SE = 0.069, P value = 0.028) and Lachnospiraceae UCG001 (β = 0.065, SE = 0.030, P value = 0.030).
Causal effects of AICP on gut microbiota
At the genus level, AICP occurrence was genetically associated with a low abundance of Candidatus soleaferrea (β = − 0.079, SE = 0.029, P value = 0.007) and Eubacterium fissicatena (β = − 0.152, SE = 0.059, P value = 0.009). Conversely, AICP occurrence was genetically associated with a high abundance of Eubacterium brachy (β = 0.152, SE = 0.069, P value = 0.029) and Lachnospiraceae UCG001 (β = 0.065, SE = 0.030, P value = 0.030). Furthermore, AICP occurrence was genetically associated with a high abundance of Eubacterium brachy (β = 0.121, SE = 0.047, P value = 0.011), Lachnospiraceae UCG010 (β = 0.048, SE = 0.020, P value = 0.016), Erysipelotrichaceae UCG003 (β = 0.047, SE = 0.021, P value = 0.024), Lachnospiraceae UCG004 (β = 0.041, SE = 0.019, P value = 0.029), Eggerthella (β = 0.067, SE = 0.032, P value = 0.035), Eggerthella (β = 0.067, SE = 0.032, P value = 0.035), and Anaerostipes (β = 0.034, SE = 0.018, P value = 0.049).
Causal effects of AP on gut microbiota
At the family level, AP occurrence was genetically associated with a high abundance of Bacteroidaceae (β = 0.060, SE = 0.030, P value = 0.049). At the phylum level, AP occurrence was genetically associated with a high abundance of Proteobacteria (β = 0.073, SE = 0.027, P value = 0.007). At the genus level, AP occurrence was genetically associated with a low abundance of Candidatus soleaferrea (β = − 0.100, SE = 00.047, P value = 0.033), and a high abundance of Odoribacter (β = 0.069, SE = 0.031, P value = 0.024), Butyricicoccus (β = 0.060, SE = 0.028, P value = 0.034), and Bacteroides (β = 0.060, SE = 0.030, P value = 0.049).
Causal effects of AIAP on gut microbiota
At the family level, AIAP occurrence was genetically associated with a high abundance of Allisonella (β = 0.124, SE = 0.051, P value = 0.0142). The Cochran’s Q-deprived and MR-Egger intercept P values showed no pleiotropy or heterogeneity in the causal effects of pancreatitis on gut microbiota. Simple reverse MR was performed, showing no reverse causality of the gut microbiota on pancreatitis. Some SNPs may overlap owing to the genus, and Butyricicoccus was a child taxon of the family Bacteroidaceae, which was revealed by their same β, SE, and P values (Fig. 3).
Causal effects of pancreatitis on metabolites
Causal effects of CP on metabolites
CP occurrence was genetically associated with a low serum level of citrate (β = − 0.032, SE = 0.014, P value = 0.025).
Causal effects of AICP on metabolites
AICP occurrence was genetically associated with a high serum level of pyruvate (β = 0.025, SE = 0.012, P value = 0.028). Contrastingly, AICP occurrence was genetically associated with a low serum level of valine (β = − 0.025, SE = 0.011, P value = 0.020), tyrosine (β = − 0.024, SE = 0.011, P value = 0.031), and leucine (β = − 0.023, SE = 0.011, P value = 0.031).
Causal effects of AP on metabolites
AP occurrence was genetically associated with a low serum level of apolipoprotein A-I (β = − 0.039, SE = 0.019, P value = 0.044); omega-3 fatty acids (β = − 0.062, SE = 0.025, P value = 0.015); omega-6 fatty acids (β = − 0.049, SE = 0.024, P value = 0.036); omega-7, omega-9 and saturated fatty acids (β = − 0.055, SE = 0.023, P value = 0.016); free cholesterol (β = − 0.078, SE = 0.027, P value = 0.036); free cholesterol (β = − 0.078, SE = 0.027, P value = 0.036); free cholesterol in intermediate-density lipoprotein (IDL) (β = − 0.053, SE = 0.027, P value = 0.046); cholesterol esters in large high-density lipoprotein (HDL) (β = − 0.039, SE = 0.019, P value = 0.040); mono-unsaturated fatty acids (β = − 0.055, SE = 0.023, P value = 0.015); total cholesterol (β = − 0.054, SE = 0.024, P value = 0.025); total fatty acids (β = − 0.060, SE = 0.024, P value = 0.011); total cholesterol in very large HDL (β = − 0.045, SE = 0.018, P value = 0.015); cholesterol esters in very large HDL (β = − 0.056, SE = 0.019, P value = 0.003); free cholesterol in very large HDL (β = − 0.051, SE = 0.018, P value = 0.006); total lipids in very large HDL (β = − 0.053, SE = 0.019, P value = 0.005); phospholipids in very large HDL (β = − 0.049, SE = 0.019, P value = 0.009); and triglycerides in very large HDL (β = − 0.049, SE = 0.019, P value = 0.009).
