Abstract
The Ataxia-telangiectasia-mutated (ATM) gene in humans was identified as the basis of a rare autosomal disorder leading to cancer susceptibility and is now well known as an important signal transducer in response to DNA damage. An approach to understanding the conserved functions of this gene is provided by the model system, Caenorhabditis elegans. In this paper we describe the structure and loss of function phenotype of the ortholog atm-1. Using bioinformatic and molecular analysis we show that the atm-1 gene was previously misannotated. We find that the transcript is in fact a product of three gene predictions, Y48G1BL.2 (atm-1), K10E9.1, and F56C11.4 that together make up the complete coding region of ATM-1. We also characterize animals that are mutant for two available knockout alleles, gk186 and tm5027. As expected, atm-1 mutant animals are sensitive to ionizing radiation. In addition, however, atm-1 mutants also display phenotypes associated with genomic instability, including low brood size, reduced viability and sterility. We document several chromosomal fusions arising from atm-1 mutant animals. This is the first time a mutator phenotype has been described for atm-1 in C. elegans. Finally we demonstrate the use of a balancer system to screen for and capture atm-1-derived mutational events. Our study establishes C. elegans as a model for the study of ATM as a mutator potentially leading to the development of screens to identify therapeutic targets in humans.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
A wide range of proteins function to protect the genome from the consequences of DNA double strand breaks (DSBs). DSBs can occur as the result of external sources such as ionizing radiation or internal causes such as defects in the repair process. The inability to properly respond to DSBs can have catastrophic consequences for the cell, a fact highlighted by the prevalence of DSB repair proteins that are mutated in cancer (Lee and Paull 2005; Negrini et al. 2010). Consequently, most known organisms have evolved a comprehensive suite of mechanisms to repair DSBs appropriately, whether they have occurred by accident or design. A better understanding of DSB repair is therefore essential for understanding the mechanisms behind genome instability for the development of therapeutic approaches to prevent it.
One key player in the DNA damage response (DDR) to DSBs is the phosphatidylinositol 3-kinase (PI3K) related protein kinase ATM (Ataxia-telangiectasia-mutated). Mutation of ATM in humans can lead to Ataxia-telangiectasia (A-T), a rare autosomal recessive disorder, one of the hallmarks of which is a predisposition to developing cancer (Kurz and Lees-Miller 2004; Lavin 2008; Lavin et al. 2006; Lavin and Shiloh 1997). ATM is activated in response to DSBs and serves as a signal transducer, phosphorylating downstream targets to activate cell-cycle checkpoints, DNA damage response (DDR) proteins, and apoptosis pathways (Lavin 2008; Shiloh 2006). Human cells mutant for ATM are known to be defective in cell-cycle checkpoint activation, sensitive to DSB-inducing agents, and display chromosomal instability (Rotman and Shiloh 1998; Kojis et al. 1989; Pandita et al. 1995; Metcalfe et al. 1996; Xia et al. 1996).
C. elegans is a powerful model for investigation of the DNA repair and has a number of conserved DDR mechanisms (O’Neil and Rose 2006; Lemmens and Tijsterman 2011). The C. elegans ATM protein, ATM-1, is known to function in the DDR and DSB repair and its activity and functional interactions are conserved between worms and humans (Lemmens and Tijsterman 2011; Lee et al. 2010). Mutations in the atm-1 gene result in animals that are sensitive to DSB inducing agents such as IR and, to a lesser extent, UV-C (Couteau and Zetka 2011; Garcia-Muse and Boulton 2005; Stergiou et al. 2007; Weidhaas et al. 2006). Genomic instability associated with mutations in ATM-1 has not, however, been documented. Furthermore ATM-1 appears structurally unique in that it is predicted to be a very small polypeptide when compared to other ATM homologs.
In this study we present the characterization of the atm-1 locus in C. elegans. Using bioinformatic and molecular approaches we have revised the gene model for the atm-1 and show that the full-length gene is a fusion of three predicted ORFs. We further describe the loss-of-function phenotype for this gene and identify a previously unrecognized mutator phenotype.
Materials and methods
Strains
C. elegans strains were maintained as described previously (Brenner 1974). Strains: VC2010 wild type, VC655 brd-1(gk297), FX5027 atm-1(tm5027), VC381 atm-1(gk186), KR4941 atm-1(gk186) (outcrossed 10 times), BC2200 eT1(III)/dpy-18(e364);eT1(V)/unc-46(e177), KR5060 atm-1(gk186); eT1(III)/dpy-18(e364);eT1(V)/unc-46(e177).
