Abstract
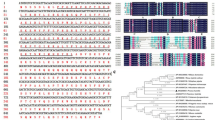
Members of the plant-specific gene family referred to as the NAC family (for NAM-ATAF-CUC-related) are involved in various functions including the regulation of plant development. However, no detailed molecular characterization of any member of the NAC family has yet been reported from monocots. Here, we report such a characterization of ONAC300, a novel NAC-family gene identified using a cDNA cloned from microdissected phloem cells of rice. The predicted ONAC300 protein sequence falls into the NAM subgroup, which also contains the proteins CUC1 and CUC2 from Arabidopsis, CUP from snapdragon, CmNACP from pumpkin and NAM from petunia. High levels of ONAC300 mRNA were detected by in situ hybridization in developing shoot apical meristem (SAM) and in the associated young leaves. The use of an ONAC300:: GUS reporter gene revealed that the ONAC300 promoter was expressed predominantly in developing vascular tissues of the leaves and roots. The construct was also expressed in anther filaments, rachis and carpel styles. RT-PCR analysis further revealed that the levels of ONAC300 transcripts were higher in leaves, roots and culms than in panicles. The observed expression pattern of ONAC300 is quite different from those of the dicot NAC genes previously reported. Thus, ONAC300 is a novel member of the NAC family which is expressed at very early developmental stages in the shoot, root and flower, as well as in the mature phloem of vascular tissues in rice.







Similar content being viewed by others
References
Aida M, Ishida T, Fukaki H, Fujisawa H, Tasaka M (1997) Genes involved in organ separation in Arabidopsis: an analysis of the cup-shaped cotyledon mutant. Plant Cell 9:841–857
Aida M, Vernoux T, Furutani M, Traas J, Tasaka M (2002) Roles of PIN-FORMED1 and MONOPTEROS in pattern formation of the apical region of the Arabidopsis embryo. Development 129:3965–3974
Asano T, Masumura T, Kusano H, Kikuchi S, Kurita A, Shimada H, Kadowaki K (2002) Construction of a specialized cDNA library from plant cells isolated by laser capture microdissection: toward comprehensive analysis of the genes expressed in the rice phloem. Plant J 32:401–408
Collinge M, Boller T (2001) Differential induction of two potato genes, Stprx2 and StNAC, in response to infection by Phytophthora infestans and to wounding. Plant Mol Biol 46:521–529
Ding B, Itaya A, Qi Y (2003) Symplasmic protein and RNA traffic: regulatory points and regulatory factors. Curr Opin Plant Biol 6:596–602
Duval M, Hsieh TF, Kim SY, Thomas TL (2002) Molecular characterization of AtNAM: a member of the Arabidopsis NAC domain superfamily. Plant Mol Biol 50:237–248
Greve K, La Cour T, Jensen MK, Poulsen FM, Skriver K (2003) Interactions between plant RING-H2 and plant-specific NAC (NAM/ATAF1/2/CUC2) proteins: RING-H2 molecular specificity and cellular localization. Biochem J 371:97–108
Ikeda K, Sunohara H, Nagato Y (2004) Developmental course of inflorescence and spikelet in rice. Breed Sci 54:147–156
Inada N, Sakai A, Kuroiwa H, Kuroiwa T (1998) Three-dimensional analysis of the senescence program in rice (Oryza sativa L.) coleoptiles. Planta 205:153–164
Ishida T, Aida M, Takada S, Tasaka M (2000) Involvement of CUP-SHAPED COTYLEDON genes in gynoecium and ovule development in Arabidopsis thaliana. Plant Cell Physiol 41:60–67
Jefferson RA, Kavanagh TA, Bevan MW (1987) GUS fusions: β-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J 6:3901–3907
Kadowaki K, Kubo N, Ozawa K, Hirai A (1996) Targeting presequence acquisition after mitochondrial gene transfer to the nucleus occurs by duplication of existing targeting signals. EMBO J 15:6652–6661