Causal effects of AIAP on metabolites
There was no causal effect of AIAP occurrence on metabolites. The MR-PRESSO test excluded one SNP in the analysis of the causal effects of AP on free cholesterol, free cholesterol in IDL, and total cholesterol, and the resulting P value remained < 0.05. There was heterogeneity in the causal effects of AP on free cholesterol in the IDL and total cholesterol, whereas the β values of the five MR methods were all negative; therefore, these two metabolites remained in the final results (Fig. 4).
Causal effects of pancreatitis on inflammatory cytokines
Causal effects of CP on inflammatory cytokines
CP occurrence was genetically associated with a high serum level of IL-4 (β = 0.048, SE = 0.024, P value = 0.048), SCGF-β (β = 0.077, SE = 0.036, P value = 0.034), IL-12-P70 (β = 0.048, SE = 0.024, P value = 0.043), and IL-10 (β = 0.049, SE = 0.025, P value = 0.045).
Causal effects of AICP on inflammatory cytokines
There was no causal effect of AICP occurrence on inflammatory cytokines.
Causal effects of AP on inflammatory cytokines
AP occurrence was genetically associated with high serum levels of IL-4 (β = 0.083, SE = 0.036, P value = 0.022) and IL-6 (β = 0.071, SE = 0.035, P value = 0.042).
Causal effects of AIAP on inflammatory cytokines
AIAP occurrence was genetically associated with a low serum level of IL-1β (β = − 0.056, SE = 0.028, P value = 0.047) (Fig. 5).
Potential intermediates in causal effects of pancreatitis on gut microbiota
After a series of processes for exploring the exposure–pancreatitis–outcome gut microbiota causal pathways, we applied a two-step MR to assess the role of inflammatory cytokines and metabolites in mediating the causal effects of pancreatitis on the gut microbiota (Tables 2, 3).
Inflammatory cytokines as intermediates in causal effects of pancreatitis on gut microbiota
After the MR-egger adjustment, the analysis showed that IL-4 (β = 0.032, SE = 0.022) accounted for 49.2% of the total effect of CP on Lachnospiraceae; SCGF-β (β = − 0.026, SE = 0.013) for 25.2% of the total effect of CP on Lentisphaerae, and IL-12-P70 (β = 0.001, SE = 0.001) for 0.6% of the total effect of CP on Eubacterium brachy. Moreover, IL-6 (β = 0.014, SE = 0.009) accounted for 19.2% of the total effect of AP on Proteobacteria (Fig. 6).
Metabolites as intermediates in causal effects of pancreatitis on gut microbiota
The analysis showed that pyruvate (β = 0.007, SE = 0.013) accounted for 10.4% of the total effect of AICP on Eggerthella and free cholesterol in very large HDL (β = − 0.005, SE = 0.003) for 0.5% of the total effect of CP on Candidatus soleaferrea (Table 3, Fig. 7).
Discussion
This study focussed on the causal effects of pancreatitis on the gut microbiota, analysing the independent causal effects of pancreatitis on gut microbiota abundance, serum metabolite levels, and inflammatory cytokine levels. Furthermore, this study addressed whether metabolites and inflammatory cytokines play mediating roles in promoting gut microbiota disruption.