Bioinformatic analysis
All nematode homology comparisons were performed using BLASTN and BLASTP (NCBI). Illumina-generated RNA-Seq data from an L1 synchronized cDNA library were analyzed for the atm-1 transcript structure. Sequence reads were aligned to the C. elegans reference sequence WS200 using Bowtie (Langmead 2010) and Tophat (Trapnell et al. 2009). Default options were used, except insert size (-r) was set to 300 and the minimum intron size (-i) to 40. The alignment file was visualized using IGV (Integrated Genomics Viewer) (Robinson et al. 2011).
cDNA library construction
A healthy mixed stage population of each strain was harvested for RNA extraction. The animals were pelleted by centrifugation and ~0.5 mL of packed worms resuspended in 2 mL of Trizol (Invitrogen, Catalog Number 15596-018). The worm/Trizol mix was vortexed, subjected to freeze–thaw, and incubated in a 37°C water bath. Nucleic acids were extracted with 1 mL chloroform followed by centrifugation and isolation of the supernatant. 1 mL isopropanol was added to the supernatant, mixed, and re-centrifugated. The nucleic acid pellet was washed with 0.5 mL RNAse-free 75% ethanol and allowed to dry and resuspended with DEPC-treated water. RNA quality was estimated on a 1% agarose gel. The nucleic acids were treated with DNase using the Fermentus DNase I, RNase-free kit and protocol (#EN0521). DNase was removed with phenol–chloroform followed by chloroform to isolate total RNA. RNA was precipitated overnight at three volumes of 100% ethanol to sample ratio with the addition of 0.3 M final concentration sodium acetate. Following centrifugation and removal of supernatant, the RNA pellet was washed with 0.5 mL 75% ethanol and resuspended in DEPC-treated water. cDNA was generated using the RevertAid H Minus M-MuLV Reverse Transcriptase kit from Fermentas using poly-dT primers (#EP0451). The quality and concentration of the total RNA and cDNA were measured using a NanoDrop spectrophotometer.
Primer design and polymerase chain reaction (PCR)
Primers were designed using the Primer3 website interface http://frodo.wi.mit.edu/primer3/. cDNA PCRs were conducted using New England BioLabs Inc Phusion High-Fidelity DNA Polymerase to ensure high fidelity products for sequencing. Suggested Phusion protocol for PCR cycling was used: 98°C denaturing, 72°C elongation at 45 s/kb, primer specific annealing temperature (between 56–60°C) and cycled for 34 cycles. 10 mM stock dNTP solution was mixed from 100 mM stock solutions of each dNTP bought from Applied Biological Materials (abm) Inc. A 20 μL reaction consisted of 4 μL 5× HF buffer, 0.5 μL 10 mM dNTPs stock solution, 0.4 μL of each 10 μM primer stock, 100 ng cDNA, 0.1 μL Phusion polymerase, and 14.4 μL of double distilled water.
Ionizing radiation sensitivity assay
For each strain, 30 adult animals were plated on 3 plates (10 worms per plate) and allowed to lay eggs for 2 h (giving approximately 50 eggs per plate.) These worms were synchronized to 1-day-old adults and subjected to different IR dosages. IR was applied by the TORREX 150D X-Ray Inspection System. After IR, worms were allowed to rest in 20°C for 20 h. 10 plates with 3 worms each were plated for each dose and a 4- to 5-h brood was collected. The number of unhatched eggs and adults was scored 24 and 72 h post brood collection, respectively. % survival was calculated by dividing the total number of adults by the total progeny (adults + unhatched eggs). Standard error was used for calculating statistical error.
DAPI staining and microscopy
20–30 adult worms for each strain were picked onto a watch glass containing 5 μL of M9 buffer. 250 μL of 150 mM DAPI in 96% ethanol was added and allowed to evaporate in the dark at room temperature for 1–2 h. 500 μL of M9 was used to destain the worms for 2 h at room temperature. Worms were mounted 3% agarose pads. Diakinetic chromosomes were observed in oocytes located before the spermatheca using a Zeiss Axioscope fluorescent microscope with 40× objective using OpenLab software.
Quantifying genomic instability
20 atm-1(gk186) L4 worms were individually plated and the subsequent progeny scored for total progeny and the presence of males. A single L4 worm from each line was plated to propagate the line. This was repeated for 20 generations. Total brood was scored and the presences of males (>10% of the total population) were recorded.
