Abstract
DNA methylation is a fundamental epigenetic process and have a critical role in many biological processes. The study of DNA methylation at a large scale of genomic levels is widely conducted by several techniques that are next-generation sequencing (NGS)-based methods. Methylome data revealed by DNA methylation next-generation sequencing (mNGS), should be always verified by another technique which they usually have a high cost. In this study, we offered a low-cost approach to corroborate the mNGS data. In this regard, mNGS was performed on 6 colorectal cancer (case group) and 6 healthy individual colon tissue (control group) samples. An R-script detected differentially methylated regions (DMRs), was further validated by high resolution melting (MS-HRM) analysis. After analyzing the data, the algorithm found 194 DMRs. Two locations with the highest level of methylation difference were verified by MS-HRM, which their results were in accordance with the mNGS. Therefore, in the present study, we suggested MS-HRM as a simple, accurate and low-cost method, useful for confirming methylation sequencing results.


Similar content being viewed by others
Data availability
Not applicable.
Code availability
Not applicable.
References
Anderson BW, Suh Y-S, Choi B, Lee H-J, Yab TC, Taylor W et al (2018) Detection of gastric cancer with novel methylated DNA markers: discovery, tissue validation, and pilot testing in plasma. Clin Cancer Res. https://doi.org/10.1158/1078-0432.CCR-17-3364
Consortium B (2016) Quantitative comparison of DNA methylation assays for biomarker development and clinical applications. Nat Biotechnol. 34(7):726–37. https://doi.org/10.1038/nbt.3605
Dor Y, Cedar H (2018) Principles of DNA methylation and their implications for biology and medicine. Lancet. https://doi.org/10.1016/S0140-6736(18)31268-6
Hussmann D, Hansen LL (2018) Methylation-sensitive high resolution melting (MS-HRM). Methods Mol Biol 1708:551–571. https://doi.org/10.1007/978-1-4939-7481-8_28
Huynh LH, Bui PTK, Nguyen TTN, Nguyen HT (2017) Developing a high resolution melting method for genotyping and predicting association of SNP rs353291 with breast cancer in the Vietnamese population. Biomed Res Ther 4(12):1812–1831
Kerachian MA, Kerachian M (2018) Long interspersed nucleotide element-1 (LINE-1) methylation in colorectal cancer. Clin Chim Acta. https://doi.org/10.1016/j.cca.2018.11.018
Kerachian MA, Javadmanesh A, Azghandi M, Shariatpanahi AM, Yassi M, Davodly ES et al (2020) Crosstalk between DNA methylation and gene expression in colorectal cancer, a potential plasma biomarker for tracing this tumor. Sci Rep 10(1):1–13
Kisiel JB, Dukek BA, Kanipakam RV, Ghoz HM, Yab TC, Berger CK et al (2018) Hepatocellular carcinoma detection by plasma methylated DNA: discovery, phase i pilot, and phase II clinical validation. Hepatology. https://doi.org/10.1002/hep.30244
Klutstein M, Nejman D, Greenfield R, Cedar H (2016) DNA methylation in cancer and aging. Can Res 76(12):3446–3450
Li LC, Dahiya R (2002) MethPrimer: designing primers for methylation PCRs. Bioinformatics 18(11):1427–1431. https://doi.org/10.1093/bioinformatics/18.11.1427
Lu J, Johnston A, Berichon P, Ru KL, Korbie D, Trau M (2017) PrimerSuite: a high-throughput web-based primer design program for multiplex bisulfite PCR. Sci Rep. 7:41328. https://doi.org/10.1038/srep41328
Ma M, Zhu H, Zhang C, Sun X, Gao X, Chen G (2015) Liquid biopsy—ctDNA detection with great potential and challenges. Ann Transl Med 3:16
Masser DR, Stanford DR, Hadad N, Giles CB, Wren JD, Sonntag WE et al (2016) Bisulfite oligonucleotide-capture sequencing for targeted base-and strand-specific absolute 5-methylcytosine quantitation. Age 38(3):49
Meissner A, Gnirke A, Bell GW, Ramsahoye B, Lander ES, Jaenisch R (2005) Reduced representation bisulfite sequencing for comparative high-resolution DNA methylation analysis. Nucleic Acids Res 33(18):5868–5877
Migheli F, Stoccoro A, Coppede F, Omar WAW, Failli A, Consolini R et al (2013) Comparison study of MS-HRM and pyrosequencing techniques for quantification of APC and CDKN2A gene methylation. PLoS One 8(1):e52501
Moore LD, Le T, Fan G (2013) DNA methylation and its basic function. Neuropsychopharmacology 38(1):23–38