Regarding metabolites, our study demonstrated that CP was genetically associated with a low serum level of citrate and AICP was genetically associated with a high serum level of pyruvate and low serum levels of valine, tyrosine, and leucine. Pyruvate is the key molecule that links glycolysis to the tricarboxylic acid (TCA) cycle (Fig. 8). In addition to its participation in the TCA cycle, pyruvate is a vital component of the phenylalanine metabolic pathway. Valine and leucine can be broken down into pyruvate for subsequent reactions, and pyruvate can be used as a reactant in amino acid conversion reactions. Another MR analysis has reported a strong association between plasma alanine and glutamic acid and the abundance of gut Proteobacteria, which complements our study and further validates the complex relationship between pancreatitis, amino acid metabolism, and gut microbiota disorders (Liu et al. 2022). Additionally, clinical studies with small sample sizes have confirmed reduced serum citrate and tyrosine concentrations in CP patients (Xu et al. 2023), and some basic studies have also shown that activated pancreatic stellate cells promote glycolysis and upregulate pyruvate production in the pancreatic tissues of CP patients (Tao et al. 2021). However, how glycolysis, the TCA cycle, and amino acid metabolism in pancreatic acinar cells influence each other during the development of pancreatitis remains unclear.
The lipids genetically associated with AP can be grouped into three main categories: unsaturated fatty acids, cholesterol, and HDL. Unsaturated fatty acids may be beneficial in the development and prognosis of AP. Alhan et al. (2006) found that feeding omega-3 fatty acids reduced mortality in rats that developed acute necrotising pancreatitis and lowered IL-6 levels, corroborating with our analysis results on the potential causal relationship between AP and inflammatory cytokines. IL-6 may reduce the expression of Apo A1, which may lead to a decrease in circulating HDL-C levels (Navarro et al. 2005). Hypertriglyceridaemia is a complication of AP; however, our study showed that low levels of triglycerides in very large HDL were genetically associated with AP, which could be explained by the low levels of very large HDL, although the specific mechanisms involved remain unclear.
In addition to the metabolites, elevated levels of IL-4, IL-12-P70, IL-10, and SCGF-β were genetically associated with CP (Fig. 9). IL-4 promotes alternative activation of macrophages into M2 cells, and activated M2 cells are associated with fibrosis (Xue et al. 2015). High levels of IL-10 have been detected in CP patients (Tanţău et al. 2021), indicating that when CP occurs, IL-10 secretion may be upregulated to reduce tissue damage. IL-12 stimulates the production of IFN-γ and TNF-α and reduces IL-4-mediated suppression of IFN-γ. IL-6 is released by various immune cells in response to tissue damage and can be a marker for predicting the severity and prognosis of AP (Li et al. 2022b). A low level of IL-1β was genetically associated with AIAP, which may be because TLR2 deficiency in a mouse model of cerulein-induced AP decreased the expression of IL-1β (Li et al. 2023).
The four types of pancreatitis are associated with different gut microbiota disorders (Fig. 10). Our study showed that CP was genetically associated with a decreased abundance of the phylum Lentisphaerae and genus Ruminococcus torques group. A decreased abundance of Lentisphaerae has been observed in patients with autoimmune hepatitis (Lou et al. 2020). Lentisphaerae may be associated with digestive system diseases; however, the exact mechanism underlying its interaction with CP requires further investigation. Ruminococcus torques belongs to the phylum Mycobacterium and has been reported to be associated with Crohn’s and non-alcoholic fatty liver disease (Zhang et al. 2023); however, it has no clear relationship with CP. Notably, CP was genetically associated with high abundance of Eubacterium brachy and Lachnospiraceae UCG001. Eubacterium produces short-chain fatty acids including butyric acid and has been reported to have significant differences between CP patients with and without pancreatic exocrine insufficiency (Jandhyala et al. 2017). Lachnospiraceae is another gut microbiota that produces butyric acid and is reported to dominate the gut microbiota in CP patients with mild exocrine pancreatic insufficiency (Maev et al. 2023).
AICP was genetically associated with a decreased abundance of the genus Candidatus soleaferrea, Eubacterium fissicatena, and Ruminococcaceae UCG009. Candidatus soleaferrea was reported in a study on mental illnesses, including delirium, schizophrenia, and autism (Yu et al. 2023; Kowalski et al. 2023); therefore, its relationship with AICP might be associated with the effects of alcohol on the nervous system. The increased abundances of Erysipelotrichaceae and Eggerthella were genetically associated with AICP, which has also been observed in sleep-deprived mice (Cammann et al. 2023). Psychotropic drugs have been shown to reduce the abundance of Erysipelotrichaceae (Ait Chait et al. 2023). Therefore, the additional effect of AICP on gut microbiota disorders may be twofold: alteration of the raw material of gut microbiota metabolism by long-term ethanol intake and further effect of alcohol on the nervous system through the brain–gut axis.