Results
C. elegans ATM-1 has a conserved kinase domain but lacks upstream sequences
Human ATM is a large protein (3056aa) with a highly conserved kinase domain and a large, poorly conserved, N-terminal region (Fig. 1a; data not shown). In contrast C. elegans ATM-1 (Ce-ATM-1) is predicted to be a relatively small 649AA polypeptide similar in structure to the C-terminal portion of ATM, having the conserved kinase domain and FATC domains (Fig. 1b). ATM-1 is, however, lacking domains known to be present in other ATM orthologs, including the FAT and TAN domains that are typically upstream of the kinase domain (Bosotti et al. 2000; Seidel et al. 2008) (Fig. 1b). The discrepancies between Ce-ATM-1 and other ATM orthologs led us to investigate its structure in more detail.
C. elegans ATM-1 is much smaller than other ATM orthologs including those in closely related nematode species. a Distant matrix tree of ATM orthologs based on conservation of the kinase domain. Percentage similarity to C. elegans is indicated. b Schematic representation of protein structure of human ATM orthologs from other species. TAN Tel1/ATM N-Terminal motif, FAT FRAP–ATM–TRAP domain, PIK phosphatidylinositol 3-kinase, FATC C-Terminal FAT domain
Cross-species analysis of nematode genomes suggests an extended C. elegans ATM-1 structure
We used BLASTP to compare the predicted sequences of ATM-1 in closely related Caenorhabditis species; C. briggsae (Cbg), C. remanei (Cr) and C. brenneri (Cbn) (Fig. 2). Interestingly the predicted sizes of Cbn-ATM-1 (1812aa) and Cr-ATM-1 (2161aa) are significantly larger than both Ce-ATM-1 (649aa) and Cbg-ATM-1 (639aa) (Fig. 1b). Furthermore, BLASTP comparison of Cbn-ATM-1 revealed matches not only with Ce-ATM-1 but also with two gene predictions located further upstream, K10E9.1 and F56C11.4 (Fig. 2a). These three gene predictions might therefore comprise a single, extended, atm-1 transcript. Likewise the equivalent genomic region in C. briggase includes three distinct gene predictions (Fig. 2b) while in C. remanei there are only two (Fig. 1c). The gene order is maintained between these nematode species. It is likely that the Cbr-atm-1 and Cr-atm-1 gene models are also misannotated.
Evidence for an extended atm-1 transcript in closely related nematode species. a–d Schematic representations of gene predictions in the genomic region of atm-1 in four nematode genomes. a In C. elegans the region contains three gene predictions. b C. briggsae also contains three gene predictions in the equivalent region. c In C. remanei two gene predictions are annotated, the genes corresponding to K10E9.1 and atm-1 are represented as a single gene, RP31208. d A single gene prediction is annotated in. C. brenneri. Figure is to scale
Transcriptome and EST analysis support the extended gene structure
Three annotated expressed sequence tags (ESTs) from the Kohara group (Tabara et al. 1996) provided evidence for an extended atm-1 transcript. The ESTs yk1752d02 and yk220a2 both span the junction between the K10E9.1 and atm-1 gene predictions while a single EST, yk1279h03, links F56C11.4 to the same transcript (Fig. 2b). To obtain further evidence of an extended Ce-atm-1 transcript we used available deep RNA-seq data to map transcript sequences to F56C11.4, K10E9.1 and atm-1 (Our unpublished data). From the data we identified four tags that spanned the junction between F56C11.6 and K10E9.1, and a further four tags that spanned the junction between K10E9.1 and atm-1 (Fig. 3c). Thus, we conclude that F56C11.4, K10E9.1 and atm-1 produce a single transcript.
Revised C. elegans atm-1 gene structure. a A schematic showing the original gene model predictions for the genomic region of atm-1. Position of primer pairs used to confirm transcript junctions by PCR is shown below the model (not to scale). Regions deleted in the knockout alleles gk186 and tm5027 are depicted as dashed bars. b Schematic of ESTs confirming the extended atm-1 gene model. c Representation of RNA-seq mapping data. Sequence tags confirming merging between gene predictions are highlighted in black. d Schematic showing the revised transcript structure for Ce-atm-1. Changes to exonic structure are listed below the figure. e Schematic of the extended ATM-1 protein. Figure is to scale
The gene predictions F56C11.4, K10E9.1, and Y48G1BL.2 (atm-1) produce a single mRNA transcript
To independently confirm the structure of the extended atm-1 transcript a wild-type (N2) cDNA library was probed with two sets of PCR primers designed to amplify the predicted junctions between adjacent open reading frames (Arrows in Fig. 3a). Both PCR assays amplified a sequence of expected size from the WT cDNA (data not shown). Full length sequencing of the cDNA transcript confirmed the extended structure for atm-1 (Fig. 3d). The majority of the transcript sequence aligned with the predicted gene models as expected. However, a previously unidentified 5′ splice site disrupts the annotated stop codon of F56C11.4. The terminal exon of F56C11.4 splices to a previously unknown exon of 163nt located 433nt upstream of the predicted first exon of K10E9.1. Exon 1 of K10E9.1 has additional exonic sequences added at both 5′ (9nt) and 3′ (42nt). Finally an alternative 5′ splice site results in removal of 42nt from the end of the last predicted exon of K10E9.1 and a 3′ splice site removes 30nt from the beginning of the first predicted exon of atm-1, joining these two ORF’s. All transcript modifications were also present in the RNA-seq data (data not shown). The revised atm-1 gene model produces an ORF of 7,137nt that is predicted to encode a protein of 2,378AA (Fig. 3e). Domain analysis of the revised ATM-1 protein prediction did not, however, reveal any additional conserved domains beyond the kinase and FATC domains (Fig. 3e).
