Abstract
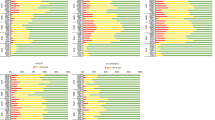
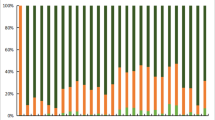
Previous studies have shown that the frequency of very important pharmacogenomic (VIP) genes varies in different populations which leads to the diversities in drug efficacy, safety, and the risk associated with adverse drug reactions (ADRs). The purpose of this study was to identify the distribution differences of VIP variants between the Li population and the other 13 populations. Based on the Pharmacogenomics Knowledgebase database (PhamGKB), we successfully genotyped 52 VIP variants within 27 genes in 200 unrelated Li population. χ2 test was used to evaluate the significant differences of genotype and allele frequencies between the Li and the other 13 populations from 1000 Genomes Project. Our study showed that the genotype frequencies of single nucleotide polymorphisms (SNPs) on KCNH2, ACE, CYP4F2, and CYP2E1 were considerably different between Li and the other 13 populations, especially in rs1805123 (KCNH2), rs4291 (ACE), rs3093105 (CYP4F2), and rs6413432 (CYP2E1) loci. Meanwhile, we found several VIP variants that might alter the drug metabolism of cisplatin-cyclophosphamide (CYP2E1), vitamin E (CYP4F2), asthma amlodipine, chlorthalidone, and lisinopril (ACE) through PharmGKB. We also identified other variants which were associated with adverse effects in isoniazid and rifampicin (CYP2E1; hepatotoxicity). The four loci rs1805123 (KCNH2), rs4291 (ACE), rs3093105 (CYP4F2), and rs6413432 (CYP2E1) provided a reliable basis for the prediction of the efficacy of certain drugs. The study complemented the existed pharmacogenomics information, which could provide theoretical basis for predicting the efficacy of certain drugs in the Li population.


Similar content being viewed by others
References
Abecasis GR, Altshuler D, Auton A, Brooks LD, Durbin RM, Gibbs RA, McVean GA (2010) A map of human genome variation from population-scale sequencing. Nature 467(7319):1061–1073. https://doi.org/10.1038/nature09534
Bardowell SA, Stec DE, Parker RS (2010) Common variants of cytochrome P450 4F2 exhibit altered vitamin E-{omega}-hydroxylase specific activity. J Nutr 140(11):1901–1906. https://doi.org/10.3945/jn.110.128579
Bose PD, Sarma MP, Medhi S, Das BC, Husain SA, Kar P (2011) Role of polymorphic N-acetyl transferase2 and cytochrome P4502E1 gene in antituberculosis treatment-induced hepatitis. J Gastroenterol Hepatol 26(2):312–318. https://doi.org/10.1111/j.1440-1746.2010.06355.x
Cederbaum AI, Lu Y, Wu D (2009) Role of oxidative stress in alcohol-induced liver injury. Arch Toxicol 83(6):519–548. https://doi.org/10.1007/s00204-009-0432-0
Ding Y, He P, He N, Li Q, Sun J, Yao J, Jin T (2016) Genetic polymorphisms of pharmacogenomic VIP variants in Li nationality of southern China. Environ Toxicol Pharmacol 42:237–242. https://doi.org/10.1016/j.etap.2016.02.003
Gabriel S, Ziaugra L, Tabbaa D (2009) SNP genotyping using the Sequenom MassARRAY iPLEX platform. Curr Protoc Hum Genet 15:14. https://doi.org/10.1002/0471142905.hg0212s60 (Chapter 2, Unit 2.12)
Holman EA (2012) Acute stress and cardiovascular health: is there an ACE gene connection? J Trauma Stress 25(5):592–597. https://doi.org/10.1002/jts.21746
Irvin MR, Lynch AI, Kabagambe EK, Tiwari HK, Barzilay JI, Eckfeldt JH, Arnett DK (2010) Pharmacogenetic association of hypertension candidate genes with fasting glucose in the GenHAT Study. J Hypertens 28(10):2076–2083. https://doi.org/10.1097/HJH.0b013e32833c7a4d
Katara P (2014) Single nucleotide polymorphism and its dynamics for pharmacogenomics. Interdiscip Sci 6(2):85–92. https://doi.org/10.1007/s12539-013-0007-x
Kaye JB, Schultz LE, Steiner HE, Kittles RA, Cavallari LH, Karnes JH (2017) Warfarin pharmacogenomics in diverse populations. Pharmacotherapy 37(9):1150–1163. https://doi.org/10.1002/phar.1982
Khrunin A, Ivanova F, Moisseev A, Khokhrin D, Sleptsova Y, Gorbunova V, Limborska S (2012) Pharmacogenomics of cisplatin-based chemotherapy in ovarian cancer patients of different ethnic origins. Pharmacogenomics 13(2):171–178. https://doi.org/10.2217/pgs.11.140
Kim TH, Chang HS, Park SM, Nam BY, Park JS, Rhim T, Shin HD (2008) Association of angiotensin I-converting enzyme gene polymorphisms with aspirin intolerance in asthmatics. Clin Exp Allergy 38(11):1727–1737. https://doi.org/10.1111/j.1365-2222.2008.03082.x