This study demonstrated that the abundances of Bacteroides, Proteobacteria, Odoribacter, and Butyricicoccus were genetically increased in AP. The small intestine and colon of AP rats had a ‘core microbiota’ composed of bacteria belonging to Bacteroidetes and Proteobacteria (Tao et al. 2019). Odoribacter is a gut microbiota associated with glucose-lipid metabolism. Huber-Ruano et al. (2022) reported the potential positive effects of Odoribacter on insulin sensitivity in individuals with glucose tolerance and obesity. However, there are no current studies on the relationship between AP and Butyricicoccus, although researchers have considered Butyricicoccus as a potentially exploitable probiotic given its ability to produce butyric acid and its tolerance to the gastrointestinal environment (Geirnaert et al. 2014). Disturbances in the gut microbiota of AP patients may be caused by several factors. Oxidative stress, which may be caused by acute inflammation, may lead to changes in the environment in which the gut microbiota grows and nutrients are available. Damage to the intestinal barrier may allow some of the gut microbiota to expand their growth and colonisation, increasing their detection rate. Additionally, a disease-induced decrease in the abundance of some microbiota may indirectly lead to an increase in the abundance of other microbiota.
The impact of metabolites and inflammatory cytokines on the gut microbiota is substantial. Our study demonstrated that pyruvate may act as a mediator between the causal effects of AICP and Eggerthella. Pyruvate is one of the downstream products of alcohol metabolism, and another study on the therapeutic effects of quercetin showed that its use in an antibiotic-treated mouse model unregulated both pyruvate metabolism and the abundance of Eggerthella (Mi et al. 2022). Free cholesterol in very large HDL may play a role as a mediator between the causal effects of AP on Candidatus soleaferrea; however, no current study has reported the underlying mechanism. SCGF-β may play a role as a mediator between the causal effects of CP on Lentisphaerae; IL-12 may play a role as a mediator between the causal effects of CP on Eubacterium, and IL-4 may play a role as a mediator between the causal effects of CP on Lachnospiraceae UCG001. Further, IL-6 may mediate the causal effects of AP on Proteobacteria. Butyric acid inhibits IL-12 production, which may explain the mediating effect observed in this study (Singh et al. 2022). The specific role of inflammatory cytokines in the association between diseases and gut microbiota needs to be explored in future research.
Notably, certain estimations varied from logical expectations. We observed that pancreatitis may lead to an increase in the abundance of certain probiotics. There are two possible reasons for this observation. First, the disease prompts this fraction of probiotics to proliferate and fulfil their protective roles. Second, the disease caused a decrease in the abundance of some gut bacteria, indirectly increasing the abundance of this probiotic fraction. Additionally, although there are few reports showing that Candidatus soleaferrea is able to metabolise lipids directly, free cholesterol in very large HDL appeared to play a mediating role in our analysis; we speculate that there may be other complex indirect roles.
This study had some limitations. First, most of the enrolled patients were European; therefore, the causal relationship between pancreatitis and gut microbiota in other populations remains unknown. Second, our findings only reported the alteration of gut microbiota in pancreatitis patients and identified several metabolites and inflammatory factors with possible mediating effects; however, the underlying mechanisms warrant further investigation. Finally, GWAS data on metabolites and inflammatory cytokines do not fully represent organ-local conditions; therefore, some changes and pathways that are uniquely significant in the pancreas may be overlooked.
To the best of our knowledge, this is the first study to implement MR analysis to address the causal relationship between gut microbiota, metabolites, inflammatory cytokines, and pancreatitis. This analysis compensates for the disadvantage of using the small sample size in previous clinical studies and adds to the doctrine of pancreas–gut axis at the level of gut microbiota. This research aimed to elucidate the alterations in gut microbiota abundance across four types of pancreatitis patients and investigate the influence of metabolites and inflammatory cytokines in this process. Our findings enhance the understanding of the ‘pancreas-gut’ axis mechanism and offer a foundation for further examination of the role of gut microbiota. Additionally, this work suggests novel avenues for developing targeted therapies for pancreatitis by examining the impact of metabolites and inflammatory cytokines. Given that gut microbiota abundance varies among different types of pancreatitis patients, assessing microbiota abundance could not only shed light on the pathogenesis of pancreatitis but also potentially form a basis for customized microbiota supplementation strategies in treatment.