Transcript analysis of the gk186 gene reveals two potential transcripts
Two knockout mutations of atm-1 are available, gk186 and tm5027. atm-1(tm5027) is a 513 bp deletion that disrupts exon 23 and deletes exon 24, resulting in 4 missense amino acids and a premature stop in exon 23 (Fig. 3a). The tm5027 mutant transcript results in a truncated protein lacking the kinase domain of the protein and is therefore expected to be null (Figure S1). atm-1(gk186) is a 548 bp deletion that removes the 3′ intron splice site for the 22nd exon of the revised atm-1 gene model (Fig. 3a), resulting in the splicing of exons 21–23 (Figure S1) (Garcia-Muse and Boulton 2005; Stergiou et al. 2007). The mutant transcript is predicted to produce a frame-shift mutation resulting in a truncated protein lacking the kinase domain of the protein. However, a second mRNA species was present in cDNA from gk186 animals (Figure S1). Sequence analysis of PCR fragments from gk186 revealed that an alternative splice acceptor allows splicing of exon 24, generating a transcript that maintains the kinase domain (Figure S1; data not shown).
Animals carrying either gk186 or tm5027 are sensitive to IR
In order to determine the radiation sensitivity of the mutant animals, we treated animals from strains carrying gk186 or tm5027 with ionizing radiation (IR). We measured the survival frequency of the progeny of parental hermaphrodites exposed to IR. Wild type and mutants in a known IR-sensitivity gene, brd-1(gk297), were used as controls. The results are shown in Fig. 4. Approximately 90% of the wild-type F1’s hatched and survived at the highest dose tested (45 Gy), compared to 2.5% F1 survival in the brd-1 mutant animals. The atm-1 mutant animals had an intermediate survival frequency that was significantly different than the wild type. Of the two atm-1 mutants, gk186 was significantly more sensitive at the higher dose than tm5027. Thus, the alternative transcript produced in gk186, which contains the kinase domain does not contribute to IR resistance. The knockout allele, tm5027 deletes the kinase domain and can be assumed to exhibit the loss-of-function phenotype. Thus, we conclude that the data shown in Fig. 4 represent the null phenotype for atm-1.
atm-1 mutants are sensitive to IR. Graph showing the ionizing radiation (IR) sensitivity of wild type, brd-1 mutant, and atm-1 mutant worms. Survival percentage is calculated by dividing the number of adults by the sum of arrested embryos and adults multiplied by 100. Error bars indicate the standard error
atm-1 mutant animals display the hallmarks of a mutator
Morphologically, atm-1 mutant animals appear superficially wild type. In maintaining these strains, however, we observed variation in the number and appearance of the offspring of specific individuals. Brood analysis of both mutant strains revealed erratic brood sizes. Some hermaphrodites were sterile and some produced arrested embryos. Embryonic arrest varied between 1 and 50% (Table 1). The appearance of a high incidence of male (Him) phenotype was also consistently observed. These male-producing lines could be maintained by picking individual hermaphrodites indicating that the high incidence of male (Him) phenotype was a heritable trait. In C. elegans, sterility and Him phenotypes are both indicators of genomic instability (Ahmed and Hodgkin 2000; Herman et al. 1982). It is likely that such phenotypes are arising due to the mutagenic consequence of persistent DSBs in the absence of ATM-1 function.