Reed K, Poulin ML, Yan L, Parissenti AM (2010) Comparison of bisulfite sequencing PCR with pyrosequencing for measuring differences in DNA methylation. Anal Biochem 397(1):96–106. https://doi.org/10.1016/j.ab.2009.10.021
Rokni P, Shariatpanahi AM, Sakhinia E, Kerachian MA (2018) BMP3 promoter hypermethylation in plasma-derived cell-free DNA in colorectal cancer patients. Genes Genomics 40(4):423–428
Sestakova S, Salek C, Remesova H (2019) DNA methylation validation methods: a coherent review with practical comparison. Biol Proced Online 21:19. https://doi.org/10.1186/s12575-019-0107-z
Soto J, Rodriguez-Antolin C, Vallespin E, De Castro CJ, De Caceres II (2016) The impact of next-generation sequencing on the DNA methylation–based translational cancer research. Transl Res 169(1–18):e1
Tost J (2010) DNA methylation: an introduction to the biology and the disease-associated changes of a promising biomarker. Mol Biotechnol 44(1):71–81
Tse MY, Ashbury JE, Zwingerman N, King WD, Taylor SA, Pang SC (2011) A refined, rapid and reproducible high resolution melt (HRM)-based method suitable for quantification of global LINE-1 repetitive element methylation. BMC Res Notes 4(1):565
Tusnady GE, Simon I, Varadi A, Aranyi T (2005) BiSearch: primer-design and search tool for PCR on bisulfite-treated genomes. Nucleic Acids Res 33(1):e9. https://doi.org/10.1093/nar/gni012
Urich MA, Nery JR, Lister R, Schmitz RJ, Ecker JR (2015) MethylC-seq library preparation for base-resolution whole-genome bisulfite sequencing. Nat Protoc 10(3):475–483
Van Wesenbeeck L, Janssens L, Meeuws H, Lagatie O, Stuyver L (2018) Droplet digital PCR is an accurate method to assess methylation status on FFPE samples. Epigenetics 13(3):207–213
Wittwer CT (2009) High-resolution DNA melting analysis: advancements and limitations. Hum Mutat 30(6):857–859
Wojdacz TK, Dobrovic A (2007) Methylation-sensitive high resolution melting (MS-HRM): a new approach for sensitive and high-throughput assessment of methylation. Nucleic Acids Res 35(6):e41. https://doi.org/10.1093/nar/gkm013
Wojdacz TK, Moller TH, Thestrup BB, Kristensen LS, Hansen LL (2010) Limitations and advantages of MS-HRM and bisulfite sequencing for single locus methylation studies. Expert Rev Mol Diagn 10(5):575. https://doi.org/10.1586/erm.10.46
Worm J, Aggerholm A, Guldberg P (2001) In-tube DNA methylation profiling by fluorescence melting curve analysis. Clin Chem 47(7):1183–1189
Yassi M, Davodly ES, Shariatpanahi AM, Heidari M, Dayyani M, Heravi-Moussavi A et al (2018) DMRFusion: a differentially methylated region detection tool based on the ranked fusion method. Genomics 110(6):366–374
Acknowledgements
Our sincere thanks go to Mr. Ebrahim Pouladin and Mrs. Nafiseh Shalchi for their close support in colorectal cancer research programs. We would also give special thanks to Dr. Abdorasoul Hayatbakhsh and Dr Ghodratollah Soltani for their cooperation in this study.
Funding
This study was supported financially by Reza Radiotherapy and Oncology Center, Mashhad University of Medical Sciences (Grant number: 961906) and Iran National Science Foundation (INSF; proposal number: 93048371), Iran.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to declare.
Ethical approval
This study was approved by ethics committee of Mashhad University of Medical Sciences, Mashhad, Iran (ethic code: IR.MUMS.MEDICAL.REC.1399.642).
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Communicated by Shuhua Xu.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
438_2022_1906_MOESM1_ESM.png
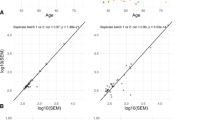
Supplementary file1 Methylation Call (MC) for case and control samples using primer A and B sets. The size of the symbols correspond to coverage (PNG 252 KB)
Rights and permissions
About this article
Cite this article
Javadmanesh, A., Mojtabanezhad Shariatpanahi, A., Shams Davodly, E. et al. MS-HRM protocol: a simple and low-cost approach for technical validation of next-generation methylation sequencing data. Mol Genet Genomics 297, 1101–1109 (2022). https://doi.org/10.1007/s00438-022-01906-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-022-01906-1