In conclusion, 19 species of gut microbiota and 7 inflammatory cytokines were genetically associated with the 4 types of pancreatitis. Metabolites involved in glucose and amino acid metabolisms were genetically associated with CP, and those involved in lipid metabolism were genetically associated with AP. Six potential mediators were identified in the causal effects of pancreatitis on the gut microbiota. Accordingly, avoiding further exacerbation of pancreatitis by correcting disturbances in the gut microbiota can be part of a new treatment plan for pancreatitis. The mechanisms underlying the association between the gut microbiota, metabolites, inflammatory cytokines, and pancreatitis require further studies.
Availability of data and materials
All data are publicly available.
References
Ait Chait Y, Mottawea W, Tompkins TA, Hammami R (2023) Evidence of the dysbiotic effect of psychotropics on gut microbiota and capacity of probiotics to alleviate related dysbiosis in a model of the human colon. Int J Mol Sci 24:7326. https://doi.org/10.3390/ijms24087326
Ahola-Olli AV, Würtz P, Havulinna AS, Aalto K, Pitkänen N, Lehtimäki T, Kähönen M, Lyytikäinen LP, Raitoharju E, Seppälä I, Sarin AP, Ripatti S, Palotie A, Perola M, Viikari JS, Jalkanen S, Maksimow M, Salomaa V, Salmi M, Kettunen J, Raitakari OT (2017) Genome-wide association study identifies 27 loci influencing concentrations of circulating cytokines and growth factors. Am J Hum Genet 100:40–50. https://doi.org/10.1016/j.ajhg.2016.11.007
Akshintala VS, Talukdar R, Singh VK, Goggins M (2019) The gut microbiome in pancreatic disease. Clin Gastroenterol Hepatol 17:290–295. https://doi.org/10.1016/j.cgh.2018.08.045
Alhan E, Türkyilmaz S, Erçin C, Kaklikkaya N, Kural BV (2006) Effects of omega-3 fatty acids on acute necrotizing pancreatitis in rats. Eur Surg Res 38:314–321. https://doi.org/10.1159/000094019
Bashir Y, Dobson M, Ryan BM, Duggan SN, Conlon KC (2018) The prevalence of small intestinal bacterial overgrowth in non-surgical patients with chronic pancreatitis and pancreatic exocrine insufficiency (PEI). Pancreatology 18:379–385. https://doi.org/10.1016/j.pan.2018.02.010
Bombardo M, Chen R, Malagola E, Saponara E, Hills AP, Graf R, Sonda S (2018) Inhibition of Class I histone deacetylases abrogates tumor growth factor β expression and development of fibrosis during chronic pancreatitis. Mol Pharmacol 94:793–801. https://doi.org/10.1124/mol.117.110924
Bowden J, Davey Smith G, Burgess S (2015) Mendelian randomization with invalid instruments: effect estimation and bias detection through Egger regression. Int J Epidemiol 44:512–525. https://doi.org/10.1093/ije/dyv080
Burgess S, Daniel RM, Butterworth AS, Thompson SG, EPIC-InterAct Consortium (2015) Network Mendelian randomization: using genetic variants as instrumental variables to investigate mediation in causal pathways. Int J Epidemiol 44:484–495. https://doi.org/10.1093/ije/dyu176
Cammann D, Lu Y, Cummings MJ, Zhang ML, Cue JM, Do J, Ebersole J, Chen X, Oh EC, Cummings JL, Chen J (2023) Genetic correlations between Alzheimer’s disease and gut microbiome genera. Sci Rep 13:5258. https://doi.org/10.1038/s41598-023-31730-5
Capurso G, Signoretti M, Archibugi L, Stigliano S, Delle Fave G (2016) Systematic review and meta-analysis: small intestinal bacterial overgrowth in chronic pancreatitis. United Eur Gastroenterol J 4:697–705
Davey Smith G, Hemani G (2014) Mendelian randomization: genetic anchors for causal inference in epidemiological studies. Hum Mol Genet 23:R89–R98. https://doi.org/10.1093/hmg/ddu328