Quantifying independent atm-1(gk186) lines indicate inherited genomic instability
To further document the variability in the phenotype of atm-1 mutants, we examined the viability of atm-1(gk186) animals over 20 generations. 20 L4 atm-1(gk186) worms were plated individually and scored for viability, brood size, and the presence of males. Each subsequent generation was established by selecting a single L4 offspring at random. This process was repeated for 20 generations and the results are summarized in Fig. 5a and Table S1. Over the course of the experiment 8 of the 20 individual lines acquired Him mutations. 16 of 20 lines became sterile; including several that had first become Him. The progeny numbers for the different lines also fluctuated unpredictably over the 20 generations (Fig. 5a; Table S1). Together these results point to a model in which heritable mutations that confer a high incidence of male and sterile phenotypes, as well as mutations that result in loss of viability are generated in atm-1 mutant animals.
atm-1(gk186) animals display pleiotrophic phenotypes associated with genome instability. a 20 individual animals were isolated and their progeny examined over 20 subsequent generations for brood size, viability, a Him phenotype and sterility. The brood size of each animal is directly proportional to size of the circle. b–e DAPI staining of diakinetic chromosomes (arrowheads) in atm-1(gk186) animals with a dominant Him phenotype. b atm-1(gk186) mutant animals have the expected compliment of six chromosomes. c atm-1(gk186);(h2681) animals have five chromosomes. d atm-1(gk186);(h2680) animals have four chromosomes. c atm-1(gk186);(h2685) animals have three chromosomes
atm-1-derived dominant him strains have a reduced chromosome number
We reasoned that since ATM is also known to function in telomere maintenance the Him phenotype observed might also be due to chromosome fusions (Ahmed and Hodgkin 2000; Herman et al. 1982; Herman and Kari 1989). Five independent atm-1(gk186) lines that became Him were isolated and their chromosome complement assessed by visualizing diakinetic chromosomes in their ooctyes. Normal wild-type animals and freshly outcrossed atm-1(gk186) mutants have the expected complement of six pairs of chromosomes, seen as distinct spots when visualized using DAPI staining (Fig. 5b; data not shown). In all of the atm-1(gk186) Him strains examined a reduced number of diakinetic chromosomes were observed (Fig. 5c–e; Table 2). A maximum of five chromosomes was seen in these strains though animals with only four or three pairs were also observed (Fig. 5). This indicated that multiple fusions were occurring with no loss of essential genetic material. The fact that less than five chromosome pairs were seen in viable Him animals also indicated that chromosome fusions were not specific to the X chromosome. This is consistent with the interpretation that loss of function of atm-1 leads to chromosome rearrangement.
Mutations produced in an atm-1(gk186) background can be captured using a translocation balancer
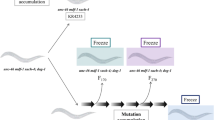
Chromosome fusions appear to be one of the consequences of loss of atm-1 function. We have also shown that fusions are likely to be occurring between autosomes also. However, the sterility and reduced brood numbers observed suggest that mutations resulting in the loss of essential genetic material are also occurring. This indicates that the mutational spectrum of atm-1 is not limited to viable chromosome fusions. To explore this further we used the eT1 balancer system (Rosenbluth and Baillie 1981) to capture mutational events induced in an atm-1(gk186) background. eT1 is a stable reciprocal translocation of the C. elegans chromosomes (Chr) III and V. Recombination is effectively suppressed in regions of Chr III and Chr V involved in the translocation and mutations present in the balanced regions are maintained heterozygously in these strains (Rosenbluth and Baillie 1981; Johnsen and Baillie 1991). Visible markers are used to follow the non-eT1 chromosomes. Loss of these markers in the viable progeny of this strain is an indication that an essential mutation has occurred within the balanced region. 1,139 atm-1(gk186); eT1 animals were plated and their progeny were screened for the absence of the visible markers, indicating that a lethal mutation had occurred. 11 independent lines harboring mutations obtained (Table S2) a number of mutational events equivalent to a 0.97% forward mutational frequency for atm-1(gk186), compared to 0.06% for the eT1 balancer system alone (Rosenbluth and Baillie 1981). This is similar mutation rate to that of dog-1 (0.9%), a known mutator that is required to maintain genome stability by resolving potentially toxic G-track secondary structures that form during replication (Cheung et al. 2002). The results show that in addition to heritable male-producing strains, non-Him-associated lethal mutations occur and can be isolated and stably maintained.
Discussion
We describe here the structure of the atm-1 gene in C. elegans and its loss of function phenotype. Revealing the structure and function of ATM is an important step in understanding the human disease A-T specifically and the DNA damage response in general. Such functional studies require a solid platform in a model organism. Here, we establish that C. elegans can be used as a model for the study of ATM by correcting the annotated gene structure of atm-1 and documenting the phenotypes arising in deletion mutants of this gene. Here, we have shown that the atm-1 gene model reported in wormbase (WS228) is incomplete, most notably lacking the large N-terminal region present in other ATM homologs.