Kamiya N, Nishimura A, Sentoku N, Takabe E, Nagato Y, Kitano H, Matsuoka M (2003) Rice globular embryo 4 (gle4) mutant is defective in radial pattern formation during embryogenesis. Plant Cell Physiol 44:875–883
Kikuchi K, Ueguchi-Tanaka M, Yoshida KT, Nagato Y, Matsusoka M, Hirano HY (2000) Molecular analysis of the NAC gene family in rice. Mol Gen Genet 262:1047–1051
Kouchi H, Hata S (1993) Isolation and characterization of novel nodulin cDNAs representing genes expressed at early stages of soybean development. Mol Gen Genet 238:106–119
Nicholas KB, Nicholas HB Jr, Deerfield DW II (1997) GeneDoc: analysis and visualization of genetic variation. EMBnet NEWS 4:14
Ooka H et al (2003) Comprehensive analysis of NAC family genes in Oryza sativa and Arabidopsis thaliana. DNA Res 10:239–247
Ren T, Qu F, Morris TJ (2000) HRT gene function requires interaction between a NAC protein and viral capsid protein to confer resistance to turnip crinkle virus. Plant Cell 12:1917–1926
Ruiz-Medrano R, Xoconostle-Cazares B, Lucas WJ (1999) Phloem long-distance transport of CmNACP mRNA: implications for supracellular regulation in plants. Development 126:4405–4419
Ruiz-Medrano R, Xoconostle-Cazares B, Lucas WJ (2001) The phloem as a conduit for inter-organ communication. Curr Opin Plant Biol 4:202–209
Sablowski RW, Meyerowitz EM (1998) A homolog of NO APICAL MERISTEM is an immediate target of the floral homeotic genes APETALA3/PISTILLATA. Cell 92:93–103
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Souer E, van Houwelingen A, Kloos D, Mol J, Koes R (1996) The No Apical Meristem gene of Petunia is required for pattern formation in embryos and flowers and is expressed at meristem and primordia boundaries. Cell 85:159–170
Swofford DL (2003) PAUP*: Phylogenetic Analysis Using Parsimony (*and Other Methods). Version 4. Sinauer Associates, Sunderland
Takada S, Hibara K, Ishida T, Tasaka M (2001) The CUP-SHAPED COTYLEDON1 gene of Arabidopsis regulates shoot apical meristem formation. Development 128:1127–1135
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucl Acids Res 25:4876–4882
Vroemen CW, Mordhorst AP, Albrecht C, Kwaaitaal MA, de Vries SC (2003) The CUP-SHAPED COTYLEDON3 gene is required for boundary and shoot meristem formation in Arabidopsis. Plant Cell 15:1563–1577
Weir I, Lu J, Cook H, Causier B, Schwarz-Sommer Z, Davies B (2004) CUPULIFORMIS establishes lateral organ boundaries in Antirrhinum. Development 131:915–922
Xie Q, Sanz-Burgos AP, Guo H, Garcia JA, Gutierrez C (1999) GRAB proteins, novel members of the NAC domain family, isolated by their interaction with a geminivirus protein. Plant Mol Biol 39:647–656
Xie Q, Frugis G, Colgan D, Chua NH (2000) Arabidopsis NAC1 transduces auxin signals downstream of TIR1 to promote lateral root development. Genes Dev 14:3024–3036
Acknowledgements
We thank CAMBIA (Canberra, Australia) for providing pCAMBIA1301 and ZENECA MOGEN (Leiden, The Netherlands) for providing the Agrobacterium tumefaciens EHA105 strain. We also thank Drs. N. Kubo, K. Ono, K. Notsu, T. Nishikawa, K. Miyashita, N. Nohara and N. Kaji for technical support; and Drs. H. Ooka, S. Kikuchi and S. Kawakami for comments on this manuscript. This work was supported in part by a fellowship from the Japan Society for the Promotion of Science to H. K.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by R. Hagemann
Rights and permissions
About this article
Cite this article
Kusano, H., Asano, T., Shimada, H. et al. Molecular characterization of ONAC300, a novel NAC gene specifically expressed at early stages in various developing tissues of rice. Mol Genet Genomics 272, 616–626 (2005). https://doi.org/10.1007/s00438-004-1097-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-004-1097-2