Diehl GE, Longman RS, Zhang JX, Breart B, Galan C, Cuesta A, Schwab SR, Littman DR (2013) Microbiota restricts trafficking of bacteria to mesenteric lymph nodes by CX(3)CR1(hi) cells. Nature 494:116–120. https://doi.org/10.1038/nature11809
Geirnaert A, Steyaert A, Eeckhaut V, Debruyne B, Arends JB, Van Immerseel F, Boon N, Van de Wiele T (2014) Butyricicoccus pullicaecorum, a butyrate producer with probiotic potential, is intrinsically tolerant to stomach and small intestine conditions. Anaerobe 30:7074. https://doi.org/10.1016/j.anaerobe.2014.08.010
Greco MFD, Minelli C, Sheehan NA, Thompson JR (2015) Detecting pleiotropy in Mendelian randomisation studies with summary data and a continuous outcome. Stat Med 34:2926–2940. https://doi.org/10.1002/sim.6522
Huber-Ruano I, Calvo E, Mayneris-Perxachs J, Rodríguez-Peña MM, Ceperuelo-Mallafré V, Cedó L, Núñez-Roa C, Miro-Blanch J, Arnoriaga-Rodríguez M, Balvay A, Maudet C, García-Roves P, Yanes O, Rabot S, Grimaud GM, De Prisco A, Amoruso A, Fernández-Real JM, Vendrell J, Fernández-Veledo S (2022) Orally administered Odoribacter laneus improves glucose control and inflammatory profile in obese mice by depleting circulating succinate. Microbiome 10:135. https://doi.org/10.1186/s40168-022-01306-y
Jandhyala SM, Madhulika A, Deepika G, Rao GV, Reddy DN, Subramanyam C, Sasikala M, Talukdar R (2017) Altered intestinal microbiota in patients with chronic pancreatitis: implications in diabetes and metabolic abnormalities. Sci Rep 7:43640. https://doi.org/10.1038/srep43640
Kowalski K, Żebrowska-Różańska P, Karpiński P, Kujawa D, Łaczmański Ł, Samochowiec J, Chęć M, Piotrowski P, Misiak B (2023) Profiling gut microbiota signatures associated with the deficit subtype of schizophrenia: findings from a case-control study. Prog Neuropsychopharmacol Biol Psychiatry 127:110834. https://doi.org/10.1016/j.pnpbp.2023.110834
Kurilshikov A, Medina-Gomez C, Bacigalupe R, Radjabzadeh D, Wang J, Demirkan A et al (2021) Large-scale association analyses identify host factors influencing human gut microbiome composition. Nat Genet 53:156–165. https://doi.org/10.1038/s41588-020-00763-1
Kurki MI, Karjalainen J, Palta P, Sipilä TP, Kristiansson K, Donner KM et al (2023) FinnGen provides genetic insights from a well-phenotyped isolated population. Nature 613:508–518. https://doi.org/10.1038/s41586-022-05473-8
Li M, van Esch BCAM, Wagenaar GTM, Garssen J, Folkerts G, Henricks PAJ (2018) Pro- and anti-inflammatory effects of short-chain fatty acids on immune and endothelial cells. Eur J Pharmacol 831:52–59. https://doi.org/10.1016/j.ejphar.2018.05.003
Li H, Xie J, Guo X, Yang G, Cai B, Liu J, Yue M, Tang Y, Wang G, Chen S, Guo J, Qi X, Wang D, Zheng H, Liu W, Yu H, Wang C, Zhu SJ, Guo F (2022a) Bifidobacterium spp. and their metabolite lactate protect against acute pancreatitis via inhibition of pancreatic and systemic inflammatory responses. Gut Microbes 14:2127456. https://doi.org/10.1080/19490976.2022.2127456
Li J, Chen Z, Li L, Lai T, Peng H, Gui L, He W (2022b) Interleukin-6 is better than C-reactive protein for the prediction of infected pancreatic necrosis and mortality in patients with acute pancreatitis. Front Cell Infect Microbiol 12:933221. https://doi.org/10.3389/fcimb.2022.933221
Li L, Liu Q, Le C, Zhang H, Liu W, Gu Y, Yang J, Zhang X (2023) Toll-like receptor 2 deficiency alleviates acute pancreatitis by inactivating the NF-κB/NLRP3 pathway. Int Immunopharmacol 121:110547. https://doi.org/10.1016/j.intimp.2023.110547