We have shown that the publically available atm-1 gene model was incomplete as it lacked a large N-terminal region present in other ATM homologs (Fig. 1b). The revised model merges the atm-1 gene prediction with the upstream gene predictions K10E9.1 and F56C11.4 leading to a protein structure that is more in keeping with other ATM homologs. In addition to the size difference, we have identified previously unpredicted exonic sequences in the atm-1 transcript, refining the model further (Fig. 3d).
Regarding the nature of the gk186 mutation, one recent report identified a transcript that utilizes a cryptic splice site located in the exon affected by the deletion (Stergiou et al. 2007). Based on this translation, the authors predicted an ATM-1 protein lacking the kinase domain and suggested that atm-1(gk186) is a null mutation. In addition to the reported transcript (atm-1.a Figure S1) we have also identified a second transcript, atm-1(gk186).b. This transcript utilizes two alternative splice sites to produce an in-frame translation resulting in an internal deletion of 112AA. The translated protein preserves the PI3K kinase domain and might therefore retain some kinase function. Two independently derived mutant strains were sensitive to treatment with IR. In addition, both strains also display the hallmarks of genomic instability. We conclude that our results represent the null phenotype for the atm-1 gene and that the alternative transcript observed in the gk186-containing mutant is either non-functional or functional at an insignificantly low level.
In contrast to previous reports (Bailly et al. 2010), we find that atm-1(gk186) mutants are sensitive to IR. We have confirmed the IR sensitivity with the tm5027 allele. Experimental factors such as the brood size collected and the rest period might account for this discrepancy. The effect of IR on atm-1 mutants is not severe when compared to brd-1(gk99) mutants. brd-1 forms a heterodimer with brc-1 and is required for all homologous recombination (HR) repair of DSBs (Boulton et al. 2004). The fact that the atm-1 mutants are only partially sensitive to IR may indicate functional redundancy in repair of DSBs as proposed by Bailly et al. (2010).
We successfully used eT1 to capture lethal mutations generated in an atm-1-deficient background. The forward mutation frequency for atm-1(gk186) in this experiment was calculated to be 0.96%, which is similar to that reported for dog-1 (Cheung et al. 2002), and comparable to low dosages of other mutagens [1.5% for 0.004 M EMS, 0.96% for 500 Roentgens of gamma radiation, and 1.58% for 0.11% formaldehyde treatment (Rosenbluth and Baillie 1981)]. Since eT1 covers approximately one-sixth of the genome, this translates into a 7% forward mutation rate for the whole gene in the absence of ATM function. Thus, although not an essential gene, atm-1 clearly has a significant role in maintaining genomic stability.
In human A-T cell lines, reduced telomere length, increased rate of telomere shortening, and telomere fusions are observed (Metcalfe et al. 1996; Pandita et al. 1995; Xia et al. 1996). In both fission and budding yeast, a role for preventing telomere fusions and chromosome translocations has also been documented (Lee et al. 2008; Naito et al. 1998). No function for ATM-1 in telomere maintenance has been reported in C. elegans until now. Here, we have shown that atm-1 mutants can give rise to a heritable Him phenotype. Him phenotypes are known to arise in animals where X:autosome rearrangements and X:autosome telomeric fusions have occurred (Ahmed and Hodgkin 2000; Herman and Kari 1989; Herman et al. 1982). Ahmed and Hodgkin, have identified and characterized a mutant (mrt-2) that results in X:autosome fusions. mrt-2 encodes a highly conserved DNA-damage checkpoint protein homologous to RAD1. Him mutants arising in a mrt-2 genetic background also have a reduced number of diakinetic chromosomes (Ahmed and Hodgkin 2000) consistent with what we have observed in atm-1-derived Him animals. We conclude that these mutations are also likely to be X:autosome telomere fusions. The observation that less than five pairs of chromosomes are observed in some of the Him animals indicates that chromosomal fusions are not restricted to the X chromosome.
Reduced viability, a Him phenotype, and sterility are all indicators of genomic instability. We have documented independent lines of atm-1 mutants that display all of these phenotypes. That these mutations lead to inviability indicates that essential genetic material has been disrupted. Our characterization of the mutant ATM phenotype in C. elegans indicates that a range of chromosomal instabilities occurs that reflect a rich-mutagenic spectrum, not limited to chromosomal fusion events. In this regard, C. elegans provides a valuable model for the study of ATM function in a living animal and its role in preventing chromosomal abnormalities, providing insight to the function of atm-1 in promoting genomic stability.