Krähenbühl-Melcher A, Schlienger R, Lampert M, Haschke M, Drewe J, Krähenbühl S (2007) Drug-related problems in hospitals: a review of the recent literature. Drug Saf 30(5):379–407. https://doi.org/10.2165/00002018-200730050-00003
Lander ES, Linton LM, Birren B, Nusbaum C, Zody MC, Baldwin J, Szustakowki J (2001) Initial sequencing and analysis of the human genome. Nature 409(6822):860–921. https://doi.org/10.1038/35057062
Liu Y, Li H, Cao K, Liu J, Sun Y, Wu J, Jin T (2020) Genetic variation of pharmacogenomic VIP variants in Zhuang nationality of southern China. Pharmacogenomics J. https://doi.org/10.1038/s41397-020-0177-y
Neafsey P, Ginsberg G, Hattis D, Johns DO, Guyton KZ, Sonawane B (2009) Genetic polymorphism in CYP2E1: population distribution of CYP2E1 activity. J Toxicol Environ Health B Crit Rev 12(5–6):362–388. https://doi.org/10.1080/10937400903158359
Pirmohamed M (2014) Personalized pharmacogenomics: predicting efficacy and adverse drug reactions. Annu Rev Genomics Hum Genet 15:349–370. https://doi.org/10.1146/annurev-genom-090413-025419
Rett A (1982) Genetic counselling and human handicap. Wien Klin Wochenschr 94(12):315–317
Roy B, Ghosh SK, Sutradhar D, Sikdar N, Mazumder S, Barman S (2006) Predisposition of antituberculosis drug induced hepatotoxicity by cytochrome P450 2E1 genotype and haplotype in pediatric patients. J Gastroenterol Hepatol 21(4):784–786. https://doi.org/10.1111/j.1440-1746.2006.04197.x
Shendre A, Brown TM, Liu N, Hill CE, Beasley TM, Nickerson DA, Limdi NA (2016) Race-specific influence of CYP4F2 on dose and risk of hemorrhage among warfarin users. Pharmacotherapy 36(3):263–272. https://doi.org/10.1002/phar.1717
Suarez-Kurtz G, Vargens DD, Santoro AB, Hutz MH, de Moraes ME, Pena SD, Struchiner CJ (2014) Global pharmacogenomics: distribution of CYP3A5 polymorphisms and phenotypes in the Brazilian population. PLoS ONE 9(1):e83472. https://doi.org/10.1371/journal.pone.0083472
Thorn CF, Klein TE, Altman RB (2010) Pharmacogenomics and bioinformatics: PharmGKB. Pharmacogenomics 11(4):501–505. https://doi.org/10.2217/pgs.10.15
van Engen CE, Ofman R, Dijkstra IM, van Goethem TJ, Verheij E, Varin J, Barbier M (2016) CYP4F2 affects phenotypic outcome in adrenoleukodystrophy by modulating the clearance of very long-chain fatty acids. Biochim Biophys Acta 1862(10):1861–1870. https://doi.org/10.1016/j.bbadis.2016.07.006
Wypasek E, Branicka A, Awsiuk M, Sadowski J, Undas A (2014) Genetic determinants of acenocoumarol and warfarin maintenance dose requirements in Slavic population: a potential role of CYP4F2 and GGCX polymorphisms. Thromb Res 134(3):604–609. https://doi.org/10.1016/j.thromres.2014.06.022
Yang ZL, Guo Y, Gao L, Wang Y, Dong YL, Tang WR, Xiao CJ (2004) Hypertension survey in Yi ethnic group in Yunnan Province, China. Zhonghua Liu Xing Bing Xue Za Zhi 25(9):817
Yap DYH, Tam CH, Yung S, Wong S, Tang CSO, Mok TMY, Chan TM (2020) Pharmacokinetics and pharmacogenomics of mycophenolic acid and its clinical correlations in maintenance immunosuppression for lupus nephritis. Nephrol Dial Transplant 35(5):810–818. https://doi.org/10.1093/ndt/gfy284
Yousef AM, Bulatova NR, Newman W, Hakooz N, Ismail S, Qusa H, Al-Diab O (2012) Allele and genotype frequencies of the polymorphic cytochrome P450 genes (CYP1A1, CYP3A4, CYP3A5, CYP2C9 and CYP2C19) in the Jordanian population. Mol Biol Rep 39(10):9423–9433. https://doi.org/10.1007/s11033-012-1807-5
Yunus Z, Liu L, Wang H, Zhang L, Li X, Geng T, Chen C (2013) Genetic polymorphisms of pharmacogenomic VIP variants in the Kyrgyz population from northwest China. Gene 529(1):88–93. https://doi.org/10.1016/j.gene.2013.07.078
Acknowledgements
We thank all participants in our study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest.
Additional information
Communicated by Shuhua Xu.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yang, S., Dou, X., Wang, Z. et al. Genetic variation of pharmacogenomic VIP variants in the Chinese Li population: an updated research. Mol Genet Genomics 297, 407–417 (2022). https://doi.org/10.1007/s00438-022-01855-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-022-01855-9