Liu X, Tong X, Zou Y, Lin X, Zhao H, Tian L, Jie Z, Wang Q, Zhang Z, Lu H, Xiao L, Qiu X, Zi J, Wang R, Xu X, Yang H, Wang J, Zong Y, Liu W, Hou Y, Zhu S, Jia H, Zhang T (2022) Mendelian randomization analyses support causal relationships between blood metabolites and the gut microbiome. Nat Genet 54:52–61. https://doi.org/10.1038/s41588-021-00968-y
Lou J, Jiang Y, Rao B, Li A, Ding S, Yan H, Zhou H, Liu Z, Shi Q, Cui G, Yu Z, Ren Z (2020) Fecal microbiomes distinguish patients with autoimmune hepatitis from healthy individuals. Front Cell Infect Microbiol 10:342. https://doi.org/10.3389/fcimb.2020.00342
Maev IV, Levchenko AI, Galeeva JS, Andreev DN, Osipenko JV, Bordin DS, Ilyina EN (2023) Comparative analysis of the intestinal microbiota in patients with exocrine pancreatic insufficiency of various severity. Ter Arkh 95:130–139. https://doi.org/10.26442/00403660.2023.02.202056
Mi W, Hu Z, Xu L, Bian X, Lian W, Yin S, Zhao S, Gao W, Guo C, Shi T (2022) Quercetin positively affects gene expression profiles and metabolic pathway of antibiotic-treated mouse gut microbiota. Front Microbiol 13:983358. https://doi.org/10.3389/fmicb.2022.983358
Mirji G, Worth A, Bhat SA, El Sayed M, Kannan T, Goldman AR, Tang HY, Liu Q, Auslander N, Dang CV, Abdel-Mohsen M, Kossenkov A, Stanger BZ, Shinde RS (2022) The microbiome-derived metabolite TMAO drives immune activation and boosts responses to immune checkpoint blockade in pancreatic cancer. Sci Immunol 7:eabn0704. https://doi.org/10.1126/sciimmunol.abn0704
Navarro MA, Carpintero R, Acín S, Arbonés-Mainar JM, Calleja L, Carnicer R, Surra JC, Guzmán-García MA, González-Ramón N, Iturralde M, Lampreave F, Piñeiro A, Osada J (2005) Immune-regulation of the apolipoprotein A-I/C-III/A-IV gene cluster in experimental inflammation. Cytokine 31:52–63. https://doi.org/10.1016/j.cyto.2005.03.002
Pang M, Zhuang S (2010) Histone deacetylase: a potential therapeutic target for fibrotic disorders. J Pharmacol Exp Ther 335:266–272. https://doi.org/10.1124/jpet.110.168385
Perry RJ, Peng L, Barry NA, Cline GW, Zhang D, Cardone RL, Petersen KF, Kibbey RG, Goodman AL, Shulman GI (2016) Acetate mediates a microbiome-brain-β-cell axis to promote metabolic syndrome. Nature 534:213–217. https://doi.org/10.1038/nature18309
Singh V, Lee G, Son H, Koh H, Kim ES, Unno T, Shin JH (2022) Butyrate producers, “The Sentinel of Gut”: their intestinal significance with and beyond butyrate, and prospective use as microbial therapeutics. Front Microbiol 13:1103836. https://doi.org/10.3389/fmicb.2022.1103836
Sonika U, Goswami P, Thakur B, Yadav R, Das P, Ahuja V, Saraya A (2017) Mechanism of increased intestinal permeability in acute pancreatitis: alteration in tight junction proteins. J Clin Gastroenterol 51:461–466. https://doi.org/10.1097/MCG.0000000000000612
Sun L, Xiu M, Wang S, Brigstock DR, Li H, Qu L, Gao R (2018) Lipopolysaccharide enhances TGF-β1 signalling pathway and rat pancreatic fibrosis. J Cell Mol Med 22:2346–2356. https://doi.org/10.1111/jcmm.13526
Tan C, Ling Z, Huang Y, Cao Y, Liu Q, Cai T, Yuan H, Liu C, Li Y, Xu K (2015) Dysbiosis of intestinal microbiota associated with inflammation involved in the progression of acute pancreatitis. Pancreas 44:868–875. https://doi.org/10.1097/MPA.0000000000000355
Tanţău A, Leucuţa DC, Tanţău M, Boţan E, Zaharie R, Mândruţiu A, Tomuleasa IC (2021) Inflammation, tumoral markers and interleukin-17, -10, and -6 profiles in pancreatic adenocarcinoma and chronic pancreatitis. Dig Dis Sci 66:3427–3438. https://doi.org/10.1007/s10620-020-06700-w