Conclusion
We provide a revised structure for the C. elegans atm-1 gene that merges three gene annotations and reinterprets the splicing structure of the gene. The reinterpreted atm-1-coding region includes the ORFs F56C11.4, K10E9.1, and the current atm-1 gene prediction. Transcript analysis indicates that there is a single transcript produced that is required for response to ionizing radiation and maintenance of genomic stability. Lethal mutational events were isolated using the eT1 balancer system. Future work on atm-1 in C. elegans would provide insight into the DDR and intracellular signalling and control. C. elegans and atm-1 can possibly be used to understand the structure and function of telomere fusions with regard to tumourigenesis. Perhaps the most significant contribution to understanding atm-1, and ATM in general, would be discovering novel therapeutic targets for ATM-driven cancers. ATM interactions for synthetic lethality could lead to targeted therapy for appropriate cancers. Research in these areas would be greatly facilitated through C. elegans because it is amendable to high throughput, cost-effective methods of study. Our results establish C. elegans as an excellent model for the study of ATM function.
References
Ahmed S, Hodgkin J (2000) MRT-2 checkpoint protein is required for germline immortality and telomere replication in C. elegans. Nature 403(6766):159–164. doi:10.1038/35003120
Bailly AP, Freeman A, Hall J, Declais AC, Alpi A, Lilley DM, Ahmed S, Gartner A (2010) The Caenorhabditis elegans homolog of Gen1/Yen1 resolvases links DNA damage signaling to DNA double-strand break repair. PLoS Genet 6(7):e1001025. doi:10.1371/journal.pgen.1001025
Bosotti R, Isacchi A, Sonnhammer EL (2000) FAT: a novel domain in PIK-related kinases. Trends Biochem Sci 25(5):225–227 pii: S0968-0004(00)01563-2
Boulton SJ, Martin JS, Polanowska J, Hill DE, Gartner A, Vidal M (2004) BRCA1/BARD1 orthologs required for DNA repair in Caenorhabditis elegans. Curr Biol 14(1):33–39 pii: S0960982203008716
Brenner S (1974) The genetics of Caenorhabditis elegans. Genetics 77(1):71–94
Cheung I, Schertzer M, Rose A, Lansdorp PM (2002) Disruption of dog-1 in Caenorhabditis elegans triggers deletions upstream of guanine-rich DNA. Nat Genet 31(4):405–409. doi:10.1038/ng928ng928
Couteau F, Zetka M (2011) DNA damage during meiosis induces chromatin remodeling and synaptonemal complex disassembly. Dev Cell 20(3):353–363. doi:10.1016/j.devcel.2011.01.015
Garcia-Muse T, Boulton SJ (2005) Distinct modes of ATR activation after replication stress and DNA double-strand breaks in Caenorhabditis elegans. EMBO J 24(24):4345–4355. doi:10.1038/sj.emboj.7600896
Herman RK, Kari CK (1989) Recombination between small X chromosome duplications and the X chromosome in Caenorhabditis elegans. Genetics 121(4):723–737
Herman RK, Kari CK, Hartman PS (1982) Dominant X-chromosome nondisjunction mutants of Caenorhabditis elegans. Genetics 102(3):379–400
Johnsen RC, Baillie DL (1991) Genetic analysis of a major segment [LGV(left)] of the genome of Caenorhabditis elegans. Genetics 129(3):735–752
Kojis TL, Schreck RR, Gatti RA, Sparkes RS (1989) Tissue specificity of chromosomal rearrangements in ataxia-telangiectasia. Hum Genet 83(4):347–352
Kurz EU, Lees-Miller SP (2004) DNA damage-induced activation of ATM and ATM-dependent signaling pathways. DNA Repair (Amst) 3(8–9):889–900. doi:10.1016/j.dnarep.2004.03.029
Langmead B (2010) Aligning short sequencing reads with Bowtie. Curr Protoc Bioinformatics Chap 11:Unit 11 17. doi:10.1002/0471250953.bi1107s32
Lavin MF (2008) Ataxia-telangiectasia: from a rare disorder to a paradigm for cell signalling and cancer. Natl Rev Mol Cell Biol 9(10):759–769. doi:10.1038/nrm2514
Lavin MF, Shiloh Y (1997) The genetic defect in ataxia-telangiectasia. Annu Rev Immunol 15:177–202. doi:10.1146/annurev.immunol.15.1.177
Lavin MF, Delia D, Chessa L (2006) ATM and the DNA damage response. Workshop on ataxia-telangiectasia and related syndromes. EMBO Rep 7(2):154–160. doi:10.1038/sj.embor.7400629
Lee JH, Paull TT (2005) ATM activation by DNA double-strand breaks through the Mre11-Rad50–Nbs1 complex. Science 308(5721):551–554. doi:10.1126/science.1108297
Lee K, Zhang Y, Lee SE (2008) Saccharomyces cerevisiae ATM orthologue suppresses break-induced chromosome translocations. Nature 454(7203):543–546. doi:10.1038/nature07054
Lee SJ, Gartner A, Hyun M, Ahn B, Koo HS (2010) The Caenorhabditis elegans Werner syndrome protein functions upstream of ATR and ATM in response to DNA replication inhibition and double-strand DNA breaks. PLoS Genet 6(1):e1000801. doi:10.1371/journal.pgen.1000801
Lemmens BB, Tijsterman M (2011) DNA double-strand break repair in Caenorhabditis elegans. Chromosoma 120(1):1–21. doi:10.1007/s00412-010-0296-3
Metcalfe JA, Parkhill J, Campbell L, Stacey M, Biggs P, Byrd PJ, Taylor AM (1996) Accelerated telomere shortening in ataxia telangiectasia. Nat Genet 13(3):350–353. doi:10.1038/ng0796-350
Naito T, Matsuura A, Ishikawa F (1998) Circular chromosome formation in a fission yeast mutant defective in two ATM homologues. Nat Genet 20(2):203–206. doi:10.1038/2517
Negrini S, Gorgoulis VG, Halazonetis TD (2010) Genomic instability—an evolving hallmark of cancer. Natl Rev Mol Cell Biol 11(3):220–228. doi:10.1038/nrm2858
O’Neil N, Rose A (2006) DNA repair. WormBook:1–12. doi:10.1895/wormbook.1.54.1
Pandita TK, Pathak S, Geard CR (1995) Chromosome end associations, telomeres and telomerase activity in ataxia telangiectasia cells. Cytogenet Cell Genet 71(1):86–93
Robinson JT, Thorvaldsdottir H, Winckler W, Guttman M, Lander ES, Getz G, Mesirov JP (2011) Integrative genomics viewer. Nat Biotechnol 29(1):24–26. doi:10.1038/nbt.1754
Rosenbluth RE, Baillie DL (1981) The genetic analysis of a reciprocal translocation, eT1(III; V), in Caenorhabditis elegans. Genetics 99(3–4):415–428
Rotman G, Shiloh Y (1998) ATM: from gene to function. Hum Mol Genet 7(10):1555–1563 pii:ddb181
Seidel JJ, Anderson CM, Blackburn EH (2008) A novel Tel1/ATM N-terminal motif, TAN, is essential for telomere length maintenance and a DNA damage response. Mol Cell Biol 28(18):5736–5746. doi:10.1128/MCB.00326-08
Shiloh Y (2006) The ATM-mediated DNA-damage response: taking shape. Trends Biochem Sci 31(7):402–410. doi:10.1016/j.tibs.2006.05.004
Stergiou L, Doukoumetzidis K, Sendoel A, Hengartner MO (2007) The nucleotide excision repair pathway is required for UV-C-induced apoptosis in Caenorhabditis elegans. Cell Death Differ 14(6):1129–1138. doi:10.1038/sj.cdd.4402115
Tabara H, Motohashi T, Kohara Y (1996) A multi-well version of in situ hybridization on whole mount embryos of Caenorhabditis elegans. Nucleic Acids Res 24(11):2119–2124 pii: 6b0019
Trapnell C, Pachter L, Salzberg SL (2009) TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25(9):1105–1111. doi:10.1093/bioinformatics/btp120
Weidhaas JB, Eisenmann DM, Holub JM, Nallur SV (2006) A Caenorhabditis elegans tissue model of radiation-induced reproductive cell death. Proc Natl Acad Sci USA 103(26):9946–9951. doi:10.1073/pnas.0603791103
Xia SJ, Shammas MA, Shmookler Reis RJ (1996) Reduced telomere length in ataxia-telangiectasia fibroblasts. Mutat Res 364(1):1–11
Acknowledgments
This work was made possible by the following funding sources; NSERC scholarship JCH. CIHR Fellowship to MRJ. Operating funds CIHR to AMR and NSERC to DLB. The authors wish to give special thanks to J. Chu, S-Y. Chua, and H. Shin for unpublished RNA-seq data, and to S. Mitani for the knockout allele, tm5027. Name.
Open Access
This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by S. Hekimi.
M. R. Jones and J. C. Huang contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 2.0 International License (https://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Jones, M.R., Huang, J.C., Chua, S.Y. et al. The atm-1 gene is required for genome stability in Caenorhabditis elegans . Mol Genet Genomics 287, 325–335 (2012). https://doi.org/10.1007/s00438-012-0681-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-012-0681-0