Tao X, Guo F, Zhou Q, Hu F, Xiang H, Xiao GG, Shang D (2019) Bacterial community mapping of the intestinal tract in acute pancreatitis rats based on 16S rDNA gene sequence analysis. RSC Adv 9:5025–5036. https://doi.org/10.1039/c8ra09547g
Tao Y, Shao F, Cai M, Liu Z, Peng Y, Huang Q, Meng F (2021) Activated pancreatic stellate cells enhance the Warburg effect to cause the malignant development in chronic pancreatitis. Front Oncol 11:714598. https://doi.org/10.3389/fonc.2021.714598
Thomas RM, Jobin C (2020) Microbiota in pancreatic health and disease: the next frontier in microbiome research. Nat Rev Gastroenterol Hepatol 17:53–64. https://doi.org/10.1038/s41575-019-0242-7
VanderWeele TJ (2016) Mediation analysis: a practitioner’s guide. Annu Rev Public Health 37:17–32. https://doi.org/10.1146/annurev-publhealth-032315-021402
Verbanck M, Chen CY, Neale B, Do R (2018) Detection of widespread horizontal pleiotropy in causal relationships inferred from Mendelian randomization between complex traits and diseases. Nat Genet 50:693–698. https://doi.org/10.1038/s41588-018-0099-7
Xu JJ, Meng YT, Zou WB, Zhao JL, Fang X, Zhang Y, Zhou W, Zhang L, Wang KX, Hu LH, Liao Z, Zhou CH, Zou DW (2023) Cross-sectional evaluation of gut microbial-host cometabolites in patients with chronic pancreatitis. J Dig Dis 24:51–59. https://doi.org/10.1111/1751-2980.13162
Xue J, Sharma V, Hsieh MH, Chawla A, Murali R, Pandol SJ, Habtezion A (2015) Alternatively activated macrophages promote pancreatic fibrosis in chronic pancreatitis. Nat Commun 6:7158. https://doi.org/10.1038/ncomms8158
Yu H, Wan X, Yang M, Xie J, Xu K, Wang J, Wang G, Xu P (2023) A large-scale causal analysis of gut microbiota and delirium: A Mendelian randomization study. J Affect Disord 329:64–71. https://doi.org/10.1016/j.jad.2023.02.078
Zhang L, Zi L, Kuang T, Wang K, Qiu Z, Wu Z, Liu L, Liu R, Wang P, Wang W (2023) Investigating causal associations among gut microbiota, metabolites, and liver diseases: a Mendelian randomization study. Front Endocrinol 14:1159148. https://doi.org/10.3389/fendo.2023.1159148
Acknowledgements
All authors are grateful to the four public databases mentioned in the article.
Funding
Financial support for this study came from the National Natural Science Foundation of China (Key Programme) (Grant nos. 82330016 [Z.L.] and 82270674 [Y.C.]) and the Youth Fund Project of the National Natural Science Foundation of China (Grant no. 81900589 [Y.C.]).
Author information
Authors and Affiliations
Contributions
Yifan Qiu, Jun Ye, and Yu Cao designed the study, conducted statistical analyses, and wrote the first draft of the manuscript. Jinjin Xie, Xiaotong Mao, Yahui Wang, and Yilong Liu played roles in acquisition and analyses of the data. Xue Fang, Qian Fang, Yangyang Qian, and Wenbin Zou participated in data interpretation. All authors revised and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
All authors declare no financial relationships with any organisations that might have an interest in this work and no other relationships or activities that could appear to have influenced this work.
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Communicated by Shuhua Xu.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Qiu, YF., Ye, J., Xie, JJ. et al. Pancreatitis affects gut microbiota via metabolites and inflammatory cytokines: an exploratory two-step Mendelian randomisation study. Mol Genet Genomics 299, 36 (2024). https://doi.org/10.1007/s00438-024-02125-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00438-024-